External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G33270 255 / 4e-83
AtCDC20.1, CDC20.1
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
AT4G33260 246 / 2e-79
AtCDC20.2, CDC20.2
cell division cycle 20.2, Transducin family protein / WD-40 repeat family protein (.1.2)
AT5G27080 230 / 3e-73
AtCDC20.3
cell division cycle 20.3, Transducin family protein / WD-40 repeat family protein (.1)
AT5G27570 228 / 7e-73
AtCDC20.5
cell division cycle 20.5, Transducin/WD40 repeat-like superfamily protein (.1)
AT5G27945 223 / 1e-70
Transducin/WD40 repeat-like superfamily protein (.1)
AT5G26900 221 / 8e-70
AtCDC20.4
cell division cycle 20.4, Transducin family protein / WD-40 repeat family protein (.1)
AT5G13840 156 / 1e-44
FZR3
FIZZY-related 3 (.1.2)
AT4G11920 153 / 1e-43
FZR1, CCS52A2
FIZZY-RELATED 1, cell cycle switch protein 52 A2 (.1)
AT4G22910 153 / 1e-43
CCS52A1, FZR2
cell cycle switch protein 52 A1, FIZZY-related 2 (.1)
AT2G21390 58 / 2e-09
Coatomer, alpha subunit (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10034687
286 / 3e-95
AT4G33270 620 / 0.0
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Lus10037595
274 / 1e-93
AT4G33270 402 / 2e-141
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Lus10037593
276 / 3e-91
AT4G33270 728 / 0.0
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Lus10006851
273 / 5e-90
AT4G33270 728 / 0.0
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Lus10006853
279 / 2e-88
AT4G33270 729 / 0.0
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Lus10042765
263 / 1e-86
AT4G33270 542 / 0.0
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Lus10042748
258 / 3e-80
AT4G33270 592 / 0.0
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Lus10025838
199 / 3e-64
AT4G33270 295 / 3e-99
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Lus10029716
189 / 3e-61
AT4G33270 193 / 1e-60
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.016G118400
255 / 4e-83
AT4G33270 771 / 0.0
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Potri.013G048900
242 / 6e-78
AT4G33270 766 / 0.0
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Potri.019G021800
242 / 7e-78
AT4G33270 767 / 0.0
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Potri.016G068700
227 / 3e-72
AT4G33270 607 / 0.0
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Potri.015G110300
156 / 5e-45
AT4G33270 363 / 5e-122
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Potri.008G057500
156 / 1e-44
AT5G13840 705 / 0.0
FIZZY-related 3 (.1.2)
Potri.001G112700
152 / 5e-43
AT4G22910 705 / 0.0
cell cycle switch protein 52 A1, FIZZY-related 2 (.1)
Potri.010G202100
150 / 2e-42
AT5G13840 707 / 0.0
FIZZY-related 3 (.1.2)
Potri.003G119500
150 / 3e-42
AT4G22910 702 / 0.0
cell cycle switch protein 52 A1, FIZZY-related 2 (.1)
Potri.005G148500
56 / 8e-09
AT3G49660 503 / 0.0
human WDR5 \(WD40 repeat\) homolog a, Transducin/WD40 repeat-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0186
Beta_propeller
PF00400
WD40
WD domain, G-beta repeat
Representative CDS sequence
>Lus10011833 pacid=23169270 polypeptide=Lus10011833 locus=Lus10011833.g ID=Lus10011833.BGIv1.0 annot-version=v1.0
ATGGATGATTTTTACTTCAACATACTGGATTGGGGCATCAATAATGTTCTTGGATTAGCACTAGATTGCACAGTTCAACTGTGGGATGCCTCCTATGGCT
CTCCTTCCGAGCTTGTCACTGTCGATGAAGAGCTCAACCCTGTTACGAGCGTTAGCTGGGCTCCGGATGGCTGCCAGATCGCCGTCGGGTTGAACAACTC
CGAAGTCCAATTGTGGGACTATACTTCTAACAAACTGGCAGCCACAGATGCCGAGCCTTCCAACCCCCGACCGACTCAGTATCTTCACAGGCTCGAGGAC
CATACCTCAGCAGTAAAGGCTCTAGCTTGGTGTCCATTCCAGCGAAACTTGGTCGCTACAGGAGGCGGCGGAGAAGACAAGACAATCAAGTTCTGGAACA
GCCAGACCGGTGCTTGCTTGAACTCGGTGGACACGGGCTCTCAGGTCTGTGCATTGCTGCGGAACAAGCACGAGAAGGAACTTTTATGCTCTCATGGGTT
CCCCGGAAATCAGCTTACCTTGTGGAAGTATCCATCCATGGTGAAGATTGCAGAGCTTACTGGCCGTACTGCCAGGGTCCTGTTCATGGCTCAGAGTCCA
GATGGATGCACGGTGGCTTCCGCAGCAGGGGATGAAAATCTGATGGTCTGGTACGTGTTTGGGGATCCTGCTACGAAGAAATCTGCTAGGAAAGCGAATC
CGGAGCCCTTCTCTTGGACGAATCGATGCACCATCCGGTGA
AA sequence
>Lus10011833 pacid=23169270 polypeptide=Lus10011833 locus=Lus10011833.g ID=Lus10011833.BGIv1.0 annot-version=v1.0
MDDFYFNILDWGINNVLGLALDCTVQLWDASYGSPSELVTVDEELNPVTSVSWAPDGCQIAVGLNNSEVQLWDYTSNKLAATDAEPSNPRPTQYLHRLED
HTSAVKALAWCPFQRNLVATGGGGEDKTIKFWNSQTGACLNSVDTGSQVCALLRNKHEKELLCSHGFPGNQLTLWKYPSMVKIAELTGRTARVLFMAQSP
DGCTVASAAGDENLMVWYVFGDPATKKSARKANPEPFSWTNRCTIR
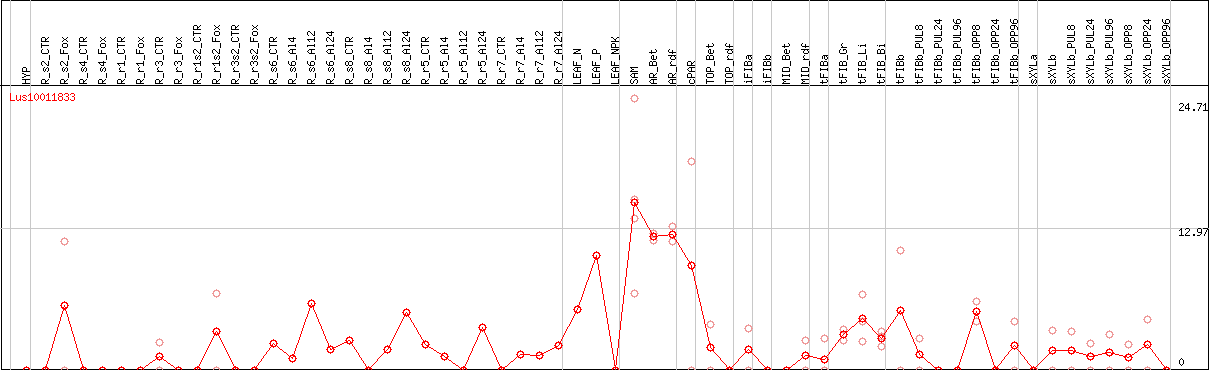
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10011833 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.