External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G65700 180 / 2e-51
BAM1
BARELY ANY MERISTEM 1, Leucine-rich receptor-like protein kinase family protein (.1.2)
AT3G49670 170 / 1e-47
BAM2
BARELY ANY MERISTEM 2, Leucine-rich receptor-like protein kinase family protein (.1)
AT1G75820 108 / 2e-26
ATCLV1, FLO5, FAS3, CLV1
FLOWER DEVELOPMENT 5, FASCIATA 3, CLAVATA 1, Leucine-rich receptor-like protein kinase family protein (.1)
AT5G62230 100 / 2e-23
ERL1
ERECTA-like 1 (.1.2)
AT5G46330 97 / 3e-22
FLS2
FLAGELLIN-SENSITIVE 2, Leucine-rich receptor-like protein kinase family protein (.1)
AT1G73080 96 / 6e-22
ATPEPR1, PEPR1
PEP1 receptor 1 (.1)
AT3G47570 95 / 7e-22
Leucine-rich repeat protein kinase family protein (.1)
AT3G57830 92 / 5e-21
Leucine-rich repeat protein kinase family protein (.1)
AT5G24100 92 / 1e-20
Leucine-rich repeat protein kinase family protein (.1)
AT4G37250 91 / 2e-20
Leucine-rich repeat protein kinase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10019248
295 / 9e-93
AT5G65700 1479 / 0.0
BARELY ANY MERISTEM 1, Leucine-rich receptor-like protein kinase family protein (.1.2)
Lus10011585
239 / 2e-72
AT5G65700 1044 / 0.0
BARELY ANY MERISTEM 1, Leucine-rich receptor-like protein kinase family protein (.1.2)
Lus10039641
236 / 3e-71
AT5G65700 1580 / 0.0
BARELY ANY MERISTEM 1, Leucine-rich receptor-like protein kinase family protein (.1.2)
Lus10017267
113 / 6e-28
AT1G75820 1305 / 0.0
FLOWER DEVELOPMENT 5, FASCIATA 3, CLAVATA 1, Leucine-rich receptor-like protein kinase family protein (.1)
Lus10013562
112 / 2e-27
AT1G75820 1297 / 0.0
FLOWER DEVELOPMENT 5, FASCIATA 3, CLAVATA 1, Leucine-rich receptor-like protein kinase family protein (.1)
Lus10040592
108 / 1e-26
AT1G75820 642 / 0.0
FLOWER DEVELOPMENT 5, FASCIATA 3, CLAVATA 1, Leucine-rich receptor-like protein kinase family protein (.1)
Lus10035429
105 / 2e-25
AT3G47570 771 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
Lus10033362
100 / 1e-23
AT3G47570 784 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
Lus10037310
99 / 3e-23
AT3G47570 744 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G009200
219 / 3e-65
AT5G65700 1530 / 0.0
BARELY ANY MERISTEM 1, Leucine-rich receptor-like protein kinase family protein (.1.2)
Potri.001G073600
115 / 9e-29
AT4G20270 1279 / 0.0
BARELY ANY MERISTEM 3, Leucine-rich receptor-like protein kinase family protein (.1)
Potri.003G157300
114 / 2e-28
AT4G20270 1285 / 0.0
BARELY ANY MERISTEM 3, Leucine-rich receptor-like protein kinase family protein (.1)
Potri.005G037400
110 / 6e-27
AT3G47570 622 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
Potri.005G241500
108 / 2e-26
AT1G75820 1268 / 0.0
FLOWER DEVELOPMENT 5, FASCIATA 3, CLAVATA 1, Leucine-rich receptor-like protein kinase family protein (.1)
Potri.002G019900
107 / 7e-26
AT1G75820 1226 / 0.0
FLOWER DEVELOPMENT 5, FASCIATA 3, CLAVATA 1, Leucine-rich receptor-like protein kinase family protein (.1)
Potri.018G045500
106 / 8e-26
AT2G25790 925 / 0.0
Leucine-rich receptor-like protein kinase family protein (.1)
Potri.004G060100
104 / 5e-25
AT3G47570 670 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
Potri.008G034000
103 / 6e-25
AT3G47570 827 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
Potri.015G037400
103 / 1e-24
AT3G47570 786 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0022
LRR
PF00560
LRR_1
Leucine Rich Repeat
CL0022
PF08263
LRRNT_2
Leucine rich repeat N-terminal domain
Representative CDS sequence
>Lus10011577 pacid=23140427 polypeptide=Lus10011577 locus=Lus10011577.g ID=Lus10011577.BGIv1.0 annot-version=v1.0
ATGAGACTCCTTTACCCTCTCCTCTTCTTCTTCCTCCTCATCCACCTCCACATCCCAACTTCCACCGGCCGCGTATTATCCGAAACCCAAGCTCTACTCT
CCCTAAAATCCGCCGTCACCGACGACCCAAACTCCTTCCTTTCCGCCTGGAACTCCTCCGCCGACCTCTGCTCCTGGCCGGGAGTCACGTGCGACTTCTC
CAGCCGTCACGTGACCTCATTAGACCTCTCTAGCCTCTCCCTCACGGGAACCCTCTCCCCGGATATCTCCCACCTCCGCTACCTCCAGAACCTCTCCCTC
GACGTCAACAAATTCTCTGGCCCCATCCCGCCGCAGCTCTCCGCCGCCACCGACCTCCGCTACCTCAATCTCTCCAACAATGGGTTTAATGATACTTTCC
CCTCGCAGCTAGCCCAGCTCAAGAACCTCGAGGTACTCGACATCTACAACAACAATATGACCGGTGACTTGCCTCTCGCCGTCGCTGAAATGCCCAACCT
CCGCCACCTCCACCTCGGAGGGAACTTCTTCGGCGGGGAGATCCCCCCGCCGGACGGGCAGTGTGAGGTGTTCTTCTCAATCTACGATGAGACTCCTTTA
CCCTCTCCTCTTCTTCTTCCTCCTCATCCACCTCCACATCCCAACTTCCACCGGCCGCGTATTATCCGAAACCCAAGCTCTACTCTCCCTAAAATCCGCC
GTCACCGACGACCCAAACTCCTTCCTTTCCGCCTGGAACTCCTCCGCCGACCTCTGCTCCTGGCCGGGAGTCACGTGCGACTTCTCCAGCCGTCACGTGA
CCTCATTAGACCTCTCTAG
AA sequence
>Lus10011577 pacid=23140427 polypeptide=Lus10011577 locus=Lus10011577.g ID=Lus10011577.BGIv1.0 annot-version=v1.0
MRLLYPLLFFFLLIHLHIPTSTGRVLSETQALLSLKSAVTDDPNSFLSAWNSSADLCSWPGVTCDFSSRHVTSLDLSSLSLTGTLSPDISHLRYLQNLSL
DVNKFSGPIPPQLSAATDLRYLNLSNNGFNDTFPSQLAQLKNLEVLDIYNNNMTGDLPLAVAEMPNLRHLHLGGNFFGGEIPPPDGQCEVFFSIYDETPL
PSPLLLPPHPPPHPNFHRPRIIRNPSSTLPKIRRHRRPKLLPFRLELLRRPLLLAGSHVRLLQPSRDLIRPL
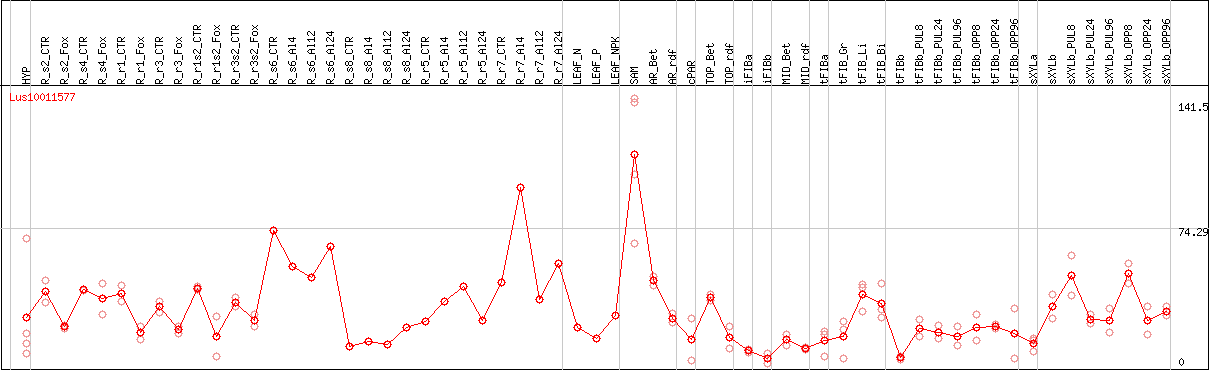
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10011577 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.