External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G51070 206 / 2e-61
SAG15, CLPD, ERD1
SENESCENCE ASSOCIATED GENE 15, EARLY RESPONSIVE TO DEHYDRATION 1, Clp ATPase (.1)
AT5G50920 194 / 3e-57
CLPC1, CLPC, ATHSP93-V, HSP93-V, DCA1
HEAT SHOCK PROTEIN 93-V, DE-REGULATED CAO ACCUMULATION 1, CLPC homologue 1 (.1)
AT3G48870 189 / 3e-55
ClpC2, ATCLPC, ATHSP93-III, HSP93-III
ClpC2, Clp ATPase (.1.2)
AT1G74310 182 / 5e-53
HOT1, ATHSP101
heat shock protein 101 (.1)
AT2G25140 172 / 3e-49
HSP98.7, CLPB-M, CLPB4
HEAT SHOCK PROTEIN 98.7, CASEIN LYTIC PROTEINASE B-M, casein lytic proteinase B4 (.1)
AT5G15450 167 / 1e-47
AtCLPB3, APG6, CLPB3, CLPB-P
CASEIN LYTIC PROTEINASE B-P, ALBINO AND PALE GREEN 6, casein lytic proteinase B3 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10038272
194 / 2e-59
AT2G19740 200 / 9e-63
Ribosomal protein L31e family protein (.1)
Lus10043013
200 / 4e-59
AT5G51070 1288 / 0.0
SENESCENCE ASSOCIATED GENE 15, EARLY RESPONSIVE TO DEHYDRATION 1, Clp ATPase (.1)
Lus10032514
198 / 3e-58
AT5G51070 1280 / 0.0
SENESCENCE ASSOCIATED GENE 15, EARLY RESPONSIVE TO DEHYDRATION 1, Clp ATPase (.1)
Lus10016723
195 / 2e-57
AT5G50920 1526 / 0.0
HEAT SHOCK PROTEIN 93-V, DE-REGULATED CAO ACCUMULATION 1, CLPC homologue 1 (.1)
Lus10032544
192 / 3e-57
AT5G50920 1271 / 0.0
HEAT SHOCK PROTEIN 93-V, DE-REGULATED CAO ACCUMULATION 1, CLPC homologue 1 (.1)
Lus10036017
192 / 2e-56
AT5G50920 1542 / 0.0
HEAT SHOCK PROTEIN 93-V, DE-REGULATED CAO ACCUMULATION 1, CLPC homologue 1 (.1)
Lus10040527
184 / 3e-55
AT1G74310 906 / 0.0
heat shock protein 101 (.1)
Lus10021159
184 / 7e-54
AT1G74310 1421 / 0.0
heat shock protein 101 (.1)
Lus10040528
184 / 1e-53
AT1G74310 1496 / 0.0
heat shock protein 101 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G105900
198 / 1e-58
AT5G50920 1505 / 0.0
HEAT SHOCK PROTEIN 93-V, DE-REGULATED CAO ACCUMULATION 1, CLPC homologue 1 (.1)
Potri.015G105100
197 / 3e-58
AT5G50920 1508 / 0.0
HEAT SHOCK PROTEIN 93-V, DE-REGULATED CAO ACCUMULATION 1, CLPC homologue 1 (.1)
Potri.015G110000
194 / 4e-57
AT5G51070 1282 / 0.0
SENESCENCE ASSOCIATED GENE 15, EARLY RESPONSIVE TO DEHYDRATION 1, Clp ATPase (.1)
Potri.012G112000
193 / 1e-56
AT5G51070 1328 / 0.0
SENESCENCE ASSOCIATED GENE 15, EARLY RESPONSIVE TO DEHYDRATION 1, Clp ATPase (.1)
Potri.015G057000
189 / 2e-55
AT1G74310 1567 / 0.0
heat shock protein 101 (.1)
Potri.015G056900
188 / 6e-55
AT1G74310 1566 / 0.0
heat shock protein 101 (.1)
Potri.005G197400
179 / 2e-53
AT5G50920 583 / 0.0
HEAT SHOCK PROTEIN 93-V, DE-REGULATED CAO ACCUMULATION 1, CLPC homologue 1 (.1)
Potri.004G124800
178 / 2e-51
AT5G15450 1567 / 0.0
CASEIN LYTIC PROTEINASE B-P, ALBINO AND PALE GREEN 6, casein lytic proteinase B3 (.1)
Potri.006G262700
173 / 1e-49
AT2G25140 1361 / 0.0
HEAT SHOCK PROTEIN 98.7, CASEIN LYTIC PROTEINASE B-M, casein lytic proteinase B4 (.1)
Potri.017G090600
169 / 3e-48
AT5G15450 1598 / 0.0
CASEIN LYTIC PROTEINASE B-P, ALBINO AND PALE GREEN 6, casein lytic proteinase B3 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF07724
AAA_2
AAA domain (Cdc48 subfamily)
CL0671
AAA_lid
PF10431
ClpB_D2-small
C-terminal, D2-small domain, of ClpB protein
Representative CDS sequence
>Lus10012953 pacid=23158566 polypeptide=Lus10012953 locus=Lus10012953.g ID=Lus10012953.BGIv1.0 annot-version=v1.0
ATGATCGAGTATATGGAGAGGCACAGCGTATCCAAACTCTTCGGGTCTCCACCTGGCTACACAAGTTTCATAGAGGGCGGCCAATTGACGGAGGCAATCG
CCCGGAAACCCAGCTCCGTCGTCCTTTTCGACGAAATCGAAAAGGCACACTGCGACGTGTGGAACACACTCCTCCAAATGTTCATGACGGACGGGGGTGA
GAATAAATTCAATTTCAGAGACGCCATCATCATCATGACCTCGAACGTCTGCCAGACAGCCGTGATCGAGGGGATTCCTAACGGTGATGAAACAGCAACA
AGGCATAAAGTTATGGAGGAGCTGAAGAAGGTGTTTAGACCGGAGTTTCTGAACAGAATTGATGACATTGTGATTTTCAAGGCGCTGTCGCAAGAGGAAT
TAATGAAGATGGTTGAGATCATGGTTAGGAAGTTGGCGGAGAGGATGAAGTCGACCAAGGGGATTCAACTGGAAATCAGCGAGGCATTGAAGAAGAAAAT
TGCGACCGAAGAGTACGATCCGTGCTACGGAGCCAGGCCATTGAAGAGGACGATGACGCGACTCCCATTGGCCGACGCAATTTTGGATGGTAGAGTTGGA
GATGGCGGCTCTATTACTCTGTTGGCCATGTCGGCGCCGACGACGACGGTCACAAAAGGTCACAGTCCATGA
AA sequence
>Lus10012953 pacid=23158566 polypeptide=Lus10012953 locus=Lus10012953.g ID=Lus10012953.BGIv1.0 annot-version=v1.0
MIEYMERHSVSKLFGSPPGYTSFIEGGQLTEAIARKPSSVVLFDEIEKAHCDVWNTLLQMFMTDGGENKFNFRDAIIIMTSNVCQTAVIEGIPNGDETAT
RHKVMEELKKVFRPEFLNRIDDIVIFKALSQEELMKMVEIMVRKLAERMKSTKGIQLEISEALKKKIATEEYDPCYGARPLKRTMTRLPLADAILDGRVG
DGGSITLLAMSAPTTTVTKGHSP
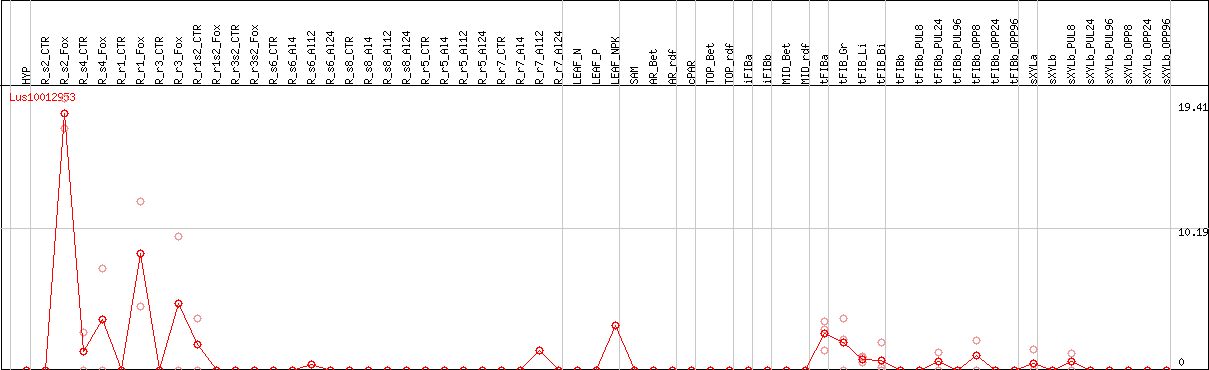
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10012953 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.