External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G45140 71 / 2e-15
ATLOX2, LOX2
ARABIODOPSIS THALIANA LIPOXYGENASE 2, lipoxygenase 2 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10041646
189 / 5e-57
AT3G45140 939 / 0.0
ARABIODOPSIS THALIANA LIPOXYGENASE 2, lipoxygenase 2 (.1)
Lus10027379
120 / 1e-32
AT1G72520 867 / 0.0
Arabidopsis thaliana lipoxygenase 4, PLAT/LH2 domain-containing lipoxygenase family protein (.1)
Lus10002547
120 / 1e-32
AT3G45140 971 / 0.0
ARABIODOPSIS THALIANA LIPOXYGENASE 2, lipoxygenase 2 (.1)
Lus10031238
102 / 7e-28
AT3G45140 226 / 1e-68
ARABIODOPSIS THALIANA LIPOXYGENASE 2, lipoxygenase 2 (.1)
Lus10031810
103 / 1e-26
AT3G45140 959 / 0.0
ARABIODOPSIS THALIANA LIPOXYGENASE 2, lipoxygenase 2 (.1)
Lus10031809
94 / 2e-23
AT3G45140 753 / 0.0
ARABIODOPSIS THALIANA LIPOXYGENASE 2, lipoxygenase 2 (.1)
Lus10031236
92 / 2e-22
AT3G45140 805 / 0.0
ARABIODOPSIS THALIANA LIPOXYGENASE 2, lipoxygenase 2 (.1)
Lus10039094
84 / 1e-19
AT3G45140 946 / 0.0
ARABIODOPSIS THALIANA LIPOXYGENASE 2, lipoxygenase 2 (.1)
Lus10002545
65 / 3e-13
AT3G45140 991 / 0.0
ARABIODOPSIS THALIANA LIPOXYGENASE 2, lipoxygenase 2 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G046200
85 / 2e-20
AT3G45140 979 / 0.0
ARABIODOPSIS THALIANA LIPOXYGENASE 2, lipoxygenase 2 (.1)
Potri.001G015500
85 / 3e-20
AT3G45140 975 / 0.0
ARABIODOPSIS THALIANA LIPOXYGENASE 2, lipoxygenase 2 (.1)
Potri.009G022400
78 / 9e-18
AT3G45140 1008 / 0.0
ARABIODOPSIS THALIANA LIPOXYGENASE 2, lipoxygenase 2 (.1)
Potri.001G015400
73 / 3e-16
AT3G45140 990 / 0.0
ARABIODOPSIS THALIANA LIPOXYGENASE 2, lipoxygenase 2 (.1)
Potri.001G015300
55 / 1e-09
AT3G45140 943 / 0.0
ARABIODOPSIS THALIANA LIPOXYGENASE 2, lipoxygenase 2 (.1)
Potri.010G057100
40 / 0.0001
AT1G67560 1144 / 0.0
Arabidopsis thaliana lipoxygenase 6, PLAT/LH2 domain-containing lipoxygenase family protein (.1)
Potri.008G178000
39 / 0.0003
AT1G67560 1155 / 0.0
Arabidopsis thaliana lipoxygenase 6, PLAT/LH2 domain-containing lipoxygenase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00305
Lipoxygenase
Lipoxygenase
Representative CDS sequence
>Lus10012570 pacid=23143863 polypeptide=Lus10012570 locus=Lus10012570.g ID=Lus10012570.BGIv1.0 annot-version=v1.0
ATGAGGGCCGCTAGTACCTTGATCGGTAACGTGGGCAGCAAGGTGCTGGGGTTCAGTAGCTTCTACGTGCCCCGGGATGAAGAGTTCTCTGAAATTAAGC
AGGCAAACTTGTCCGTCAAGAGACTCTTCTCGCTTTTGCATTCCTTGTTGCCCATCCTGGAAACCATTTTTATCGGCCCTGCCCTTGGATTCCCACACTT
TACAGCCATCAATAAGATGTTCGGCCAAGGACTCATCTTGCCTCCCATTTCCTCAACCAATCTAAAAGCCGTCATCCCAACTCTTATCAGGAATACCGCT
GAAGCAAGTGATGACATATTGCAGTTTGAGATCCCTCCAACAATGGAAAGTGAGTAA
AA sequence
>Lus10012570 pacid=23143863 polypeptide=Lus10012570 locus=Lus10012570.g ID=Lus10012570.BGIv1.0 annot-version=v1.0
MRAASTLIGNVGSKVLGFSSFYVPRDEEFSEIKQANLSVKRLFSLLHSLLPILETIFIGPALGFPHFTAINKMFGQGLILPPISSTNLKAVIPTLIRNTA
EASDDILQFEIPPTMESE
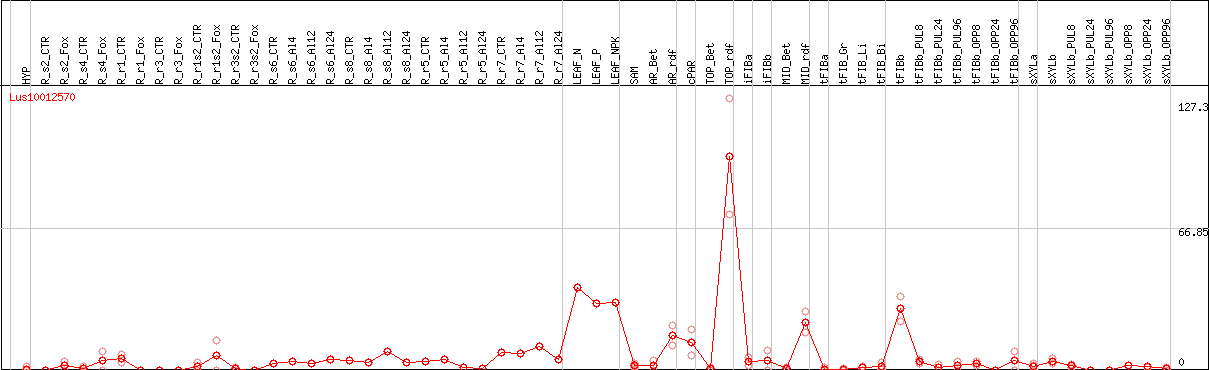
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10012570 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.