Lus10012931 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10012931 pacid=23158525 polypeptide=Lus10012931 locus=Lus10012931.g ID=Lus10012931.BGIv1.0 annot-version=v1.0
ATGTCTAGGATGTCAGGCCGTGATGTCCCGATGCCGGGGGGCGGCGGGCCATCATCATCGGCGGCCAACAATTCTTGGGTGTCGCCAACTTCAGTCTCGA
CGTCAGGGAAGAGGATACAGAGAGAAATAACAGAATTCAACATGGATCCACCGCCTGATTGTTCTGCAGGTCCAAAGGGTGACAATCTCTACCATTGGAT
CTCCACAATTATTGGACCTTCAGGAAAGCCATATGAAGGAGGTATATTTTTCCTTGACATAATATTTCCGAGTGACTATCCATTCAAACCGCCAAAGATG
GTGTTCAAGACTCGGATTTACCATTGTAATGTTGATTCTGCTGGCAACCTAATTATAGACATGTTGAAAGATAGCTGGAGTCCGGCTCTTACAATCACTA
AAGTGTTACTTGCAACAATTCTGAAATCCCTGGGATTGCACGCTTGTATTTGGCAGATAGACCCAAACATGACGAGTTGGCTTCAGAGTGGACTCGGAGG
TTTGCGAATCCCGAACAAAGCCAAAGCCGTTGCAATCAAATTCAAACACAATCTGAGTAAACGGAATTCAGAATCACTAGGGGAAGTGGAGATGATAAAG
AATGGGATATCTGAAATAGGGTTCCAGGACATAATCAGTGCAGGTCTTTCCCTCGAGGAGGCGACAAGACTCCACAGCAAGCTCAAAGATGCCATCTTTT
TATCAAATGGGCTTAGCTCTAAACCTACAGACTCGAGAGAGTTATGGCTGAGGGTGGTGGTCGGAAAAGTGCTCCAGCCATGGCATCCCCACAAGCTTCA
TCAGCTCGTTTACTATTCATGTTATGCTAATTGGGATTCCTCAACTCTTGGACCTCCTCCTTATTGGTTTCCTTCTCTGGATGAGTCAAAGTGCGCAAAC
TTGGGACGTAACTTATGGATCCAGGCTTTTTGGGACTTCTTACAAGGACCCCATTACAAGTTTTGGTCTCTTCAAAAATTTAGTGTTAAATGCCCTGAGG
TAGAAGCCTCGTATGTTCAGAAATCTGCTAGGTTGGTTGGCAATGCTCTAGATGCAACATTTCTGAAAGGGGAGACCATTGCCATTGACATGCCCATGAC
AGTTTATTCTGTTATCATCTACTTGGCCATTATATTAGCTGGATATGTTGTAGTATCAATAGCTGACAGCTTTGCCACGAAAGAAATTGCATCTCGTTTG
CGCATCTCCAATGCAAAGGGTATCTTCACTCAGGACTTCATAATTAGAGGAGGTCGGAAATTTCCGTTGTACAGCTGCGAAAGCAGGATCTTTCATGGAA
CGACTTTCTTTCCGGTGCCAACCAACTGGAAAGGTAATATATTACTTATTGCTCAACCAGCCGATTCAATGGCGAACATTCTCTTTTCGTCTGGGACTAG
AGGTGAAGAAATTGGAGACCCTAAAGCTCTCGTATGGATGCATCTATCACCAATTCGAGCCAGTAGCGGTGCTTCGGCTCACATTGATCTTGGGGTTCGT
GATGTCTACTGTTGGCCTACAAATTTAGGTTGGGTCATGGGACCCATTTTGCTCTATTCTTGCTTTTTAACTGGTGCAACTCTTGCGCTCTTTCATGGAA
CTCCTCTTGGCAGCGAGTGTGGGAAATTTGTTCAGCTAAAAGTTAGCGTCTTAGGAACTGTTCCAAGCTTAGTGAAGACTTGGAAGAATTCCAACTGCAT
GGATGGATTAAACTGGACAAAGATAAGAACTTGCATCATCCTACATCCAAGGCAGCCTGCTACAACCACAAGCTTTTGGATCATTTACACTGCATCGATG
ACCACAGGTTTAGTTATCATTGATGAAGAAGGGACTCCATATCCAGAAGACGAACCTTGTGTAGGTGAAGTGGGATTGTTTCCTTTGTACCTAGGTGCAT
CGGATCGACTTCTGAATGCTGATCACCATCAAGTTTACTTTGAGGGAATGCCAATTGTCAAGGGAATGCAACTAAGGAGGCATGCAGACATCCTGAAGAG
GACAGTTGGAGGCTACTTCATCATGCAGGGAAGGGCAGATGACACCATGAATCTCGGAGGGATTAAAACTAGTTCAGTAGAGATAGAGCGGGTATGTGAT
CGAGCTGACGAAAGCATTGTGGAGACAGCGGCAGTGAGTGTGGCACCAGTGGGTGGTGGTCCAGAAGTGTTGGTGATATTTGTGGTATTGAAGAAAGGGT
ATAAAGTGGAAGCAGAGCAGTTGAGAGTAATGTTCTCAAAGGCAATCCAGATCAATCTCAACCCACTATTCAAGGCAGAGTTGGAGTCATCCTTGGATTT
GCATCTTGCTAAATTGTTGGAGTGTCTTGCCTTGTTTGCAAGTAAGACACACATAGGATCAGATGCTAATAACCAAAGAAGAAAATCACTCATCTTAGCG
CAACCCCAAGAAGTTGCGTTAGGGAGAGACAACCAATTCCAGGGGAAAATGGAGATGATGAGAAATGGGATATCTGAAATAGAGGTCCAGGACATAATCA
GTGCAGGTCTTTCCCTTGAGGAGGCGACGAGGCTCCACGACAAGCTCAAAGATGCCATCTTTTCAACAAATGGGTTCAGTAGCTCTGAACCTGCGGACTC
GAGAGATTTATGGCAGAGGGTGGTGGTCGGAAAAGTGCTCCAGCCATGGCATCCCCATAAGCTCCACCAGCTGGTTTACTATTCTTGTTATGATAATTGG
GATTCCTCAGCTCGTGGACCTCCTCCTTATTGGTTTCCTTCTCTAGATGAGTCAATGCGTTCAAACTTGGGACTTGCAATGGAAACTTATGGATCCAAGT
TTTTTGGGACTTCTTACAAGGACCCCATTACAAGTTTTGGGCTCTTTCAAAAATTTAGTGTTAAATACCCTGAGGTTTACTGGTCCATCGTGCTACAACA
GCTTTCAGTGGCTTTCCATGAAAAGCCAAAATGCATTCTTGACCCTAGTGACGAGTCGAAACCTGGTGGAGCTTGGCTACCTGGTGCGGTCATGAATATT
GCTAATTGTTGTTTGCAGCCAGCAAGTCATCCTATGAAAAATGACAGCAGTATCGCAATTGTATGGAGGGATGAAACTGATTCCTCTAACATTAACCAAA
TGACATTGAAAGAACTCCGACAACAAGTAATGTTGGTTGCAAATGCTCTAGATGCAACATTTGTTAAAGGGGACACCATTGCCATTGACATGCCGATGAC
AGTTCATTCTGTTATCATCTACTTGGCCATTGTATTGGCTGGATATGTTGTAGTATCAATAGCTGACAGCTTTGCTGCTAAAGAAATTGCATCTCGTTTG
CGCATCTCCAATGCAAAGGGTATCTTCACTCAGGACTTCATAGTTAGAGGAGGTCGGAAGTTTCCATTGTACAGTCGTGTTACCGAAGCAAAGCCACATA
AAGCTATTGTGTTGCCTGCCGTTGGAAATGGTGTTGGAATTCAGTTGCGAAAGCAGGATCTTTCGTGGGACGACTTTCTTTCGAGTGCCAACCATCTGGA
AAGACCAAATGTTTACTCTCCTGTTTATCAACCAGCTGATTCAATGACAAACATTCTGTTTTCGTCCGGAACTACAGGAGACCCTAAAGCTATCGTATGG
ACGCATCTTTCACCAATTCGATGCAGTAGTGATGCTTGGGCTAACATTGATCTTCGGGTTGGCGATGTCTACTGTTGGCCTACAAATTTAGGTTGGGTAA
TGGGACCCATTTTGCTCTATTCTTGCTTCTTAACTGGTGCAGCTCTTGCTCTCTTTCATGGATCTCCCCTTGGCCGCGAGTTTGGAAAATTTGTTCAGGA
TGCCGGTGTTAGCGTATTAGGAACTGTTCCAAGCTTAGTGAAGACTTGGAAGAATTCCAACTGCATGGATAGATTAAACTGGACAAAGATAAGGTGCTTT
TGTTCCACTGGGGAAACTTCAAACGTTGATGATGACCTTTGGCTCTCTTCGAAATCATATTACAAACCCATTATTGAGTGTTGTGGAGGAACGGAACTTG
CATCATCCTACATCCAAGGCAGCCTGCTACAGCCACAGGCTTTTGGCTCATTTAGCACTGTGGCGATGACCACAGGTTTAGTCATCCTTGATGAAGAAGG
GATTCCATATCCAGATGATGAACCTTGTGTAGGTGAAGTGGGATTGTTTCCTTTGTACCTAGGTGCATCGGATCGACTTCTGAATGCTGATCACCATCAA
GTTTACTTCGAGGGAATGCCCATTGTCAATGGAATGCAACTAAGGAGACATGGTGACATTCTGAAGAGGACCGTTGGAGGATACTTCATTGTGCAGGGCC
GGGCAGATGACACCATGAATCTCGGAGGGATTAAGACTAGTTCAGTGGAGATAGAGCGTGTATGTGATCGAGCTGACGAAAGCATTGTGGAGACTGCAGC
AGTGAGTGTGGCACCAGTGGGTGGGGGCCCAGAAGTGTTGGTGATATTTGTGGTAGTGAAGAAAGAGTATAAAGGGGAAGCAGAGCAGTTGAGAGCCAAG
TTCTCAAAGGCAATCCAGATCAATCTCAACCCACTGTTCAAAGTTGGGCATGTTAAGATTGTAGAAGAGTTTCCCAGAACGGCCTCCAATAAGGTGCTAA
GGCGAGTTCTGAGGGACCAAATGAAGCAGCAGCTTCATGGACCAAGCAAGATATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10012931 pacid=23158525 polypeptide=Lus10012931 locus=Lus10012931.g ID=Lus10012931.BGIv1.0 annot-version=v1.0
MSRMSGRDVPMPGGGGPSSSAANNSWVSPTSVSTSGKRIQREITEFNMDPPPDCSAGPKGDNLYHWISTIIGPSGKPYEGGIFFLDIIFPSDYPFKPPKM
VFKTRIYHCNVDSAGNLIIDMLKDSWSPALTITKVLLATILKSLGLHACIWQIDPNMTSWLQSGLGGLRIPNKAKAVAIKFKHNLSKRNSESLGEVEMIK
NGISEIGFQDIISAGLSLEEATRLHSKLKDAIFLSNGLSSKPTDSRELWLRVVVGKVLQPWHPHKLHQLVYYSCYANWDSSTLGPPPYWFPSLDESKCAN
LGRNLWIQAFWDFLQGPHYKFWSLQKFSVKCPEVEASYVQKSARLVGNALDATFLKGETIAIDMPMTVYSVIIYLAIILAGYVVVSIADSFATKEIASRL
RISNAKGIFTQDFIIRGGRKFPLYSCESRIFHGTTFFPVPTNWKGNILLIAQPADSMANILFSSGTRGEEIGDPKALVWMHLSPIRASSGASAHIDLGVR
DVYCWPTNLGWVMGPILLYSCFLTGATLALFHGTPLGSECGKFVQLKVSVLGTVPSLVKTWKNSNCMDGLNWTKIRTCIILHPRQPATTTSFWIIYTASM
TTGLVIIDEEGTPYPEDEPCVGEVGLFPLYLGASDRLLNADHHQVYFEGMPIVKGMQLRRHADILKRTVGGYFIMQGRADDTMNLGGIKTSSVEIERVCD
RADESIVETAAVSVAPVGGGPEVLVIFVVLKKGYKVEAEQLRVMFSKAIQINLNPLFKAELESSLDLHLAKLLECLALFASKTHIGSDANNQRRKSLILA
QPQEVALGRDNQFQGKMEMMRNGISEIEVQDIISAGLSLEEATRLHDKLKDAIFSTNGFSSSEPADSRDLWQRVVVGKVLQPWHPHKLHQLVYYSCYDNW
DSSARGPPPYWFPSLDESMRSNLGLAMETYGSKFFGTSYKDPITSFGLFQKFSVKYPEVYWSIVLQQLSVAFHEKPKCILDPSDESKPGGAWLPGAVMNI
ANCCLQPASHPMKNDSSIAIVWRDETDSSNINQMTLKELRQQVMLVANALDATFVKGDTIAIDMPMTVHSVIIYLAIVLAGYVVVSIADSFAAKEIASRL
RISNAKGIFTQDFIVRGGRKFPLYSRVTEAKPHKAIVLPAVGNGVGIQLRKQDLSWDDFLSSANHLERPNVYSPVYQPADSMTNILFSSGTTGDPKAIVW
THLSPIRCSSDAWANIDLRVGDVYCWPTNLGWVMGPILLYSCFLTGAALALFHGSPLGREFGKFVQDAGVSVLGTVPSLVKTWKNSNCMDRLNWTKIRCF
CSTGETSNVDDDLWLSSKSYYKPIIECCGGTELASSYIQGSLLQPQAFGSFSTVAMTTGLVILDEEGIPYPDDEPCVGEVGLFPLYLGASDRLLNADHHQ
VYFEGMPIVNGMQLRRHGDILKRTVGGYFIVQGRADDTMNLGGIKTSSVEIERVCDRADESIVETAAVSVAPVGGGPEVLVIFVVVKKEYKGEAEQLRAK
FSKAIQINLNPLFKVGHVKIVEEFPRTASNKVLRRVLRDQMKQQLHGPSKI
|
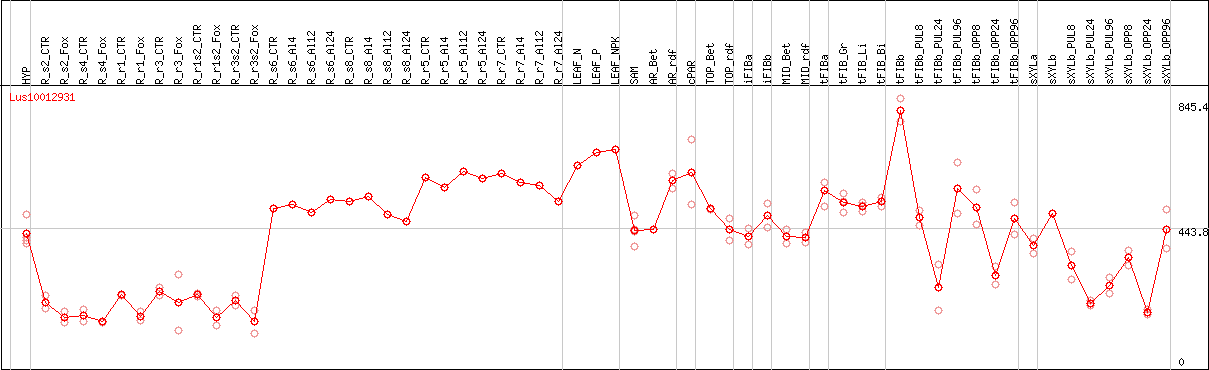
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10012931 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
