External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G23360 369 / 3e-130
MENG
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2.3)
AT5G57300 109 / 2e-28
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2.3)
AT2G41040 61 / 1e-10
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT5G54400 60 / 2e-10
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT3G15530 57 / 3e-09
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
AT3G18000 57 / 4e-09
XPL1, NMT1, XIPOTL1, PEAMT
XIPOTL 1, N-METHYLTRANSFERASE 1, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT1G73600 53 / 6e-08
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
AT1G48600 53 / 7e-08
AtPMEAMT
phosphoethanolamine N-methyltransferase, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
AT1G64970 49 / 1e-06
VTE4, TMT1, G-TMT
VITAMIN E DEFICIENT 4, gamma-tocopherol methyltransferase (.1)
AT3G63410 47 / 6e-06
VTE3, APG1, IEP37, E37
VITAMIN E DEFECTIVE 3, INNER ENVELOPE PROTEIN 37, ALBINO OR PALE GREEN MUTANT 1, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10029124
432 / 6e-155
AT1G23360 298 / 8e-102
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2.3)
Lus10011441
110 / 1e-28
AT5G57300 475 / 2e-171
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2.3)
Lus10000179
83 / 4e-19
AT5G57300 342 / 3e-120
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2.3)
Lus10037553
84 / 7e-19
AT5G57300 359 / 1e-125
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2.3)
Lus10037890
62 / 5e-11
AT5G54400 449 / 2e-160
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10038599
62 / 6e-11
AT5G54400 455 / 8e-163
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10018010
51 / 4e-07
AT3G18000 843 / 0.0
XIPOTL 1, N-METHYLTRANSFERASE 1, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10008815
50 / 6e-07
AT2G41040 451 / 9e-160
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10009056
50 / 9e-07
AT1G73600 773 / 0.0
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G188600
415 / 2e-148
AT1G23360 354 / 2e-124
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2.3)
Potri.018G088400
109 / 2e-28
AT5G57300 470 / 4e-169
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2.3)
Potri.010G043601
58 / 3e-11
AT1G23360 63 / 9e-14
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2.3)
Potri.011G128300
62 / 7e-11
AT5G54400 450 / 1e-160
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.001G405200
58 / 1e-09
AT5G54400 396 / 1e-139
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.012G047400
54 / 5e-08
AT1G48600 862 / 0.0
phosphoethanolamine N-methyltransferase, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Potri.006G060400
53 / 6e-08
AT2G41040 461 / 5e-164
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.018G120000
53 / 7e-08
AT2G41040 418 / 6e-147
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.015G039000
52 / 1e-07
AT1G48600 862 / 0.0
phosphoethanolamine N-methyltransferase, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Potri.005G245800
51 / 3e-07
AT1G20330 667 / 0.0
FRILL1, COTYLEDON VASCULAR PATTERN 1, sterol methyltransferase 2 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF01209
Ubie_methyltran
ubiE/COQ5 methyltransferase family
Representative CDS sequence
>Lus10013038 pacid=23149859 polypeptide=Lus10013038 locus=Lus10013038.g ID=Lus10013038.BGIv1.0 annot-version=v1.0
ATGGCTTCTCTTCAATTCGCATTCTCCTCATTCGCCGTCCGGCCAATCCCCCACCTCCGACGCCCAACGATACGATGCTCCGACGACCGGCAGGCACTCT
TTAACCGCATCGCTCCCGTCTACGACAATCTGAATGATTTGCTGAGTTTCGGGCAGCATCGAGTCTGGAAACGGATGGCCATTTCCTGGACTGGAGCTAA
GACTGGGGATTCTGTTCTTGATATATGCTGTGGAAGTGGGGACTTGGCTTTTCTCTTGTCCGAGAAAGTGGGTTCTGCTGGCAAGGTAATAGGACTTGAT
TTCTCCAAAGAACAGCTGGTTGTTGCTTCATCCCGGCAGAACTCGCGGTGGAAGTCCTGCTATCATAGCATAGAATGGTTAGAAGGCGATGCTACGGATT
TGCCATTCTCCGATCGTCAGTTTGATGCCATCACCATGGGGTATGGGCTGAGAAATGTGGTTGACAAAGACAAGGCGATGCGGGAGATGTTCCGAGTGCT
GAAACCAGGAGGGAAAGCTTCAATTCTCGACTTCAACAAGACAACCCAACCATTTGTTGCCTCCTTTCAGGGGTGGATGATCGACAATGTTGTTGTTCCC
GTGGCTACTGGCTATGGTCTTGCAGACGAGTACGAATACTTGAAGACCTCCATTAGAGAATTCTTGACTGGGAAGGAGCTAGAAGAACTCGCCATGGACG
CTGGTTTCTCCAACGCCGCGTATTATGAGATCAGTGGGGGGTTGATGGGGAATCTAGTTGCAACTCGTTAG
AA sequence
>Lus10013038 pacid=23149859 polypeptide=Lus10013038 locus=Lus10013038.g ID=Lus10013038.BGIv1.0 annot-version=v1.0
MASLQFAFSSFAVRPIPHLRRPTIRCSDDRQALFNRIAPVYDNLNDLLSFGQHRVWKRMAISWTGAKTGDSVLDICCGSGDLAFLLSEKVGSAGKVIGLD
FSKEQLVVASSRQNSRWKSCYHSIEWLEGDATDLPFSDRQFDAITMGYGLRNVVDKDKAMREMFRVLKPGGKASILDFNKTTQPFVASFQGWMIDNVVVP
VATGYGLADEYEYLKTSIREFLTGKELEELAMDAGFSNAAYYEISGGLMGNLVATR
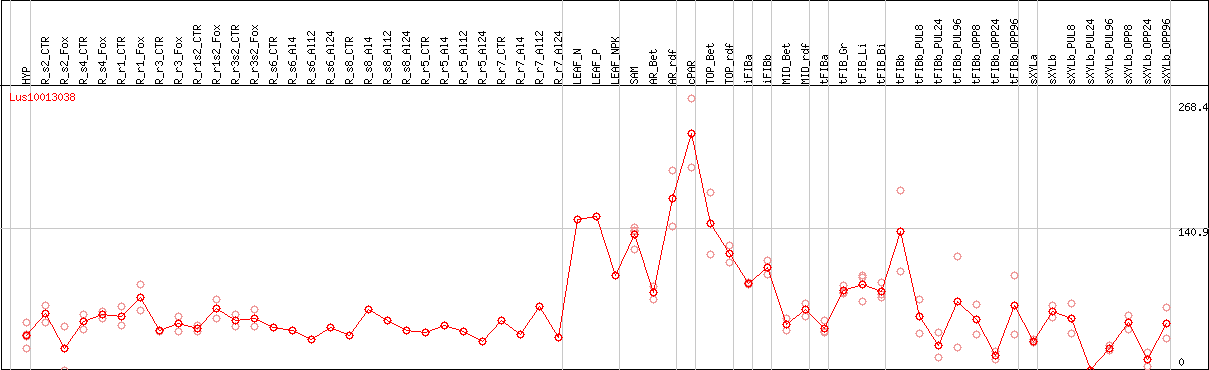
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013038 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.