External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G32720 125 / 7e-38
B5 #4, B5#4, ATCB5-B
ARABIDOPSIS CYTOCHROME B5 ISOFORM B, cytochrome B5 isoform B (.1)
AT5G53560 116 / 3e-34
B5#2, ATB5-A, ATCB5-E
ARABIDOPSIS CYTOCHROME B5 ISOFORM E, cytochrome B5 isoform E (.1)
AT2G46650 113 / 4e-33
B5 #1, B5#1, ATCB5-C
ARABIDOPSIS CYTOCHROME B5 ISOFORM C, cytochrome B5 isoform C (.1)
AT5G48810 111 / 3e-32
B5#3, ATB5-B, ATCB5-D
ARABIDOPSIS CYTOCHROME B5 ISOFORM D, cytochrome B5 isoform D (.1)
AT1G26340 107 / 8e-31
B5 #6, B5#6, ATCB5-A
ARABIDOPSIS CYTOCHROME B5 ISOFORM A, cytochrome B5 isoform A (.1)
AT1G37130 69 / 1e-14
NIA2-1, CHL3, B29, NR2, NIA2, ATNR2
CHLORATE RESISTANT 3, ARABIDOPSIS NITRATE REDUCTASE 2, nitrate reductase 2 (.1)
AT1G77760 69 / 3e-14
GNR1, NIA1
nitrate reductase 1 (.1)
AT1G60660 60 / 2e-12
B5 #5, B5#5, ATCB5LP
ARABIDOPSIS CYTOCHROME B5-LIKE PROTEIN, cytochrome B5-like protein (.1)
AT3G61580 59 / 5e-11
AtSLD1
sphingoid LCB desaturase 1, Fatty acid/sphingolipid desaturase (.1)
AT5G09680 57 / 1e-10
RLF
reduced lateral root formation (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10010111
232 / 3e-80
AT2G32720 137 / 8e-43
ARABIDOPSIS CYTOCHROME B5 ISOFORM B, cytochrome B5 isoform B (.1)
Lus10005992
153 / 6e-49
AT5G53560 144 / 2e-45
ARABIDOPSIS CYTOCHROME B5 ISOFORM E, cytochrome B5 isoform E (.1)
Lus10030219
151 / 5e-48
AT5G53560 151 / 4e-48
ARABIDOPSIS CYTOCHROME B5 ISOFORM E, cytochrome B5 isoform E (.1)
Lus10008838
126 / 3e-38
AT5G53560 232 / 3e-80
ARABIDOPSIS CYTOCHROME B5 ISOFORM E, cytochrome B5 isoform E (.1)
Lus10022357
124 / 1e-37
AT5G53560 231 / 2e-79
ARABIDOPSIS CYTOCHROME B5 ISOFORM E, cytochrome B5 isoform E (.1)
Lus10043137
108 / 3e-31
AT2G32720 218 / 2e-74
ARABIDOPSIS CYTOCHROME B5 ISOFORM B, cytochrome B5 isoform B (.1)
Lus10032612
108 / 3e-31
AT2G32720 218 / 2e-74
ARABIDOPSIS CYTOCHROME B5 ISOFORM B, cytochrome B5 isoform B (.1)
Lus10011858
107 / 1e-30
AT2G32720 211 / 5e-72
ARABIDOPSIS CYTOCHROME B5 ISOFORM B, cytochrome B5 isoform B (.1)
Lus10022794
107 / 1e-30
AT2G32720 211 / 1e-71
ARABIDOPSIS CYTOCHROME B5 ISOFORM B, cytochrome B5 isoform B (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G024600
116 / 3e-34
AT5G53560 163 / 5e-53
ARABIDOPSIS CYTOCHROME B5 ISOFORM E, cytochrome B5 isoform E (.1)
Potri.015G007600
114 / 1e-33
AT5G53560 153 / 8e-49
ARABIDOPSIS CYTOCHROME B5 ISOFORM E, cytochrome B5 isoform E (.1)
Potri.001G314200
113 / 3e-33
AT2G32720 199 / 3e-67
ARABIDOPSIS CYTOCHROME B5 ISOFORM B, cytochrome B5 isoform B (.1)
Potri.002G242500
111 / 1e-32
AT2G32720 221 / 7e-76
ARABIDOPSIS CYTOCHROME B5 ISOFORM B, cytochrome B5 isoform B (.1)
Potri.017G054300
111 / 2e-32
AT2G32720 221 / 6e-76
ARABIDOPSIS CYTOCHROME B5 ISOFORM B, cytochrome B5 isoform B (.1)
Potri.010G156900
92 / 9e-25
AT1G26340 166 / 4e-54
ARABIDOPSIS CYTOCHROME B5 ISOFORM A, cytochrome B5 isoform A (.1)
Potri.014G019200
78 / 3e-19
AT5G48810 115 / 8e-34
ARABIDOPSIS CYTOCHROME B5 ISOFORM D, cytochrome B5 isoform D (.1)
Potri.002G088600
62 / 6e-12
AT1G77760 1394 / 0.0
nitrate reductase 1 (.1)
Potri.005G172400
61 / 2e-11
AT1G37130 1400 / 0.0
CHLORATE RESISTANT 3, ARABIDOPSIS NITRATE REDUCTASE 2, nitrate reductase 2 (.1)
Potri.018G087600
56 / 7e-11
AT1G60660 155 / 4e-50
ARABIDOPSIS CYTOCHROME B5-LIKE PROTEIN, cytochrome B5-like protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00173
Cyt-b5
Cytochrome b5-like Heme/Steroid binding domain
Representative CDS sequence
>Lus10012615 pacid=23161357 polypeptide=Lus10012615 locus=Lus10012615.g ID=Lus10012615.BGIv1.0 annot-version=v1.0
ATGGCTTCAAAGCCCAGAATTCACGAGTTCGATGATTTGCGGAAGCACAGTCAACGGGACGATTGCTGGATTGTTATTCACGGCAAGGTGTACAATGTGA
CAGACTTCCTAAATGAACATCCGGGAGGAGTAGAGGTTCTTCTCTCAGCTACTGAGAAAGATGCAAGCGACGATTTCGAGGATGTAGGACACGGGAAGGC
GGCAAGGGAGCTGATGAAGAAATACTACATAGGGGACATTGATCCTAACACCTTACCCTCTGGGAGGAAGTACAATCCAACACCTGTCTTCAATGGCAAG
AAGCCAGAGAAGAGTAACTTCCTGTTGATGAAGCTTATTCTGCCTCTCCTTATCTTCGCCATTGCTTACGCTACTCATAACGCTCTCAAGCACGACTAG
AA sequence
>Lus10012615 pacid=23161357 polypeptide=Lus10012615 locus=Lus10012615.g ID=Lus10012615.BGIv1.0 annot-version=v1.0
MASKPRIHEFDDLRKHSQRDDCWIVIHGKVYNVTDFLNEHPGGVEVLLSATEKDASDDFEDVGHGKAARELMKKYYIGDIDPNTLPSGRKYNPTPVFNGK
KPEKSNFLLMKLILPLLIFAIAYATHNALKHD
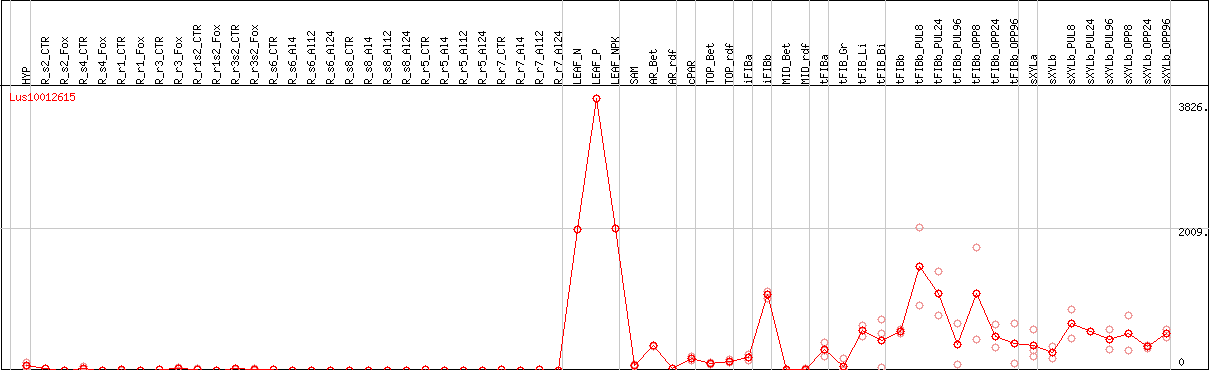
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10012615 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.