Lus10011285 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10011285 pacid=23149888 polypeptide=Lus10011285 locus=Lus10011285.g ID=Lus10011285.BGIv1.0 annot-version=v1.0
ATGGACCCGTTGATGGAGCTGTGCGAGCTCATCGCGAAGAATCCAACGCAATTCAGCGATAAACTGACGTGGCTGTGCAACCGCTGCCCGCAGCCGGAGG
CGCTGCTCGGCGGATCGCCTAGGGTTTCGAGGTCTCAGCTCAATGCGGTACTCGCGGTTGCGAGGTTCCTCTCATTGGGGACGCCAGACGCGTCCGTCGA
TCTCCGGCCGAAGTCCATCGTGTTAGAGTTCCTCCACGCGATTCCGTCGTCGTTCTCGCAGTCATTCTGGCCGCAGTCGTACACGGAGGACTCGATTGGG
TCGTTCTACGGTGAGTTCCTGGGGCATTTGGTGAGTTGCGCTGAAACGTCGGCGGATTTCGGCTCGGAGATTGCTGGCTTTTTGGGAGAGGTTGTGATGG
AAGCGGTTAATTATCACAATGCTGGTGACGGTGTGGTGATTTCGAGGGACTTTTTGCTGGCGTTGACGGAGAATTTCCCTCCGATTTTGCCTTCTGATGG
GGAGAAGCTGGTGACTTGTCTTATTGATCAGGTCTTAGGGGCAGCCGGTGATCAAGCGGGGTTAAACTCGGGTACTTCGTCGTCTCAGAGCTCGCCGTTG
AATAAAGTCAATAATCATCAGCGTAACAATGGTATTTCCAGTCCGGCAAATGATCTGAGTGGGATATCTGGCGGGTCGTCCAGTGGATTGTCTACGACGT
CAGCATTGATGAACGGAAGTGGGTTGATGTGGAAGAGTGGGTTTGATTCGATGGGCATGGGCGTAGGGTTGAACGATGGTTTGTTGCGGCAGCAAGCGAC
TTTGTTTGAGGAGGAAAGTGTGGAGGGATTGGAGAAGCATGAGATCGCATTCAAGTTGATTGCTAATGTATTGACTCATGTGAAGCTTGACGTTAAGCTT
TCGGAACAAGTAAGATTGATTGCTAAGAAGCAGCTACAGTCTCTGTCGGCGTTTTTGAAGATAAGGAAGCGCGACTGGAATGAACAAGGACAACAACTGA
AAGCTAGGGTGAACCCTAAACTCTCAGTTTATCAAGCTGCAGCTAGAATGAAGATAAGGAGCCTTGCATCCCTTGATGCTGATGGTAAAACCTTCAAAAG
ATTGGTTCTTGAAACTCTTGCATTGCTTATCGATGCTGCTGAAGCATGTCTGCTATCTGTGTGGCGGAAGTTAAGAATATGCGAAGAGCTGTTTAGGTCA
TTGCTTGCTGGAATTGCACAAATTGCTGTGAGCAGGGGTGGTCAGCCACTTCGCGTTCTTCTTATTCGTCTTAAACCCCTTGTGTTGACAGCATGTGCAC
AAGCTGATACTTGGGATAGCAATCAGGGAGCTATGTTTGAAAGTGTGATGAAGACAAGCTGCCAGATAATTGAATCTGGATGGATAAAGGACCGAGCTCC
TGTCGATACTTTTATTTTGGGATTGGCAACAAGCATACGTGAGCGGAACGACTATGATGAGCAGGCTGAGAAGGAAAAACAGGGGGTTCCTCCTGTGCAG
CTCAATCTCATACGTTTGCTAGCTGATTTGACCGTGGCTGTTAACAAGTCGGAAGTGGTTGACATGATTTTGCCACTTTTTGTTGAAAGCTTAGAGGAGG
GTGATGCTTTAACTCCTGGCATATTTCGACTTCGACTTCTTGATGCTGTATCGCGGATAGCAAGTCTAGGGTTTGAGAAATCCTATCGTGAGACAGTTGT
CCTGATGACAAGAAGTTACTTGGGTAAACTGTCAAGTGTAGGATCTGCTGAAAACACAACTTTGCCAGCAGAAGCCAATACAGAACGTGTTGAGACTCTT
CCTGCAGGATTCCGTTTGATAGCTAGTGGTCTCAAAAACCCAAAATTACGTGTAGATTACCGACATCGTCTTCTCTCTTTATGCTCAGATGTGGGGCTGG
CAGCTGAATCCAAGAGTGGAAGGAGTGGAGCAGATTTTATGGGTCCTCTACTTCCTGCTGTTGCTGAAGTCTGCTCCGATTTTGATCCTACAGTGAATGT
CGAACCGTCACTTCTGAAACTCTTCCGTAACTTGTGGTTCTATGTTGCTCTGTTTGGTTTGGCTCCACCGATTGTGAAAACTCAACAACAGGTGAAGTCC
ACATCAAGCACGCTGAATAGTGTAGGAAGCAATGGTGCACTTTCTCTTCAGGCTGTGGGTGGACCATACATGTGGAATGCACAGTGGTGTTCTGCTGTCC
AACGTATTGCTCAAGGGACTCCTCCTCTTGTTGTCAGTTCTGTCAAATGGCTTGAAGATGAACTGGAACTCAATGCACTCCACAATCCTGGAAGCCGTCA
AGCAAGTGGCAATGAGAAAGCTGCTCTAACCCAACGATCAGCTCTGTCAGCTGCATTGGGTGGCAGAGTGGAGGTTGCAGGAATGACCACAATCTCTGGT
GTTAAAGCAACATATCTGCTTGCTGTAGCTTTTCTGGAGATAATTCGGTTCAGCAGCAATGGTGGAGTTCTCAATGGGGATACCAGTTTAACTGCTTCAA
GAAGTGCCTTCAGCTGCGTATTTGAATATCTAAAAACTCCAAACTTGATGCCTGCTGTGTTTCAATGTCTGACTGCTATAGTGCATCGAGCATTTGAAGC
AGCTGTATCATGGCTGAAAGAACGGATAACTGACACTGGCAAAGATGCTGCAATCAGGGAATTAACTCTTTTTGCCCATGCATGTTTTCTGATAAAAAGC
ATGTCACTGAGAGAGGAGCACATCCGCGATATTGCTGTTAACTTGTTGACTCAGCTTCGAGATAAGTTTCCTCAGATCTTATGGAACTCGTCCTGTTTGG
ACTCCCTATTATTTTCTGTTCACAACGACTCAACTTCTACCGCTGTTAATGATCCCTCTTGGGTAGCAACTGTTCGGGCATTGTACCAGAAAATTCTCAG
GGAGTGGATCTGCATGTCGCTCTCATATGCTCCATGTACTAGCCAAGGTCTTCTTCAGGAAAAGCTCTGCAAGGCAAATACTTGGCAGAAGGCACAGCAA
ACTACTGATGTGCGGGGGGGCCGGGGCGCAGCGGCAGCTGCTGCCTCAGGAGCAAACTTGACATTGACAGAATCCTTCAACTTTGAGGTGCTCAGCACTG
GCATAGTCAGTGCAACAGTCAAGTGTAACCATGCTGGGGAAATCGCTGGTATGAGAAGGTTATATAGCAGTATTGGTGGTTTTCAATCTGGTACTCCGCC
GACAACTTTTGGTGGGGGTCTCCAGCGGTTGATAACTGGAGCATTTGCTCAACCGCCAGAGACTGAAGATGATTCTTTCAATGAGATGTTACTCAATAAG
TTTGTGCATCTACTCCAGCAATTTGTTACTGCTGCGGAGAAAGGTGGGGATGTGGAAAAATCACAGTTTCGTGATATTTGTTCTCAGTCAACAGCGTTAC
TTCTATCCAATCTGACTTCTGAATCAAAATTCAACATAGAGGGCTTTGCGCAACTGCTACGCCTTCTTTGCTGGTGTCCGGCATACATATCTACTCCAGA
TGCAATGGAGACTGGTGTCTTTATATGGACTTGGTTAGTTTCTGCAGCACCCCAACTTGGATCTCTTGTTCTTGCCGAACTCGTGGACGCCTGGTTATGG
ACTATTGACACAAAGCGAGGGCTATTTGCATATGATGTCAAGTACTCTGGACCTGCTGCGAAACTAAGGCCTCAGCTCGCTCCTGGAGAACCAGAGCCAC
AGCCTGAAATTGATCCTGTTGAACAGATCCTCGCTCATAGACTTTGGCTTGGCTTTTTCATTGATCGGTTTGAGGTAGTGCGGCATAACAGTGGCGAGCA
ACTCTTGCTCTTGGGTAGGCTATTGCAAGGGACAATGAGGTTACCTTGGAATTTTTCACGCCATCCTGCTGCTACTGGTGCCTTTTTCACAGTCATGCTC
CTTGGGCTCAAATTCAGCTCATGTCATTCTCAGGGAAACCTGCAGAATTATAAAACCGGGCTTCAACTACTGGAAGATCGGATTTATAGAGCCTGTTTGG
GCTGGTTTGCTTTTGAACCTGAGTGGTACAATGTGAACTATATGAATTTTGCTCAGAGTGAGGCTCAATCTGTATCAGTATTTGTTCACTATCTTTCAAA
TGAACGAGGGGATGCTCCTGTATCTGACTCTAAGGGACGCGGGAGTGACAATGGAAGTTCGTTGGTTGATATGAATGATGAACATCACCCTGTGTGGGGT
CACATGGAGAATTATGCTACCGGAAGGGAAAAGCGGAAACAGTTACTATTGATGCTATGTCAGCATGAGGCTGATAGGCTTGAAGTGTGGGCGCAACCTA
CTAACTCCAAGGAAAATGTTTCGCGTCCCAAAATCAGCTCAGATAAATGGATAGAATATGCTAGGACAGCTTTCTCCGTGGATCCCCATATTGCTTTGTG
CATGCCGTCAAGGTTCCCTGCAAATACTACTTTGAAGGCTGAAGTAACACAGTTGGTCCAGTCACATATTCTTGACGTGCGCACTTTACCAGAAGCACTT
CCGTACTTTGTCACCCCAAAAGCTGTTGATGAAGATTCAGTGCTTTTGCAACAACTGCCACACTGGGCTGGTTGTACAATAACACAAGCACTAGAGTTCC
TAACTCCTGCATACAAGGGTCATCCTCGTGTGATGGCTTATGTTCTGAGAGTTTTGGAGTCTTATCCTCCTGAGCGTGTAACATTTTTCATGCCACAATT
AGTCCAGTCTCTGCGGTATGATGATGGGAAGTTAGTCGAAGGATATTTGTTGAGAGCAGCTCAGAGAAGTGATATCTTTGCACATATTCTGATTTGGCAT
TTGCAGGGAGAACAACCTGAAACAGATGCAGTGAAAGATGGTGCAGTCCCAAAGGATGATGCCTTTCAAGAACTCTTGCCTGTTGTTCGTGAGCATATCG
TTGATGGTTTCAATGACAAGGCCCGTGATGTTTTCGAAAGGGAGTTTGACTTTTTCGATCAAGTTACGTCAATCTCAGGTGTCCTCCTTCCACTCCCCAA
GGATGAGCGTCGGGCTGGTATTCGGACGGAATTGGAGAAAATCAAAATGCAAGGAGAAGATCTTTATTTACCAACTGCAACGAACAAACTTGTCAGAGGT
ATCCGGGTGGATAGTGGCATACCCTTGCAATCAGCAGCGAAAGTCCCGATCATGGTTACCTTTGATGTGGTTGATCGTGATGGCAGCCCTAATGATATTA
AGCCTCAAGCTTGCATTTTCAAGGTTGGAGATGATTGCAGACAAGATGTGCTTGCCCTTCAAGTGATAACGCTTCTTAGAGACATCTTTGGTGCTGTCGG
CCTTAATCTCTATTTGTTTCCCTACGGTGTCTTGCCAACGGGGCCAGGGAGAGGTATTATCGAGGTTGTGCCTAACTCACGGAGTCGAAGTCAGATGGGC
GAAACAACAGATGGTGGTTTGTACGAGATCTTTCAGCAGGACTTTGGTCCTGTCGGCTCCCCTAGTTTCGAAGCCGCTCGTGAAAACTTCATCATCAGCA
GCGCCGGTTATGCAGTTGCCAGTCTTCTACTCCAACCGAAGGACAGACACAACGGGAACTTGCTTTTCGACAGCGATGGGAGGCTCGTCCACATTGATTT
TGGATTCATATTCGAAATCTCCCCCGGTGGGAACATGCGGTTCGAGAGTGCACATTTCAAACTCAGCCACGAGATGACTCAGTTACTCGACCCATCAGGC
GTCATGAAGAGCGAGACATGGCACCAATTCGTCAGCCTGTGCGTGAAAGGGTACCTCGCTGCACGCAGGTACATGATGATGCTCGACAGTGGCCTACCTT
GCTTCAGTCGAGGAGACCCGATCGGGAACCTCCGGAAAAGGTTCCACCCGGAAATGAGCGAGAGGGAAGCCGCGAATTTCATGATCCATGTCTGCACAGA
CGCCTACAACAAATGGTCAACTGCAGGCTACGACTTGATACAGTATCTGCAGCAAGGAATTGAGAAGTGA
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10011285 pacid=23149888 polypeptide=Lus10011285 locus=Lus10011285.g ID=Lus10011285.BGIv1.0 annot-version=v1.0
MDPLMELCELIAKNPTQFSDKLTWLCNRCPQPEALLGGSPRVSRSQLNAVLAVARFLSLGTPDASVDLRPKSIVLEFLHAIPSSFSQSFWPQSYTEDSIG
SFYGEFLGHLVSCAETSADFGSEIAGFLGEVVMEAVNYHNAGDGVVISRDFLLALTENFPPILPSDGEKLVTCLIDQVLGAAGDQAGLNSGTSSSQSSPL
NKVNNHQRNNGISSPANDLSGISGGSSSGLSTTSALMNGSGLMWKSGFDSMGMGVGLNDGLLRQQATLFEEESVEGLEKHEIAFKLIANVLTHVKLDVKL
SEQVRLIAKKQLQSLSAFLKIRKRDWNEQGQQLKARVNPKLSVYQAAARMKIRSLASLDADGKTFKRLVLETLALLIDAAEACLLSVWRKLRICEELFRS
LLAGIAQIAVSRGGQPLRVLLIRLKPLVLTACAQADTWDSNQGAMFESVMKTSCQIIESGWIKDRAPVDTFILGLATSIRERNDYDEQAEKEKQGVPPVQ
LNLIRLLADLTVAVNKSEVVDMILPLFVESLEEGDALTPGIFRLRLLDAVSRIASLGFEKSYRETVVLMTRSYLGKLSSVGSAENTTLPAEANTERVETL
PAGFRLIASGLKNPKLRVDYRHRLLSLCSDVGLAAESKSGRSGADFMGPLLPAVAEVCSDFDPTVNVEPSLLKLFRNLWFYVALFGLAPPIVKTQQQVKS
TSSTLNSVGSNGALSLQAVGGPYMWNAQWCSAVQRIAQGTPPLVVSSVKWLEDELELNALHNPGSRQASGNEKAALTQRSALSAALGGRVEVAGMTTISG
VKATYLLAVAFLEIIRFSSNGGVLNGDTSLTASRSAFSCVFEYLKTPNLMPAVFQCLTAIVHRAFEAAVSWLKERITDTGKDAAIRELTLFAHACFLIKS
MSLREEHIRDIAVNLLTQLRDKFPQILWNSSCLDSLLFSVHNDSTSTAVNDPSWVATVRALYQKILREWICMSLSYAPCTSQGLLQEKLCKANTWQKAQQ
TTDVRGGRGAAAAAASGANLTLTESFNFEVLSTGIVSATVKCNHAGEIAGMRRLYSSIGGFQSGTPPTTFGGGLQRLITGAFAQPPETEDDSFNEMLLNK
FVHLLQQFVTAAEKGGDVEKSQFRDICSQSTALLLSNLTSESKFNIEGFAQLLRLLCWCPAYISTPDAMETGVFIWTWLVSAAPQLGSLVLAELVDAWLW
TIDTKRGLFAYDVKYSGPAAKLRPQLAPGEPEPQPEIDPVEQILAHRLWLGFFIDRFEVVRHNSGEQLLLLGRLLQGTMRLPWNFSRHPAATGAFFTVML
LGLKFSSCHSQGNLQNYKTGLQLLEDRIYRACLGWFAFEPEWYNVNYMNFAQSEAQSVSVFVHYLSNERGDAPVSDSKGRGSDNGSSLVDMNDEHHPVWG
HMENYATGREKRKQLLLMLCQHEADRLEVWAQPTNSKENVSRPKISSDKWIEYARTAFSVDPHIALCMPSRFPANTTLKAEVTQLVQSHILDVRTLPEAL
PYFVTPKAVDEDSVLLQQLPHWAGCTITQALEFLTPAYKGHPRVMAYVLRVLESYPPERVTFFMPQLVQSLRYDDGKLVEGYLLRAAQRSDIFAHILIWH
LQGEQPETDAVKDGAVPKDDAFQELLPVVREHIVDGFNDKARDVFEREFDFFDQVTSISGVLLPLPKDERRAGIRTELEKIKMQGEDLYLPTATNKLVRG
IRVDSGIPLQSAAKVPIMVTFDVVDRDGSPNDIKPQACIFKVGDDCRQDVLALQVITLLRDIFGAVGLNLYLFPYGVLPTGPGRGIIEVVPNSRSRSQMG
ETTDGGLYEIFQQDFGPVGSPSFEAARENFIISSAGYAVASLLLQPKDRHNGNLLFDSDGRLVHIDFGFIFEISPGGNMRFESAHFKLSHEMTQLLDPSG
VMKSETWHQFVSLCVKGYLAARRYMMMLDSGLPCFSRGDPIGNLRKRFHPEMSEREAANFMIHVCTDAYNKWSTAGYDLIQYLQQGIEK
|
DESeq2's median of ratios [FLAX]
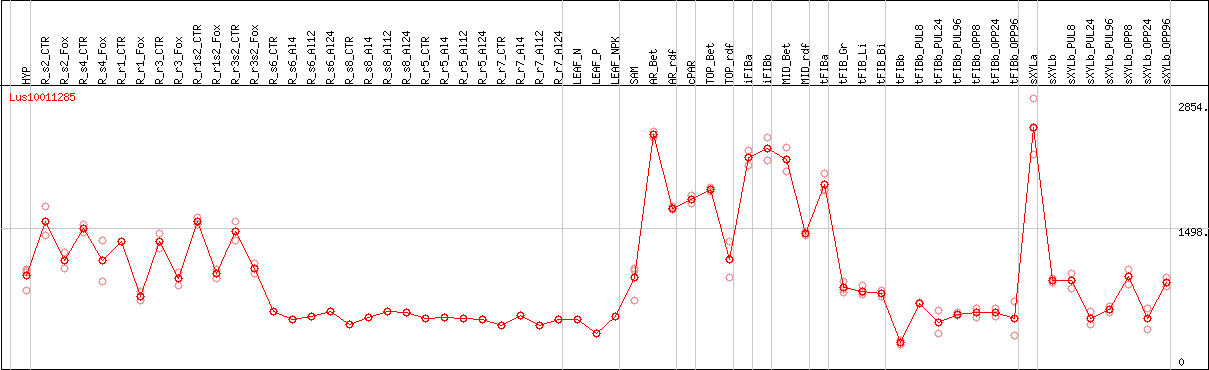
Coexpressed genes
Lus10011285 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
