External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G18990 37 / 0.0008
B3
REM39, VRN1
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10037323
46 / 5e-07
AT3G18990 79 / 2e-16
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Lus10009214
43 / 2e-06
AT1G49480 40 / 6e-05
related to vernalization1 1 (.1)
Lus10015266
43 / 7e-06
AT5G58610 558 / 7e-176
PHD finger transcription factor, putative (.1)
Lus10039303
43 / 7e-06
AT3G18990 76 / 1e-14
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Lus10002934
42 / 3e-05
AT3G18990 112 / 5e-28
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Lus10015248
41 / 4e-05
AT4G33280 50 / 3e-07
AP2/B3-like transcriptional factor family protein (.1)
Lus10009688
40 / 5e-05
AT3G18990 113 / 2e-28
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Lus10010756
40 / 5e-05
AT3G18990 99 / 2e-23
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Lus10023079
40 / 9e-05
AT3G18990 69 / 3e-13
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G147000
45 / 9e-07
AT3G18990 258 / 3e-83
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Potri.004G230100
38 / 0.0003
AT3G18990 161 / 1e-44
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0405
DNA_b-psBarrel
PF02362
B3
B3 DNA binding domain
Representative CDS sequence
>Lus10012521 pacid=23143810 polypeptide=Lus10012521 locus=Lus10012521.g ID=Lus10012521.BGIv1.0 annot-version=v1.0
ATGCAGACCATTCCAAAATCGTTCATGGACAATGTCGAGGAAGTTTACGGTGGAGCCGAGACCGCAGAGTTATGGGTAGCAGCCAAATGCCGGAGAGTGA
GGATCCGAAGAAGTGTGAAAGAAAGTCGCAGCAATTTGGGTTCTGGATGGGGCGATTTCGCAACGGACAATTCTCTGGAAGTCGGAGATGTTTGTGTTTT
CGAGCTTAAATGTGAAGAGGATGATGGTGTTGGTGGTGGAAGCAGTGGAAGAAAGGTGGTGTTTGGTGTTCATATATTCAGAAGAAGATGA
AA sequence
>Lus10012521 pacid=23143810 polypeptide=Lus10012521 locus=Lus10012521.g ID=Lus10012521.BGIv1.0 annot-version=v1.0
MQTIPKSFMDNVEEVYGGAETAELWVAAKCRRVRIRRSVKESRSNLGSGWGDFATDNSLEVGDVCVFELKCEEDDGVGGGSSGRKVVFGVHIFRRR
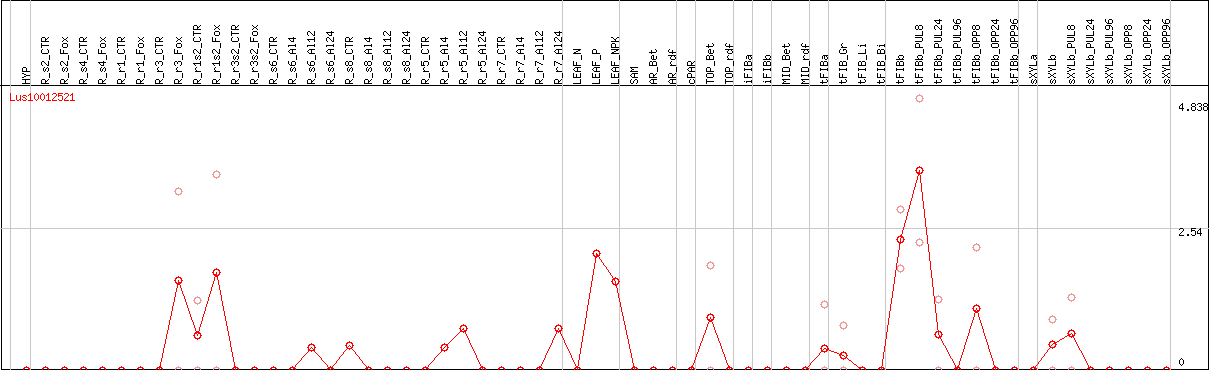
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10012521 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.