External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G52330 157 / 5e-48
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1), Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.2)
AT4G13270 129 / 4e-37
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10037534
331 / 2e-116
AT1G52330 160 / 2e-49
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1), Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.2)
Lus10043216
134 / 6e-39
AT4G13270 228 / 8e-76
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Lus10011095
132 / 3e-38
AT4G13270 234 / 3e-78
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Lus10023631
131 / 9e-38
AT4G13270 235 / 2e-78
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Lus10024264
104 / 2e-27
AT4G13270 210 / 8e-69
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Lus10003238
41 / 0.0003
AT3G54200 143 / 3e-42
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Lus10040941
40 / 0.0006
AT3G54200 63 / 1e-11
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G055000
188 / 4e-60
AT1G52330 181 / 1e-57
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1), Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.2)
Potri.001G180600
187 / 1e-59
AT1G52330 192 / 9e-62
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1), Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.2)
Potri.006G159400
127 / 1e-36
AT4G13270 228 / 3e-76
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Potri.016G142300
60 / 7e-11
AT3G54200 162 / 3e-50
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Potri.006G112800
50 / 2e-07
AT3G54200 159 / 7e-49
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0159
E-set
PF03168
LEA_2
Late embryogenesis abundant protein
Representative CDS sequence
>Lus10011458 pacid=23182637 polypeptide=Lus10011458 locus=Lus10011458.g ID=Lus10011458.BGIv1.0 annot-version=v1.0
ATGGCGGCCGCCGCCGAACAATCCGATCCAAAATCGTCACAACCCGACAGCTCAGTTCCCCTGTTATCATCCGCAGCACCCATCTCACCAACCCAAAACG
ACGCCGTTCGGTACGCTTCCCGCAGCCGACGGCGGACAATACTCACCACCACTGCCACGGTGGCAGCATCTCTCACCGTTTTCCTCCTCATCCTCGCATC
TGGCTACGAATTCTGGCCGTCGGATCCGACACTGACGATCGCACGGCTGCGGCTGACCAAGGTCAAAATCCGGACATTCCCTGTCCTGAACATCGACATA
ACAATGAACGTGACGATGGAGGTGCGGAACGCCGACGTCTACACGGCGGAGCTGAGGAGGATGGATGTGGAGGTGATGTACAGAGGGAAGCGGATCGGGG
AGGTGAAAATCGGAAAGGAGGAAGTGATCCGAGCTTATGCAACGTCGTATTTGGAGGCGGAGATGAAGCTGAACGGGATCGGAGTGATCTCCGACGCGCT
GTTGCTGATCGGAGACTTGAAGAAGGGGAAAGTGCCGTTTGGTACGGTTGTTGAAGTCGGTGGGAAGTTTGGTTTCCTCTGTTTTGGGTTCCCTTTGAAG
GCGAAATTGTCATGCCAAGTTTTGCTCAACGCCCACAATCAAACCATCGTCCGCCAGAGCTGCTACCCTCTCTAG
AA sequence
>Lus10011458 pacid=23182637 polypeptide=Lus10011458 locus=Lus10011458.g ID=Lus10011458.BGIv1.0 annot-version=v1.0
MAAAAEQSDPKSSQPDSSVPLLSSAAPISPTQNDAVRYASRSRRRTILTTTATVAASLTVFLLILASGYEFWPSDPTLTIARLRLTKVKIRTFPVLNIDI
TMNVTMEVRNADVYTAELRRMDVEVMYRGKRIGEVKIGKEEVIRAYATSYLEAEMKLNGIGVISDALLLIGDLKKGKVPFGTVVEVGGKFGFLCFGFPLK
AKLSCQVLLNAHNQTIVRQSCYPL
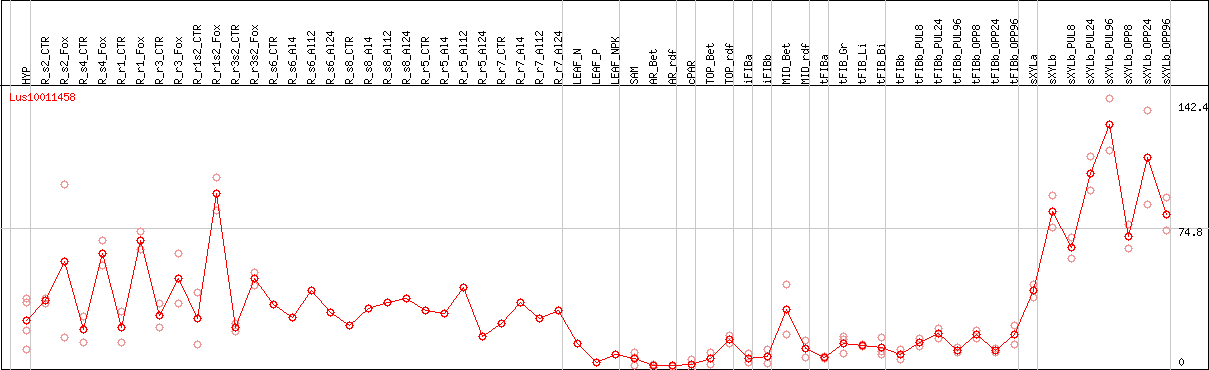
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10011458 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.