Lus10011548 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10011548 pacid=23140404 polypeptide=Lus10011548 locus=Lus10011548.g ID=Lus10011548.BGIv1.0 annot-version=v1.0
ATGGGTTCTGCAAAAACCGCTCGGAGCTCCGTTTGCATCTTCTGGGTTCTCTTGCTTTGCTCTTTCCCGTTCTTCGCGTCCTTCGCTTTCGCCACTGCAT
TAGATATTAGTGAAAGCTCTAGACTCAATGTGACTCCTAGCTTGCCTGTGGTGAACTCCCCTGGGACGAAGCCTGGGACTAGGCTGCTTTGTGAAAGAGT
TCGTATCGATGGGTTACATAGGCTTAAGAACTTGAAGAAGTTTTCACACTCGTTGAAGCTGAACCTTACGCATACTAGCTCCACTCTCCGTGGCCCAAGT
GTTGAAGTTTGTTTCCACAGGAATTCATCCGTTGGAGTTGGGATGTGTTCGAAAGGAAAGTGGGAGAAAGTTGTTAAGGGCAAATGGGGTCGTGCTATGT
CACCTTTTGATCATAAGCTCCTGGATATAAGGATGTCTGGTTCATCCCCTGAAAACTTTGAACTTCACATTGAAGAAGAATTTTTCTTGTATCGGATCGT
ATTCCTAGTAGCTGGGATCTTTATGTTGAGCATGGCATCCCCTTTGAGCGAGTCGTTAACGTTTTACTACTGCAGTGGCATGGCACTTGGCATCATCCTT
GTAATATTGGTGGTGCTTTTTCAGGGGATGAAAATTTTGCCAACTGGTCGGAGGAACTCACTTGCGATTGTCATATACGCATCTTTGGTTGGTTTGGGAT
CTTTTCTCCTGCAATACGTGCCAAACTTATTGCAGTCACTCCTCCTTGAGATGGGAATCAGCGAAGACCTGTATTTTCCTGTATTTTTTCTGGTGGCTTT
TATTGTTCTCGCTGGAGCGTGGATGGGATTCTGGGCTGTGCGGAAACTTGTCTTGGCAGAAGATGGATCAGTTGACACTAGTACTTCCATTTTTGTTGCC
TGGTCTATCAGGATTGTTGCTTGTATTATGATTCTTCAGAGCTCATTGGATCCGCTGCTAGGAGCAGGGGCATTACTATCAGGAATTATTCTCTCATCGA
TGTTGAAGAGAACTTTCCGATTGGAATTTTGGCGGCGGCTCTACACGGGGTTGCTCAATTTGGTGGACAACATTAGCTCGAAGTCTGACATTCCTGATCT
ATCTCCGTTCCAGAATCCTCATGAAAAGTACACATACAGAACTCCTGAAGATTCTCAGCCTATCTGGCCCCGGCCCAAGAGATTCAGTTCTCCTATTCAA
GATAACAAAACGTCTCCAGCATCAGATTCAGGTGTGTTCCTTTCCACCTTCTGCACTCCAGAGAAGAGGACGAAATTCTCCCAGGAAGAGTGGAAAAGGA
TCACCAAAGACTCAACACAGAAGGCGGTGAAAGAACTCGTCGGTTCCCCCGATTTCAGCAGATGGGTTGCTGCAAACGCTGATCGAATCTCTGTAACTCC
AAAGTCCTCGCGGAAGGAGGCTCCTCCGTCCGTCAAAGGTAGGGACGGCCTGAGGAAAGCGAGCGACGGTGATTATTCCAATCCAGTGGAGGAAGAGGAG
CAGTTCGTGAGGTGGTTTCGAGAAGCTTGGCCTTATCTGTGGGCTCATCGTGGGAGTACTTTTGTGGTTGTTATCTCTGGTGACATTGATGCCAGCTCTT
ACTTGGACCCAATTCTTAAGGATATAGCATTGCTTCATCACCTTGGAATCAGGTTCGTGATTGTTCCAGGGACTCATGTTCAAATCGATGATCTTTTGGC
TGAGAGAGGGCATGAGCCTAAGCATGTTGGTCCATATAGAATAACTGATCCAGAAGCTCTATCTGCATCAATGGAGGCGGCTGGAAAAATTCGAATGATG
ATAGAGGCTAAGCTTTCTCCGGGACCTTCCATATGCAAGCGATTTTCTCTAAGTTCATTATGGTTTTGTTTACAGAGACGAGGAGTTGTTGGCGGTGTTG
ATTTTGGGGCAACCGGTGAAGTGAAGAAAGTAGACGTCGATCGTATGAGTGAGAGGCTTGATAGTAGTTGCATAGTTGTGCTGAATAACTTGGGGTATTC
GAGCTCCGGCGAAGTTCTGAATTGCAACACATATGAAGTTGCTACAGCTTGTGCGTTAGCAATTGGTGCAGATAAGCTGATATGCATCATGGATGGTCCC
ATTCTTGACGAGAATGGGCATCTCATTCGTTTCTTAACTCTTGAAGAAGCAGACACATTGATTCGTGAGCGGGCTAAGCAAAGTGAGGTAGCTGCTCATT
ATGTCAAGGTGGTTGCAGAAGAAGACCCCTCTTTTGTCGAAGACACCGATTCCATCTCATTAGTTGCCAAAGATAATGCGACATTTCACAATGGAGTTGG
CTTTGGCAACGGCAGTGGTCTGTGGTCAGGAGAACAAGGTTTTGCAATTGGTGGACAGGAAAGACTTAGCCGATCACAAGGATATCTTTCGGAATTGGCT
GCCGCGGCTTTTGTTTGCAAGGGTGGTGTGCAAAGAGTACATCTATTAGACGGTACAATCGGTGGGGTTTTACTGTTAGAACTCTTTAAAAGAGATGGAA
TGGGAACTATGGTGGCTAGCGACCTATACGAAGGAACCCGTATGGCTAGGGTGTCTGACCTTGCCGGGATAAAACAGATTATACAGCCTTTACAAGAAGC
AGGCGTATTGGTCAAAAGGACCGATGAAGAGCTGCTTAAAACATTGGGTTCATACATTGTGGTGGAAAGGGAAGGTCAAATCATTGCTTGCACTGCACTC
TTCCCTTTCTTTGAAGACAAGTGTGGAGAGGTTGCTGCAATTGCAGTTTCTTCGGATTGCCGTGGACAAGGCCAAGGAGACAAATTATTAGATTATATCG
AGAAGAAGGCATCTTCACTGGGATTGGAAATATTGTTCCTTCTTACTACTCGTACAGCGGACTGGTTTACAAGACGCGGGTTCTCAGAGTGTTCGATCGA
GATGATACCGGAACAGAGGAGGAAGAAGATCAATCTGTCGCGGCACTCGAAATACTATATGAAGAGATTGCTTCCTGATACAAGTGGGATTACTCTAACT
GCCCAGCAATCTAGCTGGCAATAG
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10011548 pacid=23140404 polypeptide=Lus10011548 locus=Lus10011548.g ID=Lus10011548.BGIv1.0 annot-version=v1.0
MGSAKTARSSVCIFWVLLLCSFPFFASFAFATALDISESSRLNVTPSLPVVNSPGTKPGTRLLCERVRIDGLHRLKNLKKFSHSLKLNLTHTSSTLRGPS
VEVCFHRNSSVGVGMCSKGKWEKVVKGKWGRAMSPFDHKLLDIRMSGSSPENFELHIEEEFFLYRIVFLVAGIFMLSMASPLSESLTFYYCSGMALGIIL
VILVVLFQGMKILPTGRRNSLAIVIYASLVGLGSFLLQYVPNLLQSLLLEMGISEDLYFPVFFLVAFIVLAGAWMGFWAVRKLVLAEDGSVDTSTSIFVA
WSIRIVACIMILQSSLDPLLGAGALLSGIILSSMLKRTFRLEFWRRLYTGLLNLVDNISSKSDIPDLSPFQNPHEKYTYRTPEDSQPIWPRPKRFSSPIQ
DNKTSPASDSGVFLSTFCTPEKRTKFSQEEWKRITKDSTQKAVKELVGSPDFSRWVAANADRISVTPKSSRKEAPPSVKGRDGLRKASDGDYSNPVEEEE
QFVRWFREAWPYLWAHRGSTFVVVISGDIDASSYLDPILKDIALLHHLGIRFVIVPGTHVQIDDLLAERGHEPKHVGPYRITDPEALSASMEAAGKIRMM
IEAKLSPGPSICKRFSLSSLWFCLQRRGVVGGVDFGATGEVKKVDVDRMSERLDSSCIVVLNNLGYSSSGEVLNCNTYEVATACALAIGADKLICIMDGP
ILDENGHLIRFLTLEEADTLIRERAKQSEVAAHYVKVVAEEDPSFVEDTDSISLVAKDNATFHNGVGFGNGSGLWSGEQGFAIGGQERLSRSQGYLSELA
AAAFVCKGGVQRVHLLDGTIGGVLLLELFKRDGMGTMVASDLYEGTRMARVSDLAGIKQIIQPLQEAGVLVKRTDEELLKTLGSYIVVEREGQIIACTAL
FPFFEDKCGEVAAIAVSSDCRGQGQGDKLLDYIEKKASSLGLEILFLLTTRTADWFTRRGFSECSIEMIPEQRRKKINLSRHSKYYMKRLLPDTSGITLT
AQQSSWQ
|
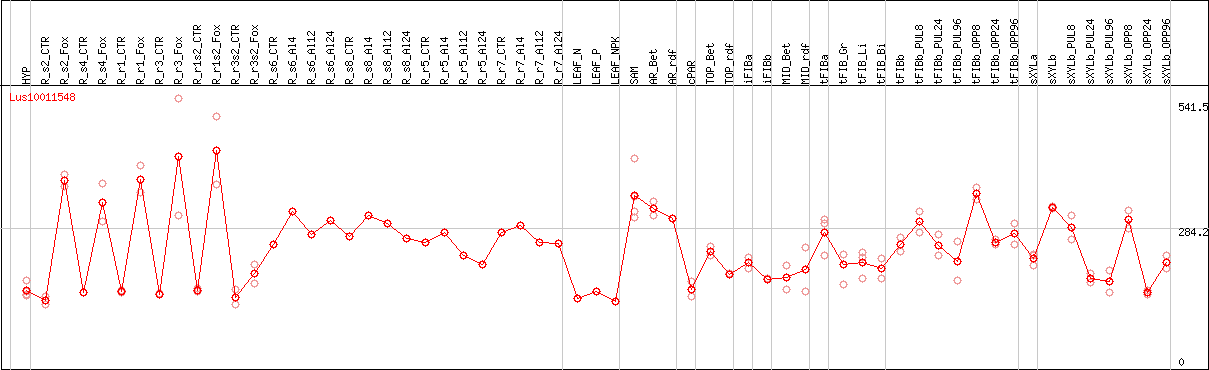
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10011548 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
