External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G47650 254 / 2e-84
ATNUDX2, ATNUDT2
ARABIDOPSIS THALIANA NUDIX HYDROLASE HOMOLOG 2, nudix hydrolase homolog 2 (.1)
AT4G12720 249 / 2e-82
ATNUDX7, GFG1, AtNUDT7, NUDT7
GROWTH FACTOR GENE 1, Arabidopsis thaliana Nudix hydrolase homolog 7, MutT/nudix family protein (.1.2.3.4)
AT4G25434 233 / 3e-76
ATNUDT10
nudix hydrolase homolog 10 (.1.2)
AT2G04450 231 / 3e-75
ATNUDX6, ATNUDT6
nucleoside diphosphates linked to some moiety X 6, Arabidopsis thaliana nucleoside diphosphate linked to some moiety X 6, nudix hydrolase homolog 6 (.1)
AT5G47240 225 / 5e-72
ATNUDT8
nudix hydrolase homolog 8 (.1)
AT2G04430 222 / 2e-71
ATNUDT5
nudix hydrolase homolog 5 (.1)
AT2G04440 115 / 5e-31
MutT/nudix family protein (.1)
AT5G06340 40 / 0.0009
ATNUDX27
nudix hydrolase homolog 27 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10006366
517 / 0
AT5G47650 281 / 4e-95
ARABIDOPSIS THALIANA NUDIX HYDROLASE HOMOLOG 2, nudix hydrolase homolog 2 (.1)
Lus10012319
428 / 1e-152
AT5G47650 310 / 4e-106
ARABIDOPSIS THALIANA NUDIX HYDROLASE HOMOLOG 2, nudix hydrolase homolog 2 (.1)
Lus10006365
413 / 2e-146
AT4G12720 300 / 1e-101
GROWTH FACTOR GENE 1, Arabidopsis thaliana Nudix hydrolase homolog 7, MutT/nudix family protein (.1.2.3.4)
Lus10039079
251 / 4e-82
AT5G47650 345 / 3e-119
ARABIDOPSIS THALIANA NUDIX HYDROLASE HOMOLOG 2, nudix hydrolase homolog 2 (.1)
Lus10038780
248 / 8e-81
AT5G47650 345 / 1e-118
ARABIDOPSIS THALIANA NUDIX HYDROLASE HOMOLOG 2, nudix hydrolase homolog 2 (.1)
Lus10040169
213 / 5e-67
AT5G47240 409 / 7e-143
nudix hydrolase homolog 8 (.1)
Lus10004371
212 / 3e-64
AT5G47240 410 / 2e-139
nudix hydrolase homolog 8 (.1)
Lus10004195
42 / 0.0003
AT5G06340 236 / 1e-78
nudix hydrolase homolog 27 (.1)
Lus10029398
42 / 0.0003
AT5G06340 223 / 3e-73
nudix hydrolase homolog 27 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G168400
291 / 5e-98
AT5G47650 331 / 2e-113
ARABIDOPSIS THALIANA NUDIX HYDROLASE HOMOLOG 2, nudix hydrolase homolog 2 (.1)
Potri.005G077500
285 / 6e-96
AT5G47650 322 / 2e-110
ARABIDOPSIS THALIANA NUDIX HYDROLASE HOMOLOG 2, nudix hydrolase homolog 2 (.1)
Potri.003G106100
269 / 5e-90
AT5G47650 332 / 9e-115
ARABIDOPSIS THALIANA NUDIX HYDROLASE HOMOLOG 2, nudix hydrolase homolog 2 (.1)
Potri.001G127700
268 / 1e-89
AT5G47650 323 / 1e-111
ARABIDOPSIS THALIANA NUDIX HYDROLASE HOMOLOG 2, nudix hydrolase homolog 2 (.1)
Potri.001G368000
262 / 4e-87
AT5G47650 342 / 6e-119
ARABIDOPSIS THALIANA NUDIX HYDROLASE HOMOLOG 2, nudix hydrolase homolog 2 (.1)
Potri.016G006000
244 / 7e-80
AT5G47650 352 / 3e-122
ARABIDOPSIS THALIANA NUDIX HYDROLASE HOMOLOG 2, nudix hydrolase homolog 2 (.1)
Potri.006G005400
244 / 1e-79
AT5G47650 347 / 7e-120
ARABIDOPSIS THALIANA NUDIX HYDROLASE HOMOLOG 2, nudix hydrolase homolog 2 (.1)
Potri.001G154300
215 / 5e-68
AT5G47240 416 / 2e-145
nudix hydrolase homolog 8 (.1)
Potri.003G080400
208 / 2e-65
AT5G47240 401 / 1e-139
nudix hydrolase homolog 8 (.1)
Potri.018G069100
197 / 2e-61
AT5G47240 316 / 3e-106
nudix hydrolase homolog 8 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0261
NUDIX
PF00293
NUDIX
NUDIX domain
Representative CDS sequence
>Lus10012320 pacid=23176829 polypeptide=Lus10012320 locus=Lus10012320.g ID=Lus10012320.BGIv1.0 annot-version=v1.0
ATGGCGTCGTTTTCCTCTTCTTCAGAAACTGCGGATGCCAATGTTCATGGAGTGGAGGGGATTGAACTGCTGGAGGCTACTCCGGATCCGTATGGTGGGC
TAACCATGGAAATCAAGGAGCCTATGGAAGCTGTGGATTTCGCAGCTCGGCTCAGGTATTCCATCTCCAAATGGATTGAACAGGAGAAGAAGGGTATTTG
GATCAAATTGCCCACTCGCCAGTCTAACCTCGTCGATTCAGTTATTAAGGAAGGGTTTAGGTTTCACCACGCTGAATCAGACTACGTGATGCTAGTGCGT
TGGCTGCCTGAGAATCTTCCTGATACTCTTCCCATCAATGCCACTCACAGGATTGGCGTTAAGGCGCTCGTGGTCAACGAAAACAAAGAGGTGCTGGTAG
TTACAGAGAAACTTGGCAAATTGGCAGGCATCGAGTATTGGAACTTGCCAACTGGTGTAGCCCAAGAGGGAGAAGACATATGGGCAGCTGCAGTTAGAGA
AGTGAAAGAAGAAACTGGAATTGATGCTGAATTTGAAGAAATGATAACATTCAGGCAAAGTCACAAGTCATACTTTAGCAAATCAGACCTGATGTTTGTC
TGCCTGTTGCGACCAGTTTCTGCTGACATCACTATTCAAGAATCAGAGATTGTAGCATCCAAGTGGATGCCGGTCGACGAGTATGTAGAGCAGCCCCACA
ATAAGAAGCATGGGCAGTTCAGATACGTAGCGGAGATTTGCAAAGCCAAGGTGGAAGGTGCATATGTGGGTTTCTCTGCTATGGCAGTGAAATCTTCTTC
AGGGAAAAAGGTTGACCTCTACTCTAACCACTGGGCAGAATGA
AA sequence
>Lus10012320 pacid=23176829 polypeptide=Lus10012320 locus=Lus10012320.g ID=Lus10012320.BGIv1.0 annot-version=v1.0
MASFSSSSETADANVHGVEGIELLEATPDPYGGLTMEIKEPMEAVDFAARLRYSISKWIEQEKKGIWIKLPTRQSNLVDSVIKEGFRFHHAESDYVMLVR
WLPENLPDTLPINATHRIGVKALVVNENKEVLVVTEKLGKLAGIEYWNLPTGVAQEGEDIWAAAVREVKEETGIDAEFEEMITFRQSHKSYFSKSDLMFV
CLLRPVSADITIQESEIVASKWMPVDEYVEQPHNKKHGQFRYVAEICKAKVEGAYVGFSAMAVKSSSGKKVDLYSNHWAE
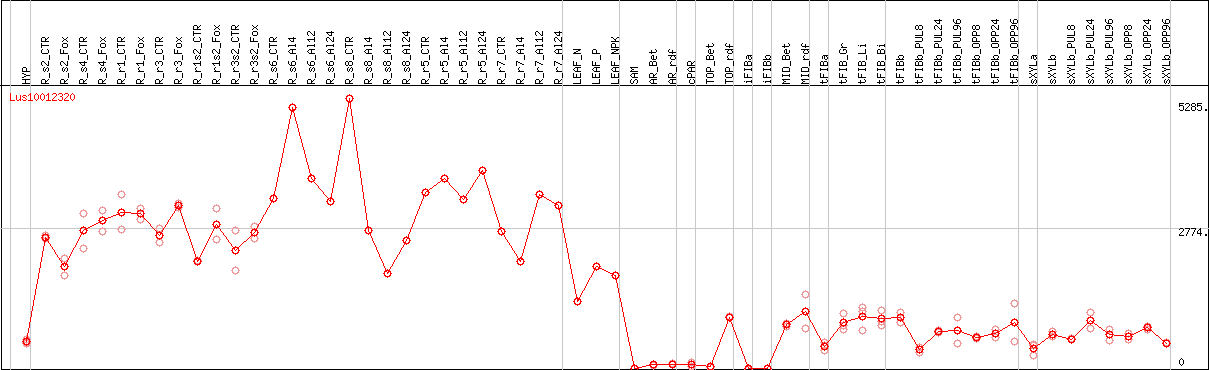
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10012320 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.