External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G26650 117 / 8e-30
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT5G55550 113 / 1e-28
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3), RNA-binding (RRM/RBD/RNP motifs) family protein (.4)
AT4G14300 112 / 3e-28
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT5G47620 110 / 1e-27
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3), RNA-binding (RRM/RBD/RNP motifs) family protein (.4)
AT2G33410 107 / 2e-26
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT3G07810 103 / 8e-25
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT1G58470 100 / 2e-24
XF41, ATRBP1
RNA-binding protein 1 (.1)
AT5G40490 96 / 2e-22
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT3G13224 92 / 3e-21
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT1G17640 92 / 5e-21
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10019276
154 / 8e-45
AT4G14300 149 / 1e-41
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10018657
116 / 5e-30
AT1G58470 233 / 1e-73
RNA-binding protein 1 (.1)
Lus10038806
116 / 2e-29
AT3G07810 417 / 2e-141
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10007721
115 / 4e-29
AT1G58470 222 / 6e-68
RNA-binding protein 1 (.1)
Lus10000537
115 / 4e-29
AT3G07810 420 / 8e-143
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10039054
115 / 4e-29
AT3G07810 428 / 4e-147
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10017555
114 / 2e-28
AT3G07810 423 / 7e-144
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10039053
113 / 2e-28
AT3G07810 417 / 1e-142
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10022889
112 / 2e-28
AT4G14300 315 / 9e-105
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G258600
121 / 1e-31
AT4G14300 277 / 1e-89
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.014G195300
118 / 1e-30
AT4G14300 305 / 2e-100
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.002G115000
114 / 2e-29
AT1G58470 219 / 7e-69
RNA-binding protein 1 (.1)
Potri.006G015700
114 / 7e-29
AT3G07810 447 / 8e-154
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.016G010200
113 / 2e-28
AT3G07810 413 / 2e-140
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.001G361300
111 / 7e-28
AT3G07810 433 / 1e-148
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.002G222400
111 / 9e-28
AT3G07810 439 / 8e-151
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.011G088100
108 / 8e-27
AT3G07810 428 / 1e-146
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.010G067500
108 / 1e-26
AT4G14300 302 / 7e-99
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.014G161600
107 / 3e-26
AT3G07810 498 / 3e-174
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0221
RRM
PF00076
RRM_1
RNA recognition motif. (a.k.a. RRM, RBD, or RNP domain)
Representative CDS sequence
>Lus10011728 pacid=23145614 polypeptide=Lus10011728 locus=Lus10011728.g ID=Lus10011728.BGIv1.0 annot-version=v1.0
ATGGTTTCTTCTCCTGCTAACTTGATGGCCATGGAAGGAGGAAAAAAACAACCGCGACCAAAACTGTTCGTGGGAGGACTTAACAATACAATAACAGGAC
TTGTCCTACGGCAGCACTTCCAGGAGTATGGCCTAGTAAAAGATGCTTTTGTCGTTCAAGACAAGCAAATTAGACTGAGCCGAGGGTTTGGTTACGTGGA
GTTTGAAGATCCATCAGCAGCTCATCATTCTCGCCAGCGCCAACACGTCATTCTTGAGAAAAAAGCTCAACCAAAGAAAAAGGGTAACCAGAACATCAAT
ATCAATGGGGAGATGAGTGCGAGGGGAAAGAAGATCTTCGTCGGAGGCTTGCTGTCAACAACTACTGAATCGGAGTTTAGGAGTTTCTTTGAGAGATTTG
GAGCAGTTAATGATGCGATTCTGGTTCATGATAAACAAAGCGGTAGACCAAGAGGGTTTGGCTTCATCACTTTTGAACCAGAGGATTCTGCAGATGGAGC
TCTTGGAAAGAGATTTCACCAACTGAATGGCTCGGATGTAGAGGTGAAGAGGGCTAATCCTAAGGATAAAAGGACTACTACCGATCTCAGATACTACTAT
GAATCAGTTCTAAATTATCATTACGTTTATGATTATTATCCTCATGGTTATTACTGGTATCACCATGCTCTGCCATTAAACTATCCGTATTATCTATTGT
CTTCAAACTATGCAACTGACTTGTCCTACCAAAGCTATACTAAACTCCACACCGCCGGTGGGACAGACACCATGTGGCATCAACAAACAACAGACGGGGA
GAATACTGCGGAAAAAAGCGGTAACTTGGTCTACTGA
AA sequence
>Lus10011728 pacid=23145614 polypeptide=Lus10011728 locus=Lus10011728.g ID=Lus10011728.BGIv1.0 annot-version=v1.0
MVSSPANLMAMEGGKKQPRPKLFVGGLNNTITGLVLRQHFQEYGLVKDAFVVQDKQIRLSRGFGYVEFEDPSAAHHSRQRQHVILEKKAQPKKKGNQNIN
INGEMSARGKKIFVGGLLSTTTESEFRSFFERFGAVNDAILVHDKQSGRPRGFGFITFEPEDSADGALGKRFHQLNGSDVEVKRANPKDKRTTTDLRYYY
ESVLNYHYVYDYYPHGYYWYHHALPLNYPYYLLSSNYATDLSYQSYTKLHTAGGTDTMWHQQTTDGENTAEKSGNLVY
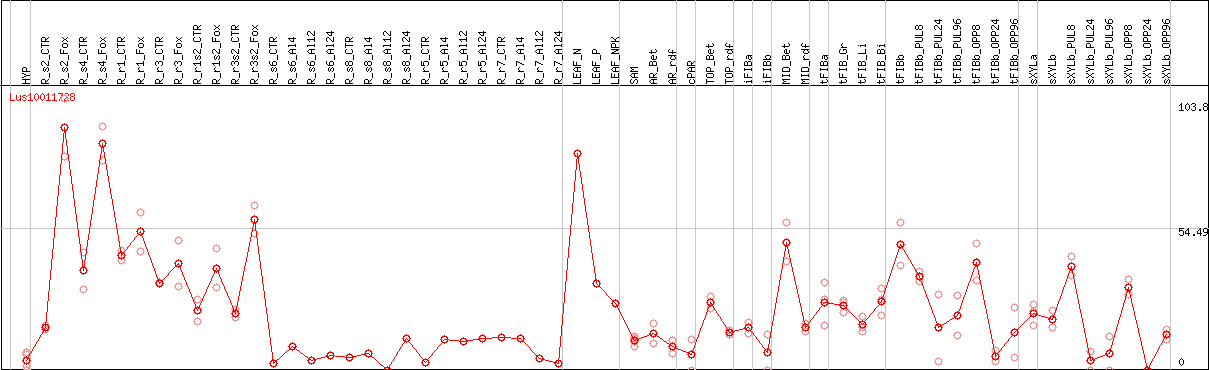
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10011728 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.