External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G49740 303 / 3e-97
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G41080 224 / 1e-68
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G16835 225 / 2e-68
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G25360 225 / 2e-67
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G09410 224 / 2e-67
pentatricopeptide (PPR) repeat-containing protein (.1)
AT4G33170 227 / 3e-67
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G49170 225 / 4e-67
EMB2261
embryo defective 2261, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G37380 221 / 6e-67
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G13600 221 / 2e-66
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT4G13650 224 / 4e-66
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10019278
573 / 0
AT3G49740 694 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10019891
235 / 2e-72
AT1G56570 666 / 0.0
PENTATRICOPEPTIDE REPEAT PROTEIN FOR GERMINATION ON NaCl, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10000597
226 / 6e-71
AT4G16835 545 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10001617
231 / 8e-71
AT2G03880 840 / 0.0
required for efficiency of mitochondrial editing 1, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10028106
230 / 2e-69
AT1G25360 916 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10010998
229 / 3e-69
AT4G16835 778 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10005064
221 / 5e-69
AT2G21090 513 / 2e-179
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Lus10007806
230 / 1e-68
AT3G03580 967 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10019098
228 / 2e-67
AT3G47840 697 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G091600
238 / 5e-73
AT4G37170 924 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.007G021700
234 / 4e-71
AT3G47840 796 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.010G136200
232 / 4e-70
AT1G68930 997 / 0.0
pentatricopeptide (PPR) repeat-containing protein (.1)
Potri.016G051300
229 / 6e-70
AT3G13770 908 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.002G155100
230 / 3e-69
AT3G61170 933 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.017G086100
224 / 9e-69
AT5G15300 675 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.007G050200
224 / 3e-68
AT4G37380 817 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G012600
224 / 9e-68
AT2G29760 540 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.014G012500
223 / 3e-67
AT3G49710 995 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.013G006700
221 / 3e-67
AT1G56570 732 / 0.0
PENTATRICOPEPTIDE REPEAT PROTEIN FOR GERMINATION ON NaCl, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10011562 pacid=23140364 polypeptide=Lus10011562 locus=Lus10011562.g ID=Lus10011562.BGIv1.0 annot-version=v1.0
ATGATTCATTCAAATGTCTTTAAAGCAGGGCTGGTTTCAAGAATCCAGGTATCAAATTCCTTGATTTCAGCATACTGCAAACATGGCACTGTCGAATCAG
GTCATAAAGTGTTTCTCACCATGTCGTCGAGAAATTTGGTGTCGTGGAACACTATAATGTCAGGTTCGTCATTGAATGGATTTCACATACAGGGGTTAGA
ACATTTCTATCAACTGCTAAAGTCTGGATTTCTCCCTAACGAGCATACCCTCACTATTGCTTTGAGCATTTGTGCAAGCATATCAGCTTCAGAACAAGCA
AAGCAGATTCACGCTCATTTCATAAGAGTCAACTCTTCTCCAAGAGGGTCTTTAGGAAACGCCCTGATTACAACTTACTCGAAATGTGTCCTTCTTAATT
CATCTTCAAAGGTCTTCAGCACCATGTCCCAGAAAGATGTCGTCTCCTGGAACGCCCTGATGTCTGCTTATGCACAACACGGTGAAGCAAACCATGCATT
AAACTGTCGTGATGAAATGTCAGATTCGTTAGATATTCGTCCTGATCAGGCTACGTTCTCTGCAGCTCTCTCTGCCTGTAGCCATGCAGGCTTGGTCGAC
GAGGGAGCTCGAGTTTTCAATTCCATGGTTGATAGGTACGGGATTAAACCCGGAGTGGATCATTTTTCGTGCGTTATTGACCTTCTGGGACGTGCTGGGT
ATTTTACTCGGGTGGATGAGATAATCCAAAGGGAGGATTTTGAAGCTCATCCTAACATCTGGTGGACGTTGATCAGTGCTTGCGCTACTCTTGGGAACTT
GGAACTGGGGAGGAGGGTTGCTGGGTATCTTCTCGAAACTGAACGACACAATTCGTCGGCTTATGTGCTGTTAGCTAATATATATGCAGGTGCTGGTCAG
TGGGAAGAAGCAGCTAATGTGAGGCGAGAAATGGATACGTTTGGGACGATGAAGCAACCTGGACATAGCTGGATTTGCAGGTAA
AA sequence
>Lus10011562 pacid=23140364 polypeptide=Lus10011562 locus=Lus10011562.g ID=Lus10011562.BGIv1.0 annot-version=v1.0
MIHSNVFKAGLVSRIQVSNSLISAYCKHGTVESGHKVFLTMSSRNLVSWNTIMSGSSLNGFHIQGLEHFYQLLKSGFLPNEHTLTIALSICASISASEQA
KQIHAHFIRVNSSPRGSLGNALITTYSKCVLLNSSSKVFSTMSQKDVVSWNALMSAYAQHGEANHALNCRDEMSDSLDIRPDQATFSAALSACSHAGLVD
EGARVFNSMVDRYGIKPGVDHFSCVIDLLGRAGYFTRVDEIIQREDFEAHPNIWWTLISACATLGNLELGRRVAGYLLETERHNSSAYVLLANIYAGAGQ
WEEAANVRREMDTFGTMKQPGHSWICR
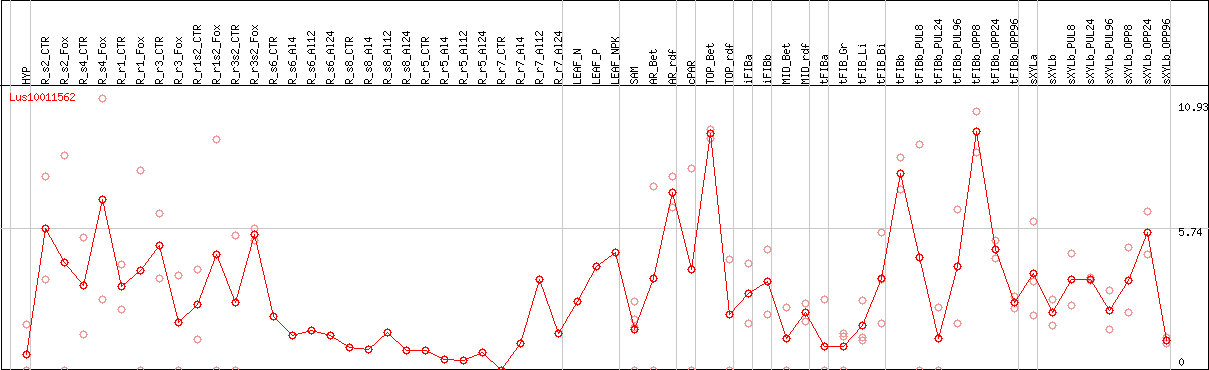
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10011562 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.