External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G39860 96 / 2e-27
bHLH
BNQ1, BHLH136, PRE1
PACLOBUTRAZOL RESISTANCE1, BANQUO 1, BASIC HELIX-LOOP-HELIX PROTEIN 136, basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
AT3G28857 96 / 4e-27
bHLH
PRE5
Paclobutrazol Resistance 5, basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
AT1G26945 94 / 1e-26
bHLH
KDR, PRE6
KIDARI, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G15160 90 / 5e-25
bHLH
BNQ2, BHLH134, PRE2
BASIC HELIX-LOOP-HELIX PROTEIN 134, BANQUO 2 (.1)
AT1G74500 83 / 5e-22
bHLH
TMO7, PRE3, ATBS1, bHLH135
TARGET OF MONOPTEROS 7, activation-tagged BRI1(brassinosteroid-insensitive 1)-suppressor 1 (.1)
AT3G47710 79 / 9e-21
bHLH
BNQ3, PRE4, bHLH161
BASIC HELIX-LOOP-HELIX PROTEIN 161, BANQUO 3 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10030490
142 / 2e-45
AT3G28857 102 / 4e-30
Paclobutrazol Resistance 5, basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Lus10037198
117 / 2e-35
AT1G26945 114 / 1e-34
KIDARI, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10036732
116 / 2e-35
AT1G26945 115 / 6e-35
KIDARI, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10024241
79 / 1e-20
AT3G47710 123 / 2e-38
BASIC HELIX-LOOP-HELIX PROTEIN 161, BANQUO 3 (.1)
Lus10023610
76 / 3e-19
AT3G28857 109 / 9e-33
Paclobutrazol Resistance 5, basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Lus10008626
66 / 2e-15
AT3G28857 71 / 1e-17
Paclobutrazol Resistance 5, basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G081300
101 / 2e-29
AT5G39860 126 / 1e-39
PACLOBUTRAZOL RESISTANCE1, BANQUO 1, BASIC HELIX-LOOP-HELIX PROTEIN 136, basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Potri.004G128400
95 / 5e-27
AT5G39860 124 / 8e-39
PACLOBUTRAZOL RESISTANCE1, BANQUO 1, BASIC HELIX-LOOP-HELIX PROTEIN 136, basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Potri.015G063300
89 / 2e-24
AT1G74500 75 / 3e-19
TARGET OF MONOPTEROS 7, activation-tagged BRI1(brassinosteroid-insensitive 1)-suppressor 1 (.1)
Potri.012G069500
85 / 5e-23
AT1G74500 81 / 2e-21
TARGET OF MONOPTEROS 7, activation-tagged BRI1(brassinosteroid-insensitive 1)-suppressor 1 (.1)
PFAM info
Representative CDS sequence
>Lus10012842 pacid=23163007 polypeptide=Lus10012842 locus=Lus10012842.g ID=Lus10012842.BGIv1.0 annot-version=v1.0
ATGTCAAGTATTAGAAGATCATCTTCGTCCCGGTCGAGACACACCACGACAAGGAGATCTAGTGCCATCTCTGATGATCAGATCATTGACCTCATTTCAA
AACTTCAACGACTCGTCCCAGAGGCAAACTCCACCTCTCACTGCTCTCGTAATAAGGGTTCGGCATCCAAGGTGTTGCAGGAGACATGCAACTACATAAG
GGAGTTGAACAGGCAGGTGGACGATCTGAGCGGAAGGCTGTCGGAGCTATTGGCCTCGATAGATTCCGACAGCGACCAAGCCGCCATTATTAGGAGCTTG
CTCATGTAG
AA sequence
>Lus10012842 pacid=23163007 polypeptide=Lus10012842 locus=Lus10012842.g ID=Lus10012842.BGIv1.0 annot-version=v1.0
MSSIRRSSSSRSRHTTTRRSSAISDDQIIDLISKLQRLVPEANSTSHCSRNKGSASKVLQETCNYIRELNRQVDDLSGRLSELLASIDSDSDQAAIIRSL
LM
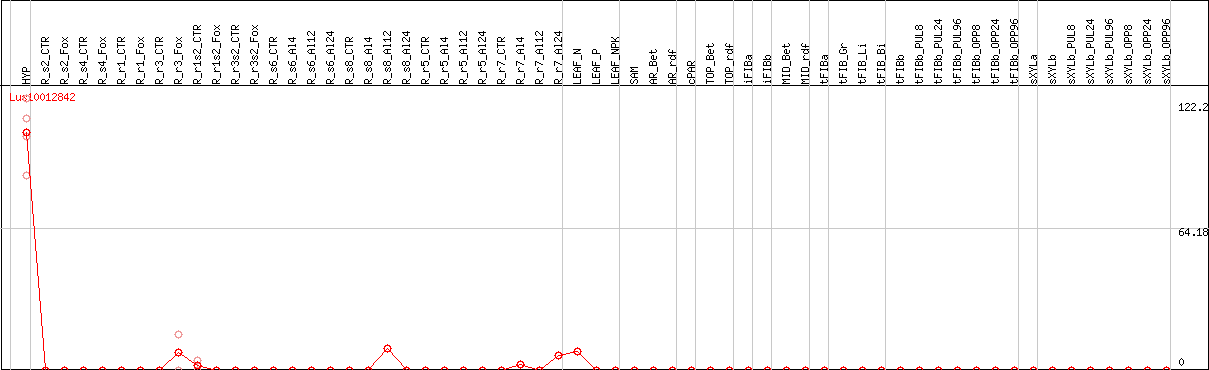
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10012842 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.