Lus10011933 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10011933 pacid=23146536 polypeptide=Lus10011933 locus=Lus10011933.g ID=Lus10011933.BGIv1.0 annot-version=v1.0
ATGGGGACTGCCAGCCTGTTGTTGCTGGTGGAGCTCTGTCTCATTGCCTCTGTTTTCAACCTTGACGGATCTACAGGATTCAGTACATCACCATCTCCTG
GAAGCATAAGTCCGGTCTATCCAGTTAGTCCTTTCGAACGTCATGCACCGTGCTCAGAGGCCTTCCTTGTTGATGGGCTTATGATAGACTCTCCAGTTGA
ATTAGCTCCTGCTCCAGCACATGAAGCAATTTCAGGGCAGGTACCATCCAACCATCCAATTCCAATTGGTTCACCGCCTTCTAGTACTATTCGTCCACCT
TCAATTATTCCAGGAGATGCACCCTCTGAATCGCCAAGCCTTTCAGTTGTACGTAGATCACATAGTGGAGCTTCTCCGCCCCAAAGTTTTGAACGAGATG
TAGCACCTATGCCATCAGATGTTTCATTCCTACCTCCGCTCCCTAATATCTCTCCTCCACTAGAGACTCGAGAAGGAAATGTAGCACCTATGCCATCTAA
CACTTCTGGTGGAAGTATTTCCTATGCACCTCCATTGCTTAACATGGCTCCTCCACCCCAGAATGTATCACCTATGCCATCTACTAACTCTTCATTGCAT
GGTATAGCTCCTCCACCCCCGATTTCTCCAGGACAAGTAGCACCTATGCCTTCAGGAGCAATCGGTTCAATTGTTCCTCCAGCACATAGTGTAGTTCCTC
CTCCCCCAATTTCTCAAGGACATGTACCACCTATGCCTTCTGGAGAAAGTTCTTCATTGGTACCTCCAATGGCAAGTGTGGCTCCTCCACCTCAGATTTC
TGAAGTGCCTCCTGCACGTAGTGTAGCTCCTCCATCCCCAGTCTTTCAAGGACAAGTAGCACCTACGCCTTCAAAAACCTCTGGAGAAAATGGTCCATTA
GTACCTCCACCAAATAGTGTAGCTCCTCCACCCCAGAATTCCGAGGGAAAACTTGCACCTCCGCCATCAAGCGATAGCATTAGCATAGCCCCTCCACCAC
ATATCATTCAAGGAACAGCAGCACCTATGCCATCAAATGATACCGGAGGAAAGGAACCTGGTGAAAAGACTCCTGCCTCCATGCCAGCTTCAGCACCTCC
AACTAAAGCATTACCAAGTATACCCTTCATCCACCCTGGTGTACCGCAACATTCGACTCTGAGTGCTCATCAGAAGAATATGTCGTTTTATGGTTCTCCT
CTTGAAAAACCAGAACCAGTTGCATTACCTCCAAGGACATTGCACAGGAAGAAACCAGCTATCCGTACACTTGCACCAGAAGCCATTATCCCATCTCTCC
CGCCTGATCCTGATGTCTCATCTCCATCATCTACTCCTGGAATGACCGATTGGAAAAGATATGGAATTCCAGTTACTGCACCCTCAACTCAAATACACAA
GCCTTTGCCACCTAGAATCCATTCACGCACTAAAGCTTCTTTTCCCGCTGTTGCTCCTTCTGAGCATCGGACAGTGAGGCATTCAAATGAACCTCTTGCT
CCATCTGTGCCTTTCCATAGTTCTCCTCGGTTCATAACGCATCAGTCTCCTTCGAATTCACCATCTACCTCTTCTCAAAAGAAACACAGTACAAGAGGTG
GAATGAGTAATGGTACTCCTGCATCATCATATGTTATACCTCCTCGCTCATCAGAAGAGCCAGCACTCCCTCCATCTCTTCCAACTACCAGGAGAACCCA
CTTTGCTTCTCCAACCAGCAATCCAGGTTTCTGTATCGCAGGTCCGGCTTCTCCTTCCCATCATTCCATCCCTAAACCAGATTTAGGGGTTTCACCAGTT
CCCTGGCCATCCAGTATAGCACCTAACTGGACCACAATACCGATTCTTCCACCAAAAGATTCTCCGAGTTCTCCATTGAAAAGTCCTATGCCCAAACTTC
CAACAATCCATGCACTGCCACCTCCACCTCCCAATCAAGATTGTTCGACGATTGTCTGCATGGAACCATATGCTAATACTCCACCAGGCTCTCCATGTGG
ATGTGTATTGCCCATGCAAGTTGGACTACGCCTTAGTGTTTCTCTCTACACCTTTTTCCCGTTGGTTTCTGAATTGGCTCAAGAGATCGCTGCCGGTGTG
TTTATGAAACAAAGTCAAGTCCGGATTATGGGAGCCAATGCCGCTAGCCAGGAACCTGAGAAGACCATTGTCCTCATTGACTTGGTTCCACTTGGACAAA
AGTTTGACAATACAACTGCTTTATTGACATATCAAAGATTCTGGCAGAAACAAGTAGTGATACAGCTTTCTTTCTTTGGTGACTATGAAGTACTGTACGT
GCGGTATCCAGGTTTACCCTCATCGCCACCTTCATCAATTTCCGGAATTACTATCATTGATGATGGGCCATATTCTGGCAATGGAAACAATGCAAGAACG
ATAAAGCCCCTAGGAATTGATGTATCAAAAAGACAACAAAAAGATGGGCTTAGTGGTAGCATGATTGCTATCATCGCCCTTTCAGCTTCCATCGTTGTAA
TCTTGAGCTCTGCAGCTGCATGGATATTTTTTTTGAGGAATCGAGGCCATGGCAGGGACCCAACTCCGACATTGGCACCTATACATGCTTCTGTTGCAAA
ACCCTTAGGGACTATTGCATCTGCTCTTAGAAGTGGTCTGAGTTCTAACTCACTGTCATTTGGCTCTAGCATTGCTCCATATACAGGATCTGCCAAGACG
TTCAGTGCAAGTGATATTGAGAGGGCGACAAATAATTTTGATTTGTCAAGACTGCTTGGGGAAGGTGGATTTGGGCGTGTTTATAGTGGTGTTCTTGAAG
ATGGGACCAAAATCGCAGTAAAAGTCCTGAAAAGAGATGACCAACAAGGTGGTCGAGAATTCTTAGCAGAAGTTGAGATGCTTAGCCGCCTTCACCACCG
AAATCTAGTTAAGTTGATTGGAATATGCACAGAAGAGTCTGCACGATGCCTCGTTTATGAACTAGTACCTAATGGCAGCGTGGAATCTCATTTGCATGGG
GATGATGAGGAATCTGCACCATTTGATTGGGATGTCCGGATCAAGATAGCACTTGGTGCTGCTAGAGGTCTGGCTTATCTTCATGAGGATTCGAGTCCCC
GTGTTATCCACAGAGATTTCAAATCCAGCAACATCTTATTGGAAAGTGATTACACACCAAAAGTTTCTGATTTTGGCTTGGCTCGAAGTGCCCTGGATGA
GGAAACCAGCCATATATCTACACGTGTCATGGGGACGTTTGGGTATGTAGCTCCAGAGTATGCAATGACCGGTCATCTTCTAGTGAAAAGTGACGTATAC
AGCTATGGGGTGGTCCTTCTTGAGCTTCTAACTGGAAGAAAGCCAGTAGATATGTTGCAACCGGCAGGTCAAGAGAATCTAGTAGCATGGGCACGACCGC
TTCTAACAAGCAAAGAAGGGCTTGCAATGATGATAGATCCATCTCTAGGCCCTGATGTGCCCTTCGATAGTGTGGCCAAGGTTGCAGCAATTGCATCAAT
GTGTGTGCAACCGGAGGTGTCTCACCGTCCATTCACGGGCGAAGTAGTTCAAGCATTGAAACTGGTATCCAATGAATGTGATGTGGCAAAGGAAGGAGGG
TCAGTAAGTTTCAGCCAAGATTTATCCCTTGACATGGAGGATGCTGGGGAGAGCACCACATCAGGCCAGGGGCAGCCATTCAACAACCAAATCATAGGGC
CAAACTATGAGTCTGATATCGAGAGAGGCGTGGCATTGTCGCATATGTTTTGTACGTCAGAGCGATTCAGTGGGCAACCGCAATCTGCATCGTTCAGGAG
GTACTCGAGTTCGGGACCACTGAGGCCCGGAAAAGGAAGAGAGTGGTGGCAGAGAATGAGACGATTGAGAGGGGAAAGTGTGAGTGAACACGGGGTGATG
TTCAGACAATGGGCAGGTTCATATTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10011933 pacid=23146536 polypeptide=Lus10011933 locus=Lus10011933.g ID=Lus10011933.BGIv1.0 annot-version=v1.0
MGTASLLLLVELCLIASVFNLDGSTGFSTSPSPGSISPVYPVSPFERHAPCSEAFLVDGLMIDSPVELAPAPAHEAISGQVPSNHPIPIGSPPSSTIRPP
SIIPGDAPSESPSLSVVRRSHSGASPPQSFERDVAPMPSDVSFLPPLPNISPPLETREGNVAPMPSNTSGGSISYAPPLLNMAPPPQNVSPMPSTNSSLH
GIAPPPPISPGQVAPMPSGAIGSIVPPAHSVVPPPPISQGHVPPMPSGESSSLVPPMASVAPPPQISEVPPARSVAPPSPVFQGQVAPTPSKTSGENGPL
VPPPNSVAPPPQNSEGKLAPPPSSDSISIAPPPHIIQGTAAPMPSNDTGGKEPGEKTPASMPASAPPTKALPSIPFIHPGVPQHSTLSAHQKNMSFYGSP
LEKPEPVALPPRTLHRKKPAIRTLAPEAIIPSLPPDPDVSSPSSTPGMTDWKRYGIPVTAPSTQIHKPLPPRIHSRTKASFPAVAPSEHRTVRHSNEPLA
PSVPFHSSPRFITHQSPSNSPSTSSQKKHSTRGGMSNGTPASSYVIPPRSSEEPALPPSLPTTRRTHFASPTSNPGFCIAGPASPSHHSIPKPDLGVSPV
PWPSSIAPNWTTIPILPPKDSPSSPLKSPMPKLPTIHALPPPPPNQDCSTIVCMEPYANTPPGSPCGCVLPMQVGLRLSVSLYTFFPLVSELAQEIAAGV
FMKQSQVRIMGANAASQEPEKTIVLIDLVPLGQKFDNTTALLTYQRFWQKQVVIQLSFFGDYEVLYVRYPGLPSSPPSSISGITIIDDGPYSGNGNNART
IKPLGIDVSKRQQKDGLSGSMIAIIALSASIVVILSSAAAWIFFLRNRGHGRDPTPTLAPIHASVAKPLGTIASALRSGLSSNSLSFGSSIAPYTGSAKT
FSASDIERATNNFDLSRLLGEGGFGRVYSGVLEDGTKIAVKVLKRDDQQGGREFLAEVEMLSRLHHRNLVKLIGICTEESARCLVYELVPNGSVESHLHG
DDEESAPFDWDVRIKIALGAARGLAYLHEDSSPRVIHRDFKSSNILLESDYTPKVSDFGLARSALDEETSHISTRVMGTFGYVAPEYAMTGHLLVKSDVY
SYGVVLLELLTGRKPVDMLQPAGQENLVAWARPLLTSKEGLAMMIDPSLGPDVPFDSVAKVAAIASMCVQPEVSHRPFTGEVVQALKLVSNECDVAKEGG
SVSFSQDLSLDMEDAGESTTSGQGQPFNNQIIGPNYESDIERGVALSHMFCTSERFSGQPQSASFRRYSSSGPLRPGKGREWWQRMRRLRGESVSEHGVM
FRQWAGSY
|
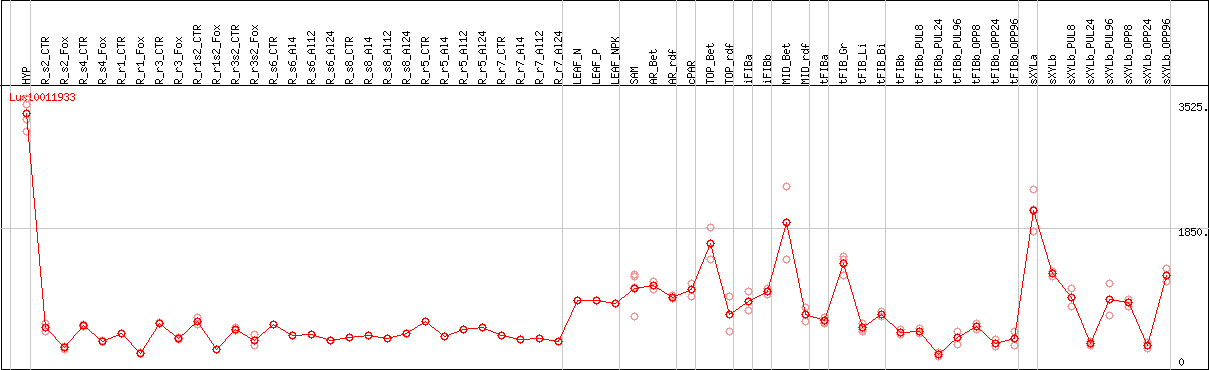
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10011933 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
