External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G13930 389 / 6e-137
ATCHS, TT4, CHS
TRANSPARENT TESTA 4, CHALCONE SYNTHASE, Chalcone and stilbene synthase family protein (.1)
AT1G02050 154 / 5e-45
LAP6
LESS ADHESIVE POLLEN 6, Chalcone and stilbene synthase family protein (.1)
AT4G00040 153 / 1e-44
Chalcone and stilbene synthase family protein (.1)
AT4G34850 152 / 2e-44
LAP5
LESS ADHESIVE POLLEN 5, Chalcone and stilbene synthase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10023670
454 / 1e-161
AT5G13930 693 / 0.0
TRANSPARENT TESTA 4, CHALCONE SYNTHASE, Chalcone and stilbene synthase family protein (.1)
Lus10031622
324 / 3e-111
AT5G13930 526 / 0.0
TRANSPARENT TESTA 4, CHALCONE SYNTHASE, Chalcone and stilbene synthase family protein (.1)
Lus10033717
320 / 2e-109
AT5G13930 518 / 0.0
TRANSPARENT TESTA 4, CHALCONE SYNTHASE, Chalcone and stilbene synthase family protein (.1)
Lus10011746
304 / 2e-103
AT5G13930 524 / 0.0
TRANSPARENT TESTA 4, CHALCONE SYNTHASE, Chalcone and stilbene synthase family protein (.1)
Lus10012586
222 / 1e-72
AT5G13930 230 / 1e-73
TRANSPARENT TESTA 4, CHALCONE SYNTHASE, Chalcone and stilbene synthase family protein (.1)
Lus10026286
162 / 3e-48
AT4G34850 650 / 0.0
LESS ADHESIVE POLLEN 5, Chalcone and stilbene synthase family protein (.1)
Lus10002187
162 / 4e-48
AT1G02050 576 / 0.0
LESS ADHESIVE POLLEN 6, Chalcone and stilbene synthase family protein (.1)
Lus10039904
161 / 2e-47
AT1G02050 577 / 0.0
LESS ADHESIVE POLLEN 6, Chalcone and stilbene synthase family protein (.1)
Lus10042388
157 / 3e-46
AT4G34850 636 / 0.0
LESS ADHESIVE POLLEN 5, Chalcone and stilbene synthase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G051500
418 / 2e-148
AT5G13930 692 / 0.0
TRANSPARENT TESTA 4, CHALCONE SYNTHASE, Chalcone and stilbene synthase family protein (.1)
Potri.001G051600
417 / 4e-148
AT5G13930 692 / 0.0
TRANSPARENT TESTA 4, CHALCONE SYNTHASE, Chalcone and stilbene synthase family protein (.1)
Potri.003G176900
414 / 7e-147
AT5G13930 694 / 0.0
TRANSPARENT TESTA 4, CHALCONE SYNTHASE, Chalcone and stilbene synthase family protein (.1)
Potri.003G176800
412 / 5e-146
AT5G13930 693 / 0.0
TRANSPARENT TESTA 4, CHALCONE SYNTHASE, Chalcone and stilbene synthase family protein (.1)
Potri.014G145100
402 / 4e-142
AT5G13930 679 / 0.0
TRANSPARENT TESTA 4, CHALCONE SYNTHASE, Chalcone and stilbene synthase family protein (.1)
Potri.003G176700
401 / 1e-141
AT5G13930 684 / 0.0
TRANSPARENT TESTA 4, CHALCONE SYNTHASE, Chalcone and stilbene synthase family protein (.1)
Potri.012G138800
336 / 4e-116
AT5G13930 531 / 0.0
TRANSPARENT TESTA 4, CHALCONE SYNTHASE, Chalcone and stilbene synthase family protein (.1)
Potri.001G028600
309 / 2e-105
AT5G13930 510 / 0.0
TRANSPARENT TESTA 4, CHALCONE SYNTHASE, Chalcone and stilbene synthase family protein (.1)
Potri.005G175200
191 / 5e-60
AT5G13930 425 / 2e-149
TRANSPARENT TESTA 4, CHALCONE SYNTHASE, Chalcone and stilbene synthase family protein (.1)
Potri.009G128800
155 / 2e-45
AT4G34850 631 / 0.0
LESS ADHESIVE POLLEN 5, Chalcone and stilbene synthase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0046
Thiolase
PF00195
Chal_sti_synt_N
Chalcone and stilbene synthases, N-terminal domain
Representative CDS sequence
>Lus10011745 pacid=23165932 polypeptide=Lus10011745 locus=Lus10011745.g ID=Lus10011745.BGIv1.0 annot-version=v1.0
ATGTCGACTATCACCGTGGATGAAGTACGCAAGGCCCAACGTGCACAAGGTCCCGCCACCGTCCTCGCCATCGGAACCGCCACTCCTGCCAACTGCTTCG
ATCAAAGCACCTACCCTGATTACTACTTCCGTATTACCAACAGCGAGCACAAGACTGAGCTCAAGGAGAAGTTCAAGCGCATGTGCGAGAAGTCGATGAT
CAAGAAACGTTACATGCACTTGACGGAAGAGTTCCTCAAGGATAATCCGACCATGTGTGCCTACATGGCAAATTCCCTCGACGCTCGCCAAGACGTCGTC
GTCCCTGAAGTTCCAAAGCTAGGAAAAGAAGCTGCCACAAAAGCCATCAAGGAATGGGGACAGCCCAAATCCAAAATCACCCACCTCATCTTCTGCACCA
CCAGTGGGGTCGACATGCCAGGCGCCGACTACCAGCTCACCAAGCTGTTGGCCCTCCGCCCGTCGGTGAAGCGCTTCATGATGTACCAACAAGGCTGCTT
CGCCGGTGGCACGGTACTCCGCCTCGCCAAGGACCTCGCCGAGAACAACAAGGGTGCACGCGTCCTCGTCGTGTGTTCCGAAATAACCGCCGTCACCTTC
CGTGGGCCGAGCGATGCCCATCTAGACAGTCTCGTGGGCCAGGCCTTGTTCGGCGACTGA
AA sequence
>Lus10011745 pacid=23165932 polypeptide=Lus10011745 locus=Lus10011745.g ID=Lus10011745.BGIv1.0 annot-version=v1.0
MSTITVDEVRKAQRAQGPATVLAIGTATPANCFDQSTYPDYYFRITNSEHKTELKEKFKRMCEKSMIKKRYMHLTEEFLKDNPTMCAYMANSLDARQDVV
VPEVPKLGKEAATKAIKEWGQPKSKITHLIFCTTSGVDMPGADYQLTKLLALRPSVKRFMMYQQGCFAGGTVLRLAKDLAENNKGARVLVVCSEITAVTF
RGPSDAHLDSLVGQALFGD
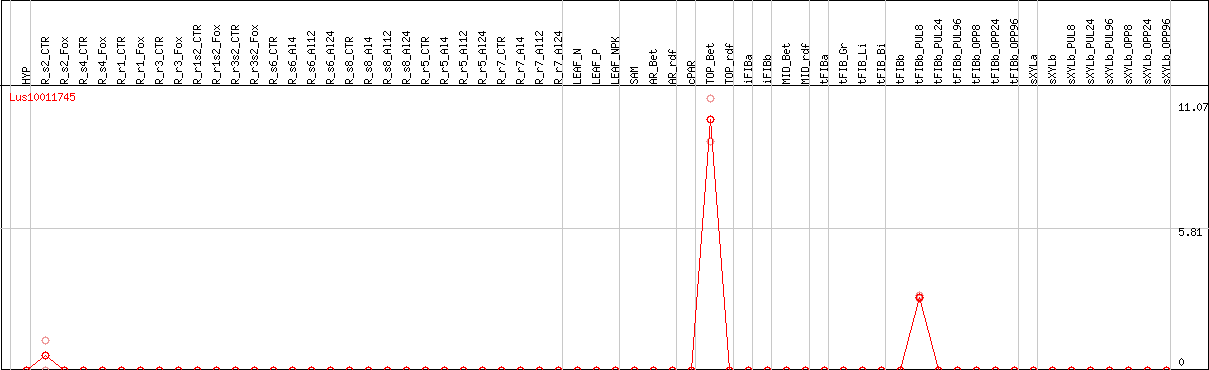
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10011745 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.