Lus10012249 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10012249 pacid=23172707 polypeptide=Lus10012249 locus=Lus10012249.g ID=Lus10012249.BGIv1.0 annot-version=v1.0
ATGGGCAAAGTCATTTCCACGCTTGAGCGAGCTCTGAGGGAAGGCCACTCACACGCTTTGCTCGAGTCTCCAACCGGCACTGGCAAATCTCTATCGCTGC
TCTGCTCAACTCTCGCATGGCAGCAGACCCAGAAGCTGAAGAATCTCTACGCCAATCTCACCCACTCCAAACCCGATCCTCAGGCGATGGCCGATCCTCT
TGCTCATGGCGGCGGATTCGTTCCTGAAACCCAGCCTTCGAGAAATCTGCCTCCCTCGATGGTAGAGCCAGGTAATCCAGCACCGGGGATGAAAAAGAAA
AAATCAGCACCTACCATCTTCTATGCATCGAGGACACATGCACAAATTTCTCAACTGGTGCGCGAGTATAAGAAAACTACATATCGCGTGCCAATGACAG
TATTGGCTTCGAGGAAACATTACTGCACAAACAAACTTGTAATTCAGACGGGGAATATTGATGAAGAATGTAAGCTGCTAATGAAGGATATTGAAGATGG
AGGTGGATGTTCATACTTCAAAAATGTGCATAAGCTCAAGGCTCATTCCTCACTTCAAAAGGGAGGTTGCTACGAGGTTCATGATATTGAAGATCTTGTG
AAAGTTGGAAAGGCTGTTAAAGGTTGTCCATATTACGCTGCCCGCTCAATGGCAGATGACGCGCAGTTGGTGTTTTGCCCCTACAATTACGTAATAAATC
CAATCATTCGAGGTGCGATGGAAATCGATATCAAAGGAGCCATTATTGTACTCGATGAAGCCCACAATATTGAAGACATCGCTCGTGACGCTGGTAGTGT
GGATCTAGAAGAAGATGTTTTACAGAAATTACAAACGGAGTTGGAAGAACTTTGCTCAAGTGATGGATTAACATACCAGCCCTTGTACGAGATGACACAG
GGCCTAATAAGTTGGATGGATCGGAAGAAAAGTACGCTTGAAATGCGTGACTTTCAGCAGTACTATTCTTGTTGGAGCGGTGATCAAGCTTTAAGGGAGC
TTCAAGAAGCTAATATTTCACAGCAGTATTTCCCTATCTTGTTAGAGTGTGCCCTAAAGGCTATTAAAGCTGCAACTGATGTGGAGTCAGAATTGCCTCT
TTTAAGCGGCATGTCAGTGACAATTTTGGAAGGTTTGTTCTCTGCGCTCACATATTTTTTTGCAAGAAATGGATCCCATATTTCTGATTACCAGCTTGTT
GTGCAACGAAAGGTCAAGAAAGACTCAAAAAACGATGTTGGTGGCTGGACACATATTTTTAGCTTGTGGTGCTTGAATCCGGCTGTTGTGTTCAAGGACA
TTGCTGATCTATCTTTGTCAGTCATCCTAACATCCGGGACTCTGTCACCTATGAATTCCTTCTCATCTGAGCTTGGAGTGAACTTTGGAAATTGTCTAGA
GGCCCCACATGTGGTTAATATTAAATCACAGGTGTGGACTGGTGTAATATCCTCTGGTCCTGGAAACTATCCCTTAAATGCCAGTTTCAAGACTGCAGGT
GCATTTTCCTTCCAGGATGCACTTGGGAAATCTTTGGAGGAAATTTTCAAAGTTGTTCCTGCTGGTTCCCTTGCCTTCTTCCCAAGTTATAATTTGCTGG
AAAAACTGTCCAAGCGGTGGCGTGAAACAGGTCAATGGGTGAGACTTAATGCGCAGAAGTCCCTCTTTGTCGAGCCAAGAGGAGGAACTCAAGAGGATTT
TGATTCTCTTATAAAGCGTTACTACAATTCAGTTTCCCGGCATAGCAAAGTTCCTGTTGGAACTAAGAAAAAGAACAAAAAACCAGATCTAAAGAACACT
AAGACACCTGAGCCTCCCACGACTTCTAAGAAAGATGGAGGTGCTTTTCTTGCCGTTTGCAGAGGAAAGGCCTCTGAAGGAATAGACTTTTCAGATGATA
ATGCCCGAGTAGTTGATATCCAAGTCTGTTTGAAGAAGAAATACAACGACAGTTACAAATCAAGTAAGAACCTTCTAAGTGGGAGTGAGTGGTACTGTCA
CCAAGCTTTCCGAGCACTGAATCAAGCTGCAGGACGTTGCATACGGCATAAGTCTGACTATGGAGCCATCATCTTCTTAGATGAACGTTTTCAGGAAGAG
AGGAACACAGTATACGTTTCCAAGTGGCTCAGACAGTCCATTCAAAACTATAGAAGCTTTGATGAGTCACTTGAACGATTGAAATGCTTTTTCAGAGACG
CCAAGAATATACTTGGTGAAACTACACCTACTCCAGTGCCAAATTATACGGTTACGGAGGAGAAAGATGTACCTAAAGATCTAGAGAATAGCTGTACTGG
GGGAAGCTCTCATAGATTGAGTGGACCTTGTTGCCGTGGACAGAAGCTAGTTGCTGCACCAAAGTGTAGCCCCTCCGTGAACTTAGACCTCGAGCCTGAA
GCCCAGCCAATCGTAATAGAAGACGACGGCATTGAGAGTTGCCCAGTTACTGATCTAGAATGTAATTCACCAATGGGCACAAGGTTTGATGATGCATTAA
CCATTGTCAAGGAAACTCCTGATATAGATAGAAATAGCCCAAAAACTTTATCTACAGGTATGGATTATGGCTCAAGCATGATTGAACCCTCTTATGAGAT
GACAACTCCCACAACAAGCCTTCTCCAAGTCCAGGGTTCATCAATAGGCAGTCCCAAGAAACTGTTCTCTATATCCACCTTCAGCCCGTCAGAAGAAGTT
GAAACTTCTCTTATTCGAAGTGTTAATTCTCGCACCCAGAAAAGAAGGAAATTGGTAGACTCATCACCTTTGGTTCTTCCTTCAGATGACAACTCTGGTA
TCTGCAATGTCGCAACTTTGGGATGTGTCAGTTCCGAAAGACGTCCTTTCTCTGGTTTGGAGTTAAATGGCACGATGCAACGTTTATCTTTTGGTCCATC
TGTAAATAAAAACCTGAAAATTTCCTGCTTTCTTTGTGAAAAACCATTGGGTCGTCCAGAGAACGACCTTTTTGTTGAGTGCCTGGCAACTTCGTCGTCA
AAAGTTCATTTAGTGTCCCTTGCAAAGGACGGTTTGGGAGATGAAGCGGAGAATACTTCAACAAAGGTGCCTGTTGTAGTAGCAGAAAGACTATCAGTCG
ATCATATGCTATGTAATAGTGCTCGAGAAGATGTTCCTGCTGAAGGAGTTTGGTGCGAGGAAGATGGATGTGTTTTCAAGTCAATCTATTGTCCATTCTG
CACTAATCCCCATTGTCTTGGTGTTCAAGTCATGGCAACTGATTCATCAAATGTTCAGTTACTTAACAAGATAGTTCTTTTGTACGATCGCCTCCAAGTC
CAGACTGTGGGTACTTCAGAGGACAAGGCTTCAAAAAATCATCAGAACCACGTCCCAGTGAATGGTTGTGTTGGTAAAACTGCAGACCCGGATCAATTCA
AGCGATTTTCATTTTCACCACAACCGAATTCTAATGGCTGGAGAAGTACAAAGCCAAATAATCCGCAACACAATTCGCTTCACGATAAATATGAGATAAA
CGAACCTCCCAAAGCCGACGGTAGAGGTCTTGAAATTGGAGGACTAGTGCAGCGTGTTGATGATCCAGGTAGCGTGAAGAAGAAAGTTCTTGAGCTGAAT
GTCAATAGCGCTCGCAGTTTCTCGTTACGTGAGCTGGCGGCTGCCACTCAGAATTTCAGGGATGCGAATCTGATCGGAGAAGGAGGGTTCGGTAGGGTTT
ATAAAGGCCGCCTGGAAACTGGCGAGATTGTTGCGGTGAAACAATTGAACCAAGATGGTATGCAGGGGAATCAAGAATTCATCATCGAGGTTCTTATGCT
GAGTCTCTTACACCACCCTAATCTCGTCACCTTGACTGGCTACTGTGCTGCCGGCGACCAGAGACTGTTGGTTTACGAGTACATGCCCATGGGAAGCTTG
GAAACTCATCTTTTTGACGCGGATCCTGGTAAAGAGCCACTTAGTTGGGGCACAAGGATGAAGATTGCCGTTGATGCAGCAAGAGGCCTTCAACACCTGC
ATTGCAAGACAACTCCACCAGTAATTTACCGCGACCTGAAATCAGCAAACATCTTGCTAGACAACGATTTCCACGCGAAGCTATCAGATTTCGGGCTAGC
GAAACTAGGGCCCACTGGCGATAACACCCACGTTTCAACCAGGGTTATGGGAACGTACGGCTACTGTGCCCCAAAGTATGCCATGACTGGGAAATTGACT
CTCAAATCCGACATCTATAGCTTCGGTGTCCTTCTGTTGGAGCTTATTACTGGCCGGAAGGCTATGGATTATGAAAGAAGGCAACGGGAGCAGAACCTGG
TAGCTTGGTGTCGGCCATTTCTGAAAGATCCGAAGAAGTACTTACAAATGGCCGATCCACAGCTTCAAAGACGCTACCCTCGCAGGTGCTTAACCTACGC
TGTTGCCATAACCGAAATGTGTCTGCACGAGGAAGCACATTTCCGGCCACTAATCGGGGACATAGTAGTGGCCCTGGAGTACTTGGAAGCACAGAGTCTA
CAGGCTGTGCAGAACACAAGCTCGAAAGAAAGGCACGAAGCGACATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10012249 pacid=23172707 polypeptide=Lus10012249 locus=Lus10012249.g ID=Lus10012249.BGIv1.0 annot-version=v1.0
MGKVISTLERALREGHSHALLESPTGTGKSLSLLCSTLAWQQTQKLKNLYANLTHSKPDPQAMADPLAHGGGFVPETQPSRNLPPSMVEPGNPAPGMKKK
KSAPTIFYASRTHAQISQLVREYKKTTYRVPMTVLASRKHYCTNKLVIQTGNIDEECKLLMKDIEDGGGCSYFKNVHKLKAHSSLQKGGCYEVHDIEDLV
KVGKAVKGCPYYAARSMADDAQLVFCPYNYVINPIIRGAMEIDIKGAIIVLDEAHNIEDIARDAGSVDLEEDVLQKLQTELEELCSSDGLTYQPLYEMTQ
GLISWMDRKKSTLEMRDFQQYYSCWSGDQALRELQEANISQQYFPILLECALKAIKAATDVESELPLLSGMSVTILEGLFSALTYFFARNGSHISDYQLV
VQRKVKKDSKNDVGGWTHIFSLWCLNPAVVFKDIADLSLSVILTSGTLSPMNSFSSELGVNFGNCLEAPHVVNIKSQVWTGVISSGPGNYPLNASFKTAG
AFSFQDALGKSLEEIFKVVPAGSLAFFPSYNLLEKLSKRWRETGQWVRLNAQKSLFVEPRGGTQEDFDSLIKRYYNSVSRHSKVPVGTKKKNKKPDLKNT
KTPEPPTTSKKDGGAFLAVCRGKASEGIDFSDDNARVVDIQVCLKKKYNDSYKSSKNLLSGSEWYCHQAFRALNQAAGRCIRHKSDYGAIIFLDERFQEE
RNTVYVSKWLRQSIQNYRSFDESLERLKCFFRDAKNILGETTPTPVPNYTVTEEKDVPKDLENSCTGGSSHRLSGPCCRGQKLVAAPKCSPSVNLDLEPE
AQPIVIEDDGIESCPVTDLECNSPMGTRFDDALTIVKETPDIDRNSPKTLSTGMDYGSSMIEPSYEMTTPTTSLLQVQGSSIGSPKKLFSISTFSPSEEV
ETSLIRSVNSRTQKRRKLVDSSPLVLPSDDNSGICNVATLGCVSSERRPFSGLELNGTMQRLSFGPSVNKNLKISCFLCEKPLGRPENDLFVECLATSSS
KVHLVSLAKDGLGDEAENTSTKVPVVVAERLSVDHMLCNSAREDVPAEGVWCEEDGCVFKSIYCPFCTNPHCLGVQVMATDSSNVQLLNKIVLLYDRLQV
QTVGTSEDKASKNHQNHVPVNGCVGKTADPDQFKRFSFSPQPNSNGWRSTKPNNPQHNSLHDKYEINEPPKADGRGLEIGGLVQRVDDPGSVKKKVLELN
VNSARSFSLRELAAATQNFRDANLIGEGGFGRVYKGRLETGEIVAVKQLNQDGMQGNQEFIIEVLMLSLLHHPNLVTLTGYCAAGDQRLLVYEYMPMGSL
ETHLFDADPGKEPLSWGTRMKIAVDAARGLQHLHCKTTPPVIYRDLKSANILLDNDFHAKLSDFGLAKLGPTGDNTHVSTRVMGTYGYCAPKYAMTGKLT
LKSDIYSFGVLLLELITGRKAMDYERRQREQNLVAWCRPFLKDPKKYLQMADPQLQRRYPRRCLTYAVAITEMCLHEEAHFRPLIGDIVVALEYLEAQSL
QAVQNTSSKERHEAT
|
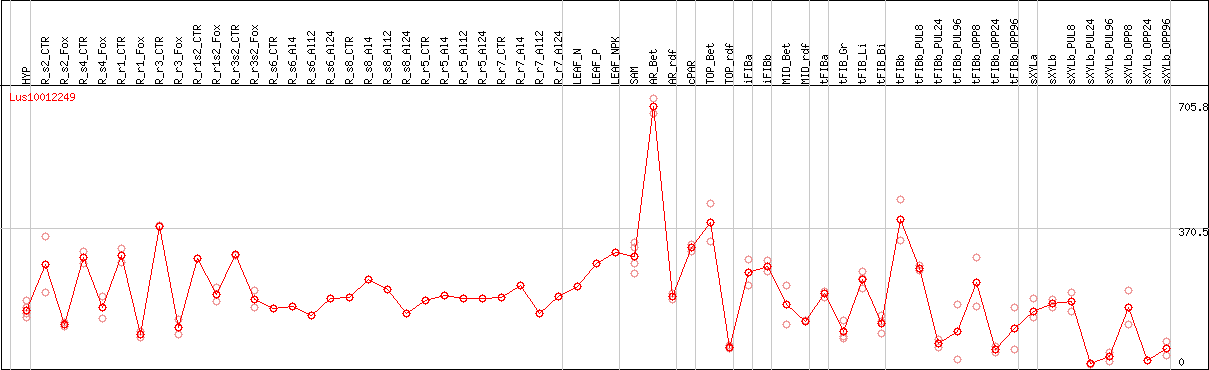
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10012249 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
