Lus10012317 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10012317 pacid=23176792 polypeptide=Lus10012317 locus=Lus10012317.g ID=Lus10012317.BGIv1.0 annot-version=v1.0
ATGGCGGCACTACTGGCTTCACACAGCTGTTATTGCCTTCATGTGGAGTCCTTGAATCAAGGTAGAAATATCTCGGAGAGCTTCAGTTTCTCAAGCTCAG
TTTCCAGCTCATCATTTGCAGTGTCTAAGAGGCAGCATGATCATACTCCGTCGAGATCTCGATTTCAAGTGCGGATGACTCAGACTGAATCATCCCCAGC
ATCCAAGTTAGGCCCAAATGGACGGCCGGTTAAAATGGTGCCAGTGAATGAAGTGGTGAAGAGGAGGACGCCAAATGGTAGTAGTAGGAGTAGTAAGGTA
GATTTAGTGAATGGGAAAAGGCAGGTGATTAATGGAGCTGCTAGTATAGTAAAGAAAAGTTCTTCAGTAGTTCCTGTGAGAAGATTGGCAGATAATAAAC
TTCCTCCTCTTGAGGGTTTGAAAGCTCTTCCTACTGATGAAGGCTTCAGTTGGGCTGATGACAATTACAACAATGTTCAGAGGAATATTGACATCTGGTC
TTTCTTCCTTGCTTTTCGGGTTCGTCTCACGTTGGATAATGCCAAATGGGCATATTTGGGGAGTTTCAGTGAAGAGAAGCAGAAATTGCGAAGGAAGAAC
ACAGCTGGGTGGCTAAGGGAGCAAGTATTGCAGCTCGGTCCGACTTTTATCAAACTCGGACAGTTGTCTTCGACAAGGTCGGACCTTTTCCCCAAGGAAT
TTGTTGATGAGCTTGCCAAGTTGCAGGATAGAGTCCCAGCATTCCCGCCTAGTAAAGCAAGAAGCTTCGTTGAGCGAGAACTGGGAGCTCCTATAGATGT
GCTGTTCAAGGAGTTTGAAGACAGGCCGATAGCAGCAGCTAGCCTTGGTCAGGTGCATCGAGCTATCCTACACAACGGAGAGAAAGTCGTCGTCAAAATT
CAGAGGCCTGGCCTCAAGAAGCTTTTTGATATTGATCTAAATAATATGAAGATAGTTGCAGGATACTTGCAGAAAAGTGACACTTTTGGTGGTCCGACTA
ACGATTGGGTTGGTATATACGAGGAATGCGCCACGATTTTATACCAGGAGATTGACTACATCAATGAAGGAAAAAATGCAGATAGATTCCGTCGAGATTT
CAGGAATATAAAGTGGGTCCGAGTACCTATGGTATTTTGGGACTACACATCTACAAAGGTCTTGACTTTGGAATATGTACCAGGCATTAAGATTAATGAG
TTGGACATGCTAGATTCCCGGGGATACAATCGTTCTCGGGTTTCATCACGAGCCATCGAGGCATACTTGATTCAGATTCTGAAAACCGGTTTCTTCCATG
CGGATCCTCATCCCGGAAATCTCGCAGTTGATATCGACGAATCTCTCATCTATTACGACTTCGGTATGATGGGAGAGATAAAGGCTTTTACTAGGGAGAG
GTTGCTTGGACTTTTCTATGCAGTTTATGAGAAAGATGCCAGAAAGGTTATGGAGAGCCTGATAGACCTTGGAGCACTTCAGCCCACTGGGGACTTGTCT
TCTGTGAGGAGATCCGTTCAATACTTCTTAGACAACTTGTTAGGGCAGACGCCTGATCAACAGCAGACTTTAGCAGCCATTGGGGAGGATTTGTTTTCAA
TAGCTCAGGATCAACCATTTCGGTTCCCATCCACCTTTACGTTCGTTATAAGGGCGTTTTCAACCCTTGAAGGCATAGGCTACATTCTCGATCCGGACTT
TTCATTCGTGAAGATCGCTGCCCCTTATGCCCAGGAGCTTTTAGATATCAGACAGAGGCAGCAGCCTGGAACACAACTAGTAGAGCAAATAAGGAAACAG
GCCAACGATGCTAGAAGTTCCACAATGTCCATGCCGTATAGAGTCCAGCGGATAGAAGAGTTCGTGAAGCAACTCGAATCCGGAGACTTCAAACTCCGCG
TCCGGGTACTAGAGTCGGAGAGAGCAGCAAGGAAAGCAACCATCTTACAGATGGCAACAATGTACACAGTGTTAGGAGGCACACTTGTAAACCTTGGGGT
AACCTTTGGCAGTCAGGGGAGCCAACTTGTTGCCAATGGTTCATTTGTAGGCGCAGGAGTATTCTTGTTACTGTTGATAAGGTCAATGCAAAGGCATTGT
ATTGTGCTGCTTCGACTTTCCCCTTCGCTGCTTCTTATCAGAGTTGCATCTAAAGTAACATCCATGGCTTCAGCTCCATCAGCTTCAGATATAAAGCCTA
AAGATGTTTGCATTGTTGGAGTTGCACGAACCCCAATGGGTGGATTTCTTGGAACCTTGTCTTCTGTTCCTGCAACCAAGCTTGGATCTATAGCTATTGA
AGCTGCTCTTAAGAGAGCCAATGTGGATCCATCCCTAGTACAGGAAGTTATCTTTGGTAATGTTCTGAGTGCGAATCTAGGACAGGCTCCAGCCAGGCAG
GCGGCATTGGGTGCAGGAATTCCTAGCACAGTCATCTGCACAACTGTCAACAAAGTGTGTGCTTCAGGGATGAAGGCAACTATGCTTGCGGCCCAGAGTA
TTCAGTTGGGCATCAACAATGTTGTTGTCGCTGGAGGCATGGAGAGCATGTCCAATGTACCTAAATATCTAGCGGAAGCAAGGAAGGGATCTCGGCTTGG
ACATGATTCACTTGTTGATGGGATGCTTAAGGACGGACTATGGGATGTGTACAATGATTTTGGCATGGGAAACTGTGCTGAAATATGTGCAGAACAGCAC
TCCATAACAAGGGAGCAGCAGGATGACTTTGCAGTTCAAAGTTTTGAGCGTGGCATTGCTGCACAGGACAGTGCTGCTTTTGCATGGGAAATAGTTCCGG
TTGAGGTTTCTGGCGGGAGGGGAAAGCCGTCAACCCTTGTTGATAAAGATGAAGGTTTAGGGAAGTTTGATGCTGCTAAATTAAGGAAGCTCCGACCAAG
TTTCAAACCAAATGGAGGAACCGTTACTGCTGGGAATGCTTCTAGCATCAGTGACGGTGCAGCTGCTCTAGTTCTGGTGAGTGGAGAGAAGGCATTAGAG
CTCGGGTTACACGTGATTGCAAAGATCACTGGATATGCCGATGCTGCACAGGCACCAGAGCTTTTCACCACAGCTCCAGCTCTGGCCATACCTAAAGCTA
TTTCAAACGCTGGATTGGATGCTTCCCGAGTAGATTATTACGAGATAAACGAAGCCTTCGCTGTTGTAGCCCTTGCTAATCAGAAGTTGCTGAGTCTTAA
TCCGGAAAAGATTAACGTACATGGCGGAGCAGTCTCTTTGGGGCATCCACTAGGATGCAGTGGCGCTCGGATCCTGGTCACGCTTTTGGGAGTGTTGAGG
CAGAAGAATGCGAAATACGGCGTGGGCGGAGTATGCAATGGGGGAGGTGGAGCATCAGCACTGGTTGTGGAGCTTCTATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10012317 pacid=23176792 polypeptide=Lus10012317 locus=Lus10012317.g ID=Lus10012317.BGIv1.0 annot-version=v1.0
MAALLASHSCYCLHVESLNQGRNISESFSFSSSVSSSSFAVSKRQHDHTPSRSRFQVRMTQTESSPASKLGPNGRPVKMVPVNEVVKRRTPNGSSRSSKV
DLVNGKRQVINGAASIVKKSSSVVPVRRLADNKLPPLEGLKALPTDEGFSWADDNYNNVQRNIDIWSFFLAFRVRLTLDNAKWAYLGSFSEEKQKLRRKN
TAGWLREQVLQLGPTFIKLGQLSSTRSDLFPKEFVDELAKLQDRVPAFPPSKARSFVERELGAPIDVLFKEFEDRPIAAASLGQVHRAILHNGEKVVVKI
QRPGLKKLFDIDLNNMKIVAGYLQKSDTFGGPTNDWVGIYEECATILYQEIDYINEGKNADRFRRDFRNIKWVRVPMVFWDYTSTKVLTLEYVPGIKINE
LDMLDSRGYNRSRVSSRAIEAYLIQILKTGFFHADPHPGNLAVDIDESLIYYDFGMMGEIKAFTRERLLGLFYAVYEKDARKVMESLIDLGALQPTGDLS
SVRRSVQYFLDNLLGQTPDQQQTLAAIGEDLFSIAQDQPFRFPSTFTFVIRAFSTLEGIGYILDPDFSFVKIAAPYAQELLDIRQRQQPGTQLVEQIRKQ
ANDARSSTMSMPYRVQRIEEFVKQLESGDFKLRVRVLESERAARKATILQMATMYTVLGGTLVNLGVTFGSQGSQLVANGSFVGAGVFLLLLIRSMQRHC
IVLLRLSPSLLLIRVASKVTSMASAPSASDIKPKDVCIVGVARTPMGGFLGTLSSVPATKLGSIAIEAALKRANVDPSLVQEVIFGNVLSANLGQAPARQ
AALGAGIPSTVICTTVNKVCASGMKATMLAAQSIQLGINNVVVAGGMESMSNVPKYLAEARKGSRLGHDSLVDGMLKDGLWDVYNDFGMGNCAEICAEQH
SITREQQDDFAVQSFERGIAAQDSAAFAWEIVPVEVSGGRGKPSTLVDKDEGLGKFDAAKLRKLRPSFKPNGGTVTAGNASSISDGAAALVLVSGEKALE
LGLHVIAKITGYADAAQAPELFTTAPALAIPKAISNAGLDASRVDYYEINEAFAVVALANQKLLSLNPEKINVHGGAVSLGHPLGCSGARILVTLLGVLR
QKNAKYGVGGVCNGGGGASALVVELL
|
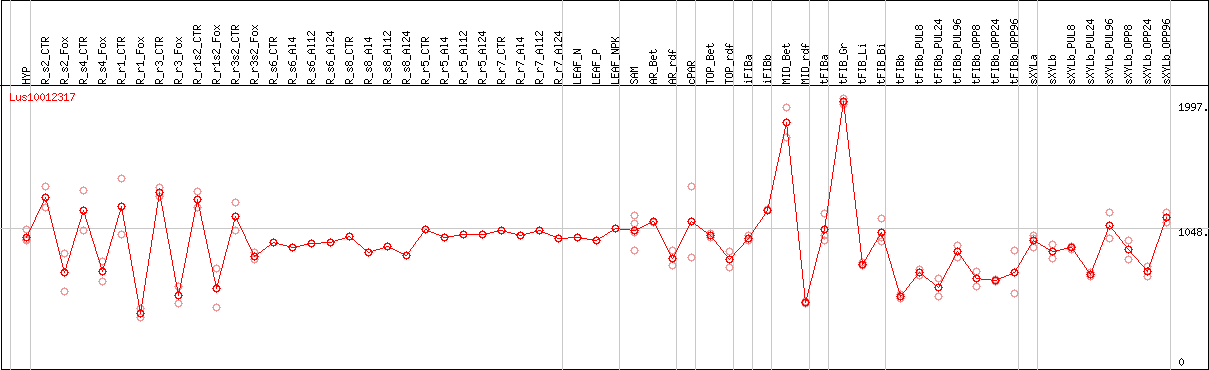
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10012317 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
