Lus10013044 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10013044 pacid=23149849 polypeptide=Lus10013044 locus=Lus10013044.g ID=Lus10013044.BGIv1.0 annot-version=v1.0
ATGAGTAATAGCATAAGTCATAATTTATTCTACCAGAGTTTGCTTCATCCTAAGGTTGTCGACCACAGAAGCAAATCCAAAGCTTCTGGCGTTGTTGCAA
GCTCCTCGTTTCAATCTGCTTCTGTCAGCCAAGCTACATTGCCTACGCTTGGGGTTACTACCCCGTCGTCTAGTTTCTATGGAGGAAGATTGAAAACAAA
GAAATCAAAGGTGTCTGCTGGGAGAAGTCGTCCTGTTGCAGTCACACCGCGAGCTGTACTGGCGATGGAACAACCATCCGAGCTTGCTGGTAAATTTAAA
CTCGATCACAATGCAGAGTTGCAGGTTTTCATTAGCAATTCTCCTACTGCACCGGTTACTACTGTAAATATTCAGTTGACATATACTAGTGACTCGTTGA
TCCTCCACTGGGGTGGGATACATGATAAAAGAGAGAAATGGGTGCTTCCTGCTCGTTGCCCAGATGGGACGAAAAATCATAAGAACAGGGCTCTTAGAAC
GCCTTTCTCAAAGGCTGGTTCAAGTTCTTTCCTCAAACTAGAGATCGATGATCCTAAGATCCAAGCTATTGAGTTCCTCATATTTGATGAAGCCAAAAAC
CAATGGTTTAAAAATAGTGGCCAGAACTTTCATGTGAAGTTCTCCCCTAGAGAGAAGTTTATCAGTTCAAATGTTTCAGTTCCTGAAGAGCTTGTACAAG
TCCAGGCATATCTCAGGTGGGAAAGAAAAGGAAAACAAATGTATACGCCTCAGCAAGAGAAGGCAACTGCTCGTGCTGAGTTATTGGAAGAAGTAGCCAG
AGGCTCTTCTGTAGAAGACCTCCGAGCAAAGCTGAATAGTAAAAATGATGGAACTTCAATTAAGGAGTCCCCAGTCTCTGAAGTAAAGAGCAACATACCT
GAGGATCTTGTGCAAGTGCAATCATATTTACGATGGGAAAGAGCAGGGAAACCCGATTACTCCACGGAACAGCAACTGCGAGAATTCGAGGAGGCGAGAG
CAGAGTTGCAAGCTGAGCTTGACAGGGGTGTGTCTCTGGATGAGATCAGAAAGAAAATCACTAAAGGCAATACTCATACAAATGCTGGTGTTAATCCAAA
GAAGGAAAACTACGTCAGCACTGAAAGGATTCAGCGCAAAAGTAGGGATTTGACACAAATTGTCACTAAATATAAAGCTGACTCAACAATGGAAGTAGTT
CCTGCGGAACCAAAATCTCCAAAGCCAGTTGAGCTATATGCCAATGCTATTGAAGAACAAGCTGGTGACACTATCCTCAACAAGAAAAAATTTAAGCTTG
GGGATAAAGAGATTCTGGTTCTTGTAAGCAAGCCTGGCGAGAAAATCAAGGTTTATGTTGTCACTGATCTTGAAGAATCAGCTATTCTTCATTGGGCTTT
AGCTAAAAGTCCTGGAGAGTGGTCGGCACCACCTGCAAGCGTCTTGCCTCCAGATTCAGTTCCTCTGAAGGAAGCTGCTGAAACACCACTCACGAAACCG
TCTACCGAACTTCCTTATCAGGTCCAGTCGTTTCAAATAGAGATCGAAGATAGTAGTTTCGTAGGAATGCCTTTCGTTCTTTTGTCTAACGGCAATTGGA
TGAAGAATAATGGCTCAGACTTCTACGTTGAATTTTCCAGCCCAACTAAGAAAGTGCAAAAGGATGCTGGTGATGGGAGAGGTACGGCAAAGACATTGTT
GGACAGAATTGCAGAGATGGAAGGGGAGGCACAGAAGTCTTTCATGCACCGGTTCAATATTGCGTCTGATTTAATGCATCAAGCCAAGGATGCTGGAGTG
CTGGGTATTGCGGCTATCTTCGTGTGGAAGAGGTTTATGGCGACCCGACAACTTATATGGAACAAGAACTACAATGTCAAACCACGTGAGATCAGTCAAG
CCCAGGATAGGCTTACTGACATGCTCCAAGATATTTATTCTAGCTATCCACAGTACCGAGAGTTTTTGCGGTTGATTCTGTCAACTGTTGGTAGAGGAGG
TGAAGGAGATGTGGGGCAGCGTATACGAGATGAAATACTGGTTATTCAGAGAAACAATGATTGCAAAGGTGGAATGATGGAAGAATGGCACCAAAAATTA
CACAATAACACTAGCCCCGATGATGTTATAATCTGCCAGGCTTTGATTGATCACATAAGTAGCGACTTCGACATGAGTGTCTACTGGAAAACTTTGAATG
ACAATGGCATTACAAAAGAGCGACTTTTAAGTTATGATCGTGCCATACATAATGAACCGAATTTTAGGAGGGATCAAAAGGATGGCCTTTTGCGCGATCT
TGGAAGCTATATGAGGACATTGAAGGCAGTGCATTCAGGTGCAGATCTTGAGTCTGCTATTGCAAATTGCATGGGCTATAAAGCAGAGGGGCAAGGTTTT
ATGGTGGGAGTGAATATTAATCCAGTTTCAGGCTTGCCTTCTGGATTTCCGGAATTGCTTCAATTTGTCCTGAAACATGTTGAGGACAGGAATGTACAAG
CACTTCTGGAGGGTTTACTTGAGGCTCGTCAGGAGCTTAGACCATCACTGTCCCAAGCTAACCCTCGCCTGAAGGATGTTTTGTTTTTGGACATTGCACT
TGATTCTGCAGTTAGGACAGCCATTGAAAGAGGGTATGAGGAATTAAACGATGCTGGCCCTGAGATTTCACAAGGCACTGACTGCCTACCCATTTTGCAG
AAATTTATGTATTTCATTTCTCTGGTAGTTGAGAATCTTGCACTTTCGTCAGATGATAATGAGGATCTTATCTACTGCTTAAAGGGATGGAACCAAGCCA
TGAAGATGTCCAAAAGAAATAACAACTGGGCATTATATGCAAAGTCGATCCTGGACAGAACTCGCCTTGCCCTTGCTGGCAAGGCCGAGTCATATCAGTA
TATTCTGCAGCCTTCAGCTGAGTATCTTGGATCACTGCTTGGAGTGGATCAGTGGGCTGTAGAAATTTTCACGGAAGAGGTCATCCGTGGGGGTTCAGCA
GCTGCTTTGTCGTCACTTCTTAATAGACTAGATCCAGTTCTACGTCAAACAGCTAATCTTGGAAGTTGGCAAGTTATCTGCCCAGCTGAAGCAGTTGGGT
ACGTTGTTGTTGTGGATGAGTTGCTCTCAGTACAGAACAAAACATATGAGCGGCCTACTATATTGATTGCAAAAAAAGTAAAAGGTGAAGAGGAAATACC
AGACGGTACAGTTGCCGTGCTAACACCAGATATGCCTGATGTCCTATCACATGTATCCGTCCGGGCAAGAAATGGCAAGGTTTGCTTTGCTACATGTTTT
GACCCTAACATTCTGGACGATCTCCAAGCATGCGAAGGGAAACTACTGCGTCTGAAACCTACTTCTGCAGAAGTAGTTTACAGTGAAGTAAAAGAGGATG
AGCTGGAAGGTCCTAGTTCAACTAGCTCAAAAGATGTTGATTCTTCATCTCTGAAGTTGGTCAAAAAGCAATTTACAGGAAAATATGCCATATCATCAGA
CGAGTTCACAGGCGAAATGGTTGGTGCCAAGTCCCGCAATATCTCGAATTTGAGGGGAAAAGTTCCATCATGGATCGGAATTCCAACATCAGTTGCCTTG
CCTTTTGGAGTTTTTGAGAAGGTTCTTGCCGATGGTTTAAATAAGGATGTTGAGAAGAAGCTACAGCTATTGAAGGAGAAGTTGGACGCAGGAGAATTCG
GTGTTCTTGGAGAGATTCGGGAGACTGTTTTAGACCTGTTACCTCCACCAGAGCTGGTGGCAGAGCTGAAATCCAAAATGCAGAGTTCTGAGATGCCTTG
GCCAGGTGATGAAGGTGAAAAACGCTGGGAACAAGCCTGGACGGCTATTAAGAAGGTCTGGGCTTCGAAATGGAATGAGAGAGCATACTTCAGCACAAAG
AAAATAAAGCTGGATCATGACTATCTCTGCATGGCAGTACTGGTGCAAGAAATCATAAACGCTGATTACGCGTTCGTCATCCATACCACCAATCCATCCT
CCGGAGACACGTCAGAGATATATGCAGAGGTGGTCAAAGGACTCGGAGAAACTCTCGTCGGAGCTTATCCTGGCCGTGCTTTGAGTTTCATCTGCAAGAA
AAATAACTTGGATTCTCCGCAGGTTCTGGGTTATCCCAGCAAACCAATCGGTCTCTTCATAAGACGTTCGATCATCTTTCGATCCGACTCCAATGGAGAA
GATCTAGAAGGCTATGCCGGTGCAGGCTTGTACGACAGTGTGCCAATGGACGAAGAGGAGAAAGTAGTGCTGGATTACTCGTCCGATCCACTGATGATCG
ACGAGAACTTCCAGAAATCGACACTCGCAAAGATCGCCAGTGCAGGGCACGCAATTGAAGAACTCTATGGATCTGCACAAGACATTGAAGGCGTGGTACG
AGATGGCAAAATCTACGTCGTTCAGACCAGACCCCAGGTGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10013044 pacid=23149849 polypeptide=Lus10013044 locus=Lus10013044.g ID=Lus10013044.BGIv1.0 annot-version=v1.0
MSNSISHNLFYQSLLHPKVVDHRSKSKASGVVASSSFQSASVSQATLPTLGVTTPSSSFYGGRLKTKKSKVSAGRSRPVAVTPRAVLAMEQPSELAGKFK
LDHNAELQVFISNSPTAPVTTVNIQLTYTSDSLILHWGGIHDKREKWVLPARCPDGTKNHKNRALRTPFSKAGSSSFLKLEIDDPKIQAIEFLIFDEAKN
QWFKNSGQNFHVKFSPREKFISSNVSVPEELVQVQAYLRWERKGKQMYTPQQEKATARAELLEEVARGSSVEDLRAKLNSKNDGTSIKESPVSEVKSNIP
EDLVQVQSYLRWERAGKPDYSTEQQLREFEEARAELQAELDRGVSLDEIRKKITKGNTHTNAGVNPKKENYVSTERIQRKSRDLTQIVTKYKADSTMEVV
PAEPKSPKPVELYANAIEEQAGDTILNKKKFKLGDKEILVLVSKPGEKIKVYVVTDLEESAILHWALAKSPGEWSAPPASVLPPDSVPLKEAAETPLTKP
STELPYQVQSFQIEIEDSSFVGMPFVLLSNGNWMKNNGSDFYVEFSSPTKKVQKDAGDGRGTAKTLLDRIAEMEGEAQKSFMHRFNIASDLMHQAKDAGV
LGIAAIFVWKRFMATRQLIWNKNYNVKPREISQAQDRLTDMLQDIYSSYPQYREFLRLILSTVGRGGEGDVGQRIRDEILVIQRNNDCKGGMMEEWHQKL
HNNTSPDDVIICQALIDHISSDFDMSVYWKTLNDNGITKERLLSYDRAIHNEPNFRRDQKDGLLRDLGSYMRTLKAVHSGADLESAIANCMGYKAEGQGF
MVGVNINPVSGLPSGFPELLQFVLKHVEDRNVQALLEGLLEARQELRPSLSQANPRLKDVLFLDIALDSAVRTAIERGYEELNDAGPEISQGTDCLPILQ
KFMYFISLVVENLALSSDDNEDLIYCLKGWNQAMKMSKRNNNWALYAKSILDRTRLALAGKAESYQYILQPSAEYLGSLLGVDQWAVEIFTEEVIRGGSA
AALSSLLNRLDPVLRQTANLGSWQVICPAEAVGYVVVVDELLSVQNKTYERPTILIAKKVKGEEEIPDGTVAVLTPDMPDVLSHVSVRARNGKVCFATCF
DPNILDDLQACEGKLLRLKPTSAEVVYSEVKEDELEGPSSTSSKDVDSSSLKLVKKQFTGKYAISSDEFTGEMVGAKSRNISNLRGKVPSWIGIPTSVAL
PFGVFEKVLADGLNKDVEKKLQLLKEKLDAGEFGVLGEIRETVLDLLPPPELVAELKSKMQSSEMPWPGDEGEKRWEQAWTAIKKVWASKWNERAYFSTK
KIKLDHDYLCMAVLVQEIINADYAFVIHTTNPSSGDTSEIYAEVVKGLGETLVGAYPGRALSFICKKNNLDSPQVLGYPSKPIGLFIRRSIIFRSDSNGE
DLEGYAGAGLYDSVPMDEEEKVVLDYSSDPLMIDENFQKSTLAKIASAGHAIEELYGSAQDIEGVVRDGKIYVVQTRPQV
|
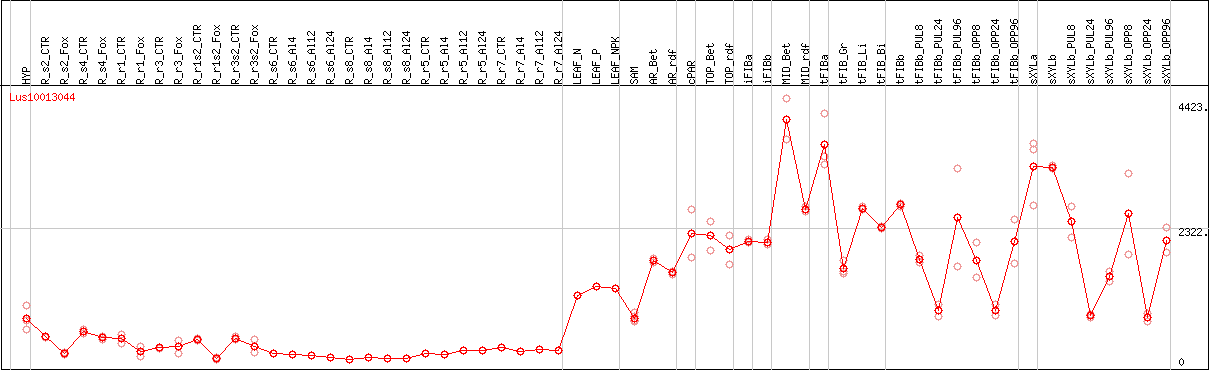
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013044 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
