Lus10011466 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10011466 pacid=23182630 polypeptide=Lus10011466 locus=Lus10011466.g ID=Lus10011466.BGIv1.0 annot-version=v1.0
ATGTCCAAGAGAAAATTCGGATTCGAAGGATTCGGCATCAACCGTCAAACCAGCTACAACTTCGAGCGATCTCAAGCTCCACAGCGGCTCTACGTCCCAC
CTTCATCTCGCCACAGCCATGACAACTACGAAGATACGGACATCGACAACATCGATTACGACGACACCAAGGAGACCACAAGCGATGGCGAGCCTGACAT
CAGCAACAACAACAATAGCAACAGCAATACCGCTGGCGTTGAAGATGACGAAGTTGATCCACTGGACGCGTTCATGGAGGGGATACACGAAGAGATGAAA
GCAGCGCCTCCGCCGAAGGCTAAGGATAAAGTGGACAAGTATCGGGATGACGAGGAGGATGATCCCATGGAGAGCTTCTTGAGTGCGAAGAAAGATATAG
GGTTGACGTTGGCTGCCGATGCTCTCAACGCTGGATACAATTCCGACGAGGAGGTTTACGCAGCTGCAAAAGCTGTGGATGCTGGATTGCTGGATTATGA
TTCGGATGATAATCCGGTGGTGGTTGATAAGAAGAAGATTGAGCCCCTTACCGCTTTAGACCATAGTTCCATTGATTATGAACCTTTCAACAAGGATTTT
TACGATGAAAAGGAGTCCATAGCAGGGATGAGCGGGCAGGATGTGGCTGAGTACAGGAACACTCTAGCTATTCGTGTTTCGGGCTTTGATGTTCCGAGGC
CAGTTAAAACGTTTGAAGATTGTGGATTTTCTACACAACTGATGGGTGCTATTGCTAAACAAGGTTATGAAAAGCCTACCCCCATACAATGTCAAGCTTT
GCCTATTGTGCTCTCTGGCAGAGACATCATTGGTATAGCAAAAACAGGCTCTGGTAAGACTGCTGCTTTTATACTTCCCATGATTGTACATATCATGGAC
CAACCTGAACTTCAGAAAGAAGAAGGGCCAATTGGAGTTATTTGTGCACCTACTAGGGAGTTAGCTCATCAAATATACTTGGAGGCAAAGAAATTTGCTA
AACCGTATGGGATACGGGTCTCTGCTGTCTATGGTGGAATGTCGAAGCTTGAGCAGTTTAAAGAACTCAAAGCTGGTTGTGAAATGGTTGTTGCTACTCC
TGGTAGATTGATTGACATGATAAAAATGAAAGCAGTGAATATGTTGAGAGCAACTTGCTTGGTACTTGATGAGGCAGATAGGATGTTTGACCTAGGGTTT
GAGCCTCAAATAAGGTCTATTGTTGGACAGATTAGGCCAGATCGGCAGACTCTACTCTTCTCGGCTACCATGCCGTTCAAAATTGAGAAGTTAGCCAGAG
AGATTCTTTCTGATCCCGTTAGAGTCATCGTAGGAGAGGTGGGTATGGCCAATGAAGATATTACTCAAGTAGTCCAAGTGATTCCCTCTGATGCAGAGAA
GCTTCCCTGGCTTCTCGAAAAGCTACCCCCTATGATTGATGATGGAGATGTTCTAGTTTTTGCTTCGAAAAAGGCCACTGTTGATGAGATTGAATCACAG
TTGGTCAAGAAGGAGTTCAAAGTTGCAGCTCTTCATGGCGACAAAGATCAGTCTTCAAGAATGGAGATCTTGCAGAAATTCAAATCTGGGGTCTACCATG
TTCTTATTGCAACTGATGTTGCTGCTCGTGGATTAGATATTAAATCAATCAAGTCGGTTGTGAACTTTGACGTTGCAAAAGATATGGACATGCATGTTCA
TCGTATTGGCAGGACTGGTCGTGCTGGCAATAAAGATGGGATTGCTTATACTCTTATAACCCAGAAGGAGGCACGGTTTGCTGGGGAACTGGTTAACAAT
CTGGTTGCTGCTGGTCAAAATGTCTCAGCAGAACTCATGGATCTTGCTATGAAGGATGGAAGATTCAGGTCGAAACGTGACGCAAGAAAAGGAGGTGGTA
AGAAAGGCAAAGGCAGGGGTGGGAGTGGTCGAGGAGTTCGTGGCGTGGATTTCGGTCTTGGCATCGGCTACAATTCTGAAACCACCGGCACCTCATCTCA
CGACAATAGACCCAGTAAATCTGCAGGTTTCAGTTCAGTCCGGACAGGGATGACGGGACAGTTCAAGAACAACTTTGTTTCTGCTTCGTCAAATTCTCAG
TCTGCTGGATTTAGTAACAACGGCTCGAGTGCTGCTCCTCCCAGCAAAAGGCCAGCATTACGTGGATTCGTGTCCGGCGGTTCAATAGGAGGCGATGCAA
ATAGAGTCCAGCCGACTACCAATACACTATCTGGGTTCGTCTCAGCCGGCTCAATAACTGGAGATGCAAACAGAACATCTCAGAACCCCTCCGGAAATTC
CAACCAGAAAACACCAGAGAGAGCCGCTCTCAACTTGTTGGCTAGCTTAGGTGTGGTTCAAAGTCTTATTTGTCCTACTGTTGACACTGTGAGGGGTGGG
AGTCGGTCGGCCGTTCCGCAGGGCGAGAAAGGCGGAGACGCTCCGGTTGGGACCGTTGAGGACAATGTTAAGAACATCAAAACACAGGGTAGCCGATCTG
CACGAAGCCTCGCATGGACCGACCGGAAGTCTACGTCGTCGGTGTTGTCACTAAGGTGGACGGGTAGAGTCATTCAACTCACATGCAGCTCGAAGCTCGG
ATTCTGTGGCTCAGTCGACCGTGATGCCAGTCTGTTGGGTTCCTTCAAGCCAGTTGCGCCGATTGAAGAGTTGGTGATCCCCGCTTGTGGTGTGAATCCG
CGAGGAGCAACTAATAATATCACAGGCTGTTGGGCAAGGTTGACAATTAGGGCAGCTGTTGAGAGCAAAACCGGCAACTATACTCGCAATATTAATTGCG
TCGCAGGACCACCGAGAAGGCTGTGGTATTTATGGAGGATTCTATTTGTAATCTTCGGCTCAGTGATTCTCATCACTGATACAGTTGATTCAGCTGGCGT
TAGTGTGGCAGAGCATCATAAATCTCACAGGAAGCATTTCTCGCCTCTTGGTTCACCGGCATTTGTGGTTGCTCCAGCTCAGCCTCCTGACTATGATCCA
TTGATAACTTCTGGTCACCCACCAACGAGTTCGAGCTTGTCGAAGCCTTCGATGAAGAAGACACCGGGTGTTGGGTTGGTTGCTCTTGCCCCAACGCAGC
CCCGTGCTAGCGACAACAATTCGGGCCTAGCTCAGCCTCCTTTGTCTCCAGATGTCTCTGATTGTTGTGGACCAGAGCTAGTGGTGAAGAGAGGGGTTCA
TGGCTGTCATTGTGTGTATCCTATAAAGCTTGACCTTCTGCTGTCGAATGTTTCCCAAAATCATAATTGGGATTTGTTCCTTCAGGAGTTAGCTTCTGAG
CTGGGTTTGTTGGTTTCTCAAATTGAGCTGAGAAACTTCTATTTTCTCACCCTATCGAGATTAAACATTTCGATGGATATTATTCCTCACTCCGGTGACA
GCTTTCCTGCCAGTGATGCATACAAAATAAACTCTTCCCTTTCCTTGCATCAAGTCCGCTTCGACCCTAACTTTGTTGGTGACTACAAACTTCTTAATCT
TACATGGTTTGAACCGCCACCGCCTAGCCCAGCTCCTGTTGCCATTTCATCACCTCCTCAAGCACCATCGCATCCTTTGTCAAATTCTACATCAGCAAGC
ATTCCAAACAGAAGCAAGCATTCATATACACTTTTAGTACTCATTTTTGTGGGCATTGCAGTCGTGATTGTAACCATTATATGTACGCTTGTGGTTTGCT
CCCGCTTGTCACGCAGAAGGAGAACTGAACCACCATCTGAAGAACCTGAGAAGCCGAGGCCTACATATGCAGTTCCTGTTGCAGGACCCATGCCACATCC
TTCAAGTACACGTTTCCTATCTTATGAAGAACTCAGGGAAGCAACCGGTAACTTCTCGACTGCTAGCATACTTGGAGAAGGTGGGTTTGGTAAGGTTTTC
AAAGGTGTCCTTAGTGACGGTACAGCTGTTGCGATCAAGAGGCTCACCTGTGGAGGCCAACAAGGGGATAAAGAGTTCTTAGTTGAGGTTGATATGTTGA
GTAGATTGCATCATCGCAATCTTGTCAAACTCGTTGGCTACTATAGTAGTCGTGAAACTTCACAGAACCTCCTTTGTTATGAGCTTGTTCCCAATGGAAG
CTTAGAGGAGTGGCTCCATGGTCCTCTAGGTGCTAATTCTCCTCTTCTTTGGGACACCAGAATGAAGATCGCTTTGGATGCCGCGAGAGGACTCGCTTAC
CTACATGAGGACTCGCAGCCGAGTGTTATCCATAGGGATTTCAAGGCATCCAACATCCTTCTGGAGAACAATTTCCTTGCTAAAGTCGCGGATTTCGGAC
TTGCGAAACAGGCACCAGAAGGCAGAGTGAACTATCTATCTACTCGAGTGATGGGAACATTCGGATATGTTGCGCCCGAGTATGCCATGACCGGACATTT
ACTGGTGAAGAGTGATGTATATAGCTATGGAGTTGTGCTGCTCGAGTTGCTTACTGGGAGGAAGCCTGTGGATATGTCGCAACCATCTGGACAGGAGAAC
CTTGTTACTTGGACAAGACCCATTCTCAGAGACAAGGACAGGCTAGAGGAGCTCGCGGATCCGAGGCTAGTCGGGAAGTACCCGAAGGAAGACTTCGTGC
GAGTCTGCACCATAGCAGCAGCCTGCGTAGCCCCTGAGGCAAGCCAGCGGCCCACCATGGGAGAAGTGGTGCAGTCGCTGAAAATGGTGCAGCGGGTGAC
CGAGTACCAAGACGCTGCTGCTGCGGCAGCTGCTGCATCGAGTGCCAGGCCGAACACGAGACAGTCGTGCACCACCTTCGACTCGGACGGGACATCATCG
ATGTTCTCGTCGGGTCCTTTCTCCGGGTTGAGCAACTTCGACAACGAGCGGACTGCAGCCTTCTCGGAGGATCTCCACGAAGGAAGGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10011466 pacid=23182630 polypeptide=Lus10011466 locus=Lus10011466.g ID=Lus10011466.BGIv1.0 annot-version=v1.0
MSKRKFGFEGFGINRQTSYNFERSQAPQRLYVPPSSRHSHDNYEDTDIDNIDYDDTKETTSDGEPDISNNNNSNSNTAGVEDDEVDPLDAFMEGIHEEMK
AAPPPKAKDKVDKYRDDEEDDPMESFLSAKKDIGLTLAADALNAGYNSDEEVYAAAKAVDAGLLDYDSDDNPVVVDKKKIEPLTALDHSSIDYEPFNKDF
YDEKESIAGMSGQDVAEYRNTLAIRVSGFDVPRPVKTFEDCGFSTQLMGAIAKQGYEKPTPIQCQALPIVLSGRDIIGIAKTGSGKTAAFILPMIVHIMD
QPELQKEEGPIGVICAPTRELAHQIYLEAKKFAKPYGIRVSAVYGGMSKLEQFKELKAGCEMVVATPGRLIDMIKMKAVNMLRATCLVLDEADRMFDLGF
EPQIRSIVGQIRPDRQTLLFSATMPFKIEKLAREILSDPVRVIVGEVGMANEDITQVVQVIPSDAEKLPWLLEKLPPMIDDGDVLVFASKKATVDEIESQ
LVKKEFKVAALHGDKDQSSRMEILQKFKSGVYHVLIATDVAARGLDIKSIKSVVNFDVAKDMDMHVHRIGRTGRAGNKDGIAYTLITQKEARFAGELVNN
LVAAGQNVSAELMDLAMKDGRFRSKRDARKGGGKKGKGRGGSGRGVRGVDFGLGIGYNSETTGTSSHDNRPSKSAGFSSVRTGMTGQFKNNFVSASSNSQ
SAGFSNNGSSAAPPSKRPALRGFVSGGSIGGDANRVQPTTNTLSGFVSAGSITGDANRTSQNPSGNSNQKTPERAALNLLASLGVVQSLICPTVDTVRGG
SRSAVPQGEKGGDAPVGTVEDNVKNIKTQGSRSARSLAWTDRKSTSSVLSLRWTGRVIQLTCSSKLGFCGSVDRDASLLGSFKPVAPIEELVIPACGVNP
RGATNNITGCWARLTIRAAVESKTGNYTRNINCVAGPPRRLWYLWRILFVIFGSVILITDTVDSAGVSVAEHHKSHRKHFSPLGSPAFVVAPAQPPDYDP
LITSGHPPTSSSLSKPSMKKTPGVGLVALAPTQPRASDNNSGLAQPPLSPDVSDCCGPELVVKRGVHGCHCVYPIKLDLLLSNVSQNHNWDLFLQELASE
LGLLVSQIELRNFYFLTLSRLNISMDIIPHSGDSFPASDAYKINSSLSLHQVRFDPNFVGDYKLLNLTWFEPPPPSPAPVAISSPPQAPSHPLSNSTSAS
IPNRSKHSYTLLVLIFVGIAVVIVTIICTLVVCSRLSRRRRTEPPSEEPEKPRPTYAVPVAGPMPHPSSTRFLSYEELREATGNFSTASILGEGGFGKVF
KGVLSDGTAVAIKRLTCGGQQGDKEFLVEVDMLSRLHHRNLVKLVGYYSSRETSQNLLCYELVPNGSLEEWLHGPLGANSPLLWDTRMKIALDAARGLAY
LHEDSQPSVIHRDFKASNILLENNFLAKVADFGLAKQAPEGRVNYLSTRVMGTFGYVAPEYAMTGHLLVKSDVYSYGVVLLELLTGRKPVDMSQPSGQEN
LVTWTRPILRDKDRLEELADPRLVGKYPKEDFVRVCTIAAACVAPEASQRPTMGEVVQSLKMVQRVTEYQDAAAAAAAASSARPNTRQSCTTFDSDGTSS
MFSSGPFSGLSNFDNERTAAFSEDLHEGR
|
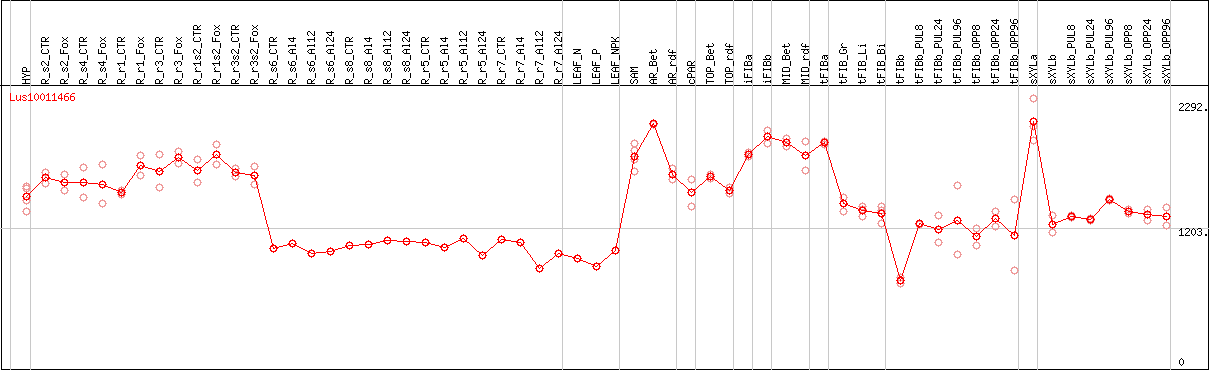
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10011466 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
