External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G56400 135 / 2e-37
WRKY
ATWRKY70, WRKY70
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 70, WRKY DNA-binding protein 70 (.1)
AT2G40750 116 / 5e-30
WRKY
ATWRKY54, WRKY54
WRKY DNA-binding protein 54 (.1)
AT4G23810 92 / 2e-21
WRKY
ATWRKY53, WRKY53
WRKY family transcription factor (.1)
AT2G40740 89 / 4e-20
WRKY
ATWRKY55, WRKY55
WRKY DNA-binding protein 55 (.1.2)
AT2G46400 87 / 2e-19
WRKY
ATWRKY46, WRKY46
WRKY DNA-binding protein 46 (.1)
AT5G24110 85 / 8e-19
WRKY
ATWRKY30, WRKY30
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 30, WRKY DNA-binding protein 30 (.1)
AT4G11070 81 / 3e-17
WRKY
ATWRKY41, WRKY41
WRKY family transcription factor (.1.2)
AT5G45260 80 / 2e-16
WRKY
ATWRKY52, SLH1, RRS1
SENSITIVE TO LOW HUMIDITY 1, RESISTANT TO RALSTONIA SOLANACEARUM 1, ARABIDOPSIS THALIANA WRKY DOMAIN PROTEIN 52, Disease resistance protein (TIR-NBS-LRR class) (.1), Disease resistance protein (TIR-NBS-LRR class) (.2)
AT1G66600 74 / 5e-15
WRKY
ABO3, ATWRKY63, WRKY63
WRKY DNA-binding protein 63, ABA overly sensitive mutant 3 (.1)
AT5G01900 71 / 3e-14
WRKY
ATWRKY62, WRKY62
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 62, WRKY DNA-binding protein 62 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10030517
448 / 8e-160
AT3G56400 162 / 5e-48
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 70, WRKY DNA-binding protein 70 (.1)
Lus10034244
219 / 4e-70
AT3G56400 151 / 6e-44
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 70, WRKY DNA-binding protein 70 (.1)
Lus10029021
207 / 2e-65
AT3G56400 149 / 3e-43
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 70, WRKY DNA-binding protein 70 (.1)
Lus10025216
108 / 4e-27
AT3G56400 123 / 8e-33
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 70, WRKY DNA-binding protein 70 (.1)
Lus10022736
106 / 2e-26
AT5G01900 105 / 1e-26
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 62, WRKY DNA-binding protein 62 (.1)
Lus10014177
102 / 7e-25
AT3G56400 105 / 3e-26
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 70, WRKY DNA-binding protein 70 (.1)
Lus10025133
101 / 1e-24
AT3G56400 114 / 2e-29
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 70, WRKY DNA-binding protein 70 (.1)
Lus10001062
93 / 3e-21
AT4G11070 169 / 5e-50
WRKY family transcription factor (.1.2)
Lus10039331
92 / 7e-21
AT4G11070 174 / 1e-51
WRKY family transcription factor (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0274
WRKY-GCM1
PF03106
WRKY
WRKY DNA -binding domain
Representative CDS sequence
>Lus10012870 pacid=23162984 polypeptide=Lus10012870 locus=Lus10012870.g ID=Lus10012870.BGIv1.0 annot-version=v1.0
ATGGGTATGCAGGGGAACAACAAGCTGCGGCAGGAGATGGCGATGAAGGAGCTTGCTCAAGGCCATGGATTTGCATCCCAGCTCCAGCTCCTCCTCAAGA
ACAATAAAGACCATCCTGCAGCTGCAGATGAACTTGTCAACAAGATCTTGGAATCTTTCACTTGTGGTATCTCTGCTCTGATGAGTACCGCTGCTTCGAC
TTCCATGGTGGCCATTAGCGATCAGGCAGGGGAGCAGCAGTCCTCTGTTTCCGGGATGTACGAGGATTCCGGCGAGAGTTGCCGGAAAAGGTCGTCGGCG
GCGGCGGCGGGGAAGAGTGGCGGAAGGGACAATAGAGGGTCCTATAAGAGAAGAAAGAGTGGACACTCTGATGTAAAAATCTCATCGTCGATGCACGACG
AACATTCGTGGAGGAAGTATGGCCAGAAGGAGATCCTCAATGCCAAATTCCCTAGGAGCTACTATAGATGTACACACAAGTACGACCAAGGCTGCAAGGC
CACCAAGCAAGTCCAGATGCTTGAACAAGAAACCAAAGAGACCATGTACTCCATAACTTACATGGGGCTCCACACTTGTCGGAACGCCACCGGGGTGTCT
CGATCGTCGGTGGAGCCTCTTTCAACGGCGGTGAATGTGAAGAAAGAGATTCGTACCATGGTCAATGTAGATGACATCAATAACAATGTGAACAACGAAG
AGAACCCGAGTGAAGTGATGACCGAGATTAATGATAATGTGGACTTGTGGATGCAGGGTTTGGAGTCATTTGATCAAGCAGAGGAGGCAATCAGTAACTA
TGATTACCATCATTATAGTAATTATGGGTTCTGTGGGGATCAAAGTGGGCAAGTTTTGGATTTTGATTTCGGGGTTAAGGATATTGATGATTTCGAGACC
GACTTTCGGTTCGATTATCACTAG
AA sequence
>Lus10012870 pacid=23162984 polypeptide=Lus10012870 locus=Lus10012870.g ID=Lus10012870.BGIv1.0 annot-version=v1.0
MGMQGNNKLRQEMAMKELAQGHGFASQLQLLLKNNKDHPAAADELVNKILESFTCGISALMSTAASTSMVAISDQAGEQQSSVSGMYEDSGESCRKRSSA
AAAGKSGGRDNRGSYKRRKSGHSDVKISSSMHDEHSWRKYGQKEILNAKFPRSYYRCTHKYDQGCKATKQVQMLEQETKETMYSITYMGLHTCRNATGVS
RSSVEPLSTAVNVKKEIRTMVNVDDINNNVNNEENPSEVMTEINDNVDLWMQGLESFDQAEEAISNYDYHHYSNYGFCGDQSGQVLDFDFGVKDIDDFET
DFRFDYH
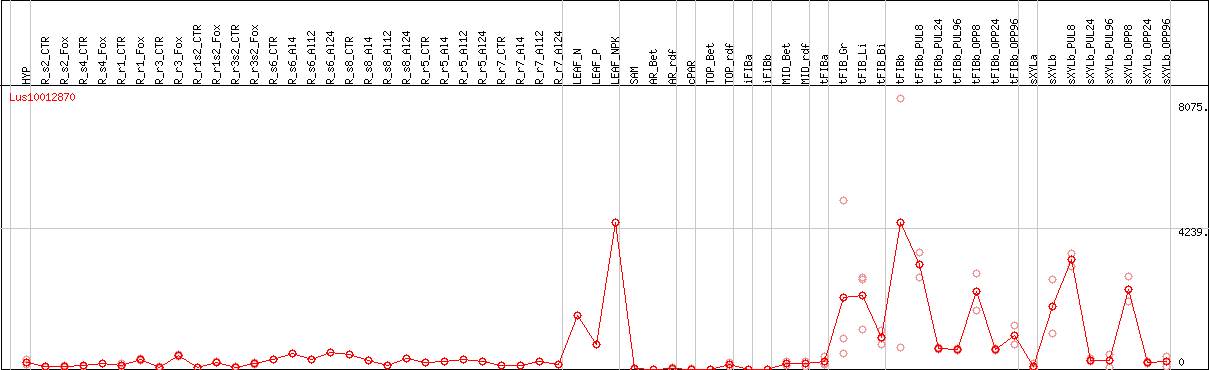
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10012870 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.