External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G15550 253 / 8e-82
ATGA3OX1, GA4
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
AT4G21690 239 / 7e-77
ATGA3OX3
ARABIDOPSIS THALIANA GIBBERELLIN 3-OXIDASE 3, gibberellin 3-oxidase 3 (.1)
AT1G80330 239 / 1e-76
ATGA3OX4
ARABIDOPSIS THALIANA GIBBERELLIN 3-OXIDASE 4, gibberellin 3-oxidase 4 (.1)
AT1G80340 226 / 2e-71
ATGA3OX2, GA4H
ARABIDOPSIS THALIANA GIBBERELLIN-3-OXIDASE 2, gibberellin 3-oxidase 2 (.1)
AT1G78550 148 / 2e-41
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT3G13610 145 / 2e-40
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G55290 144 / 5e-40
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G62380 142 / 1e-39
ATACO2, ACO2
ACC oxidase 2 (.1)
AT1G12010 140 / 9e-39
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G60980 140 / 2e-38
ATGA20OX4
gibberellin 20-oxidase 4 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10008100
486 / 1e-173
AT4G21690 273 / 8e-90
ARABIDOPSIS THALIANA GIBBERELLIN 3-OXIDASE 3, gibberellin 3-oxidase 3 (.1)
Lus10013134
463 / 2e-164
AT4G21690 242 / 3e-77
ARABIDOPSIS THALIANA GIBBERELLIN 3-OXIDASE 3, gibberellin 3-oxidase 3 (.1)
Lus10011476
266 / 1e-86
AT1G15550 412 / 1e-143
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
Lus10008099
227 / 3e-73
AT4G21690 137 / 3e-38
ARABIDOPSIS THALIANA GIBBERELLIN 3-OXIDASE 3, gibberellin 3-oxidase 3 (.1)
Lus10013137
215 / 1e-69
AT4G21690 140 / 1e-40
ARABIDOPSIS THALIANA GIBBERELLIN 3-OXIDASE 3, gibberellin 3-oxidase 3 (.1)
Lus10013136
204 / 9e-66
AT1G15550 97 / 6e-25
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
Lus10022292
149 / 1e-41
AT1G17020 449 / 5e-159
senescence-related gene 1 (.1)
Lus10028068
143 / 6e-40
AT5G08640 429 / 1e-151
flavonol synthase 1 (.1.2)
Lus10025620
143 / 7e-40
AT5G08640 424 / 2e-149
flavonol synthase 1 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G033600
333 / 2e-113
AT1G15550 328 / 2e-111
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
Potri.006G247700
328 / 1e-111
AT1G15550 328 / 2e-111
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
Potri.001G176600
264 / 3e-86
AT1G15550 452 / 1e-159
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
Potri.003G057400
261 / 5e-85
AT1G15550 451 / 2e-159
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
Potri.009G022800
155 / 3e-44
AT1G17020 302 / 6e-101
senescence-related gene 1 (.1)
Potri.005G113900
145 / 3e-40
AT3G51240 598 / 0.0
TRANSPARENT TESTA 6, flavanone 3-hydroxylase (.1.2)
Potri.010G023600
143 / 2e-39
AT3G21420 511 / 0.0
LATERAL BRANCHING OXIDOREDUCTASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.019G014454
142 / 3e-39
AT5G08640 454 / 1e-161
flavonol synthase 1 (.1.2)
Potri.005G113700
141 / 9e-39
AT3G51240 590 / 0.0
TRANSPARENT TESTA 6, flavanone 3-hydroxylase (.1.2)
Potri.017G136000
139 / 5e-38
AT1G06620 314 / 2e-105
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0029
Cupin
PF03171
2OG-FeII_Oxy
2OG-Fe(II) oxygenase superfamily
Representative CDS sequence
>Lus10013135 pacid=23164173 polypeptide=Lus10013135 locus=Lus10013135.g ID=Lus10013135.BGIv1.0 annot-version=v1.0
ATGCCTACAACTACTACTACCTCCGAATCCCACCAAACAGTCCCTCTCGACTTCCACTCAATTCACACCCTCCCAGAGACCCACGTCTGGCCACCTGGCG
CCAACGATAAAGTGTTCAGGATTCCTGTCGTTGACTTCTCGGGTCCTTTTGAGAGTAGCCGGAAGTTGGTCGTTGAGGCGTGCGAGAAGTGGGGCGTGTT
CCAATTGACAAACCACGGGATTCCTTCCGAATTGTTGACGGAGGTTGAATCTCAGTCGAGACAGCTGTTTGCCCTCCCGGTGAGCCGGAAACTGAAGGCT
TTACGGGCCCCGAACGGAGTCAGCGGGTATGGCCTGCCTCCCATTTCGCAGTTCTTTGAGAAGCGTATGTGGAATGAAGGGTTTACCATTGCTGGTTCTC
CAAGGGATCATGTTGCGAAACTTTGGCCGGATGATCAATCCCGAGTGTTTTGCGATGTCATCGAAGACTTTCAACGGCGAATGACGGAGCTCGCCGCCAA
AATTTTCCGACTAATCCTCGAGTCCCTCAACACCTCGGAGCAAGAAATCCGGCGTATCTCATCGCCGGAAAGCATCATAACCAGCGTACAAATGAACTCA
TATCCGCCATGCCCCGACCCGAATCGGGCAATGGGTATGGTGGCCCACACCGACACTTCGTTGCTCACCCTCCTATACCAAGGCAGCGTCAAGGGTCTCC
AAATCTTCAACGGCGGGGAGTGGGGAATGGTACCTCCGGTTGACGGAGCATTGACGGTGAACGTCGGGGATTTCCTCCACGTCATTTCCAACAGCCAGTT
TCACCCGGTTCGGCACCGGGTGGTGATTAAGGAGGCCATGCTGCAAAGGGTTTCTGTTGCGTTGTTTTATTTTCCGAAGCCGGAGTTTGTTTTATCACCG
TCCGGGGTAGCCTCCGGCGACGTCGAAGCTCCAATGAACCATTCCTTGACTGTGAAGGAGTATAGAGATTTGAAGTACAAAAGCTATCAGACTGCACTTG
ATGCAATAAAGAAATAG
AA sequence
>Lus10013135 pacid=23164173 polypeptide=Lus10013135 locus=Lus10013135.g ID=Lus10013135.BGIv1.0 annot-version=v1.0
MPTTTTTSESHQTVPLDFHSIHTLPETHVWPPGANDKVFRIPVVDFSGPFESSRKLVVEACEKWGVFQLTNHGIPSELLTEVESQSRQLFALPVSRKLKA
LRAPNGVSGYGLPPISQFFEKRMWNEGFTIAGSPRDHVAKLWPDDQSRVFCDVIEDFQRRMTELAAKIFRLILESLNTSEQEIRRISSPESIITSVQMNS
YPPCPDPNRAMGMVAHTDTSLLTLLYQGSVKGLQIFNGGEWGMVPPVDGALTVNVGDFLHVISNSQFHPVRHRVVIKEAMLQRVSVALFYFPKPEFVLSP
SGVASGDVEAPMNHSLTVKEYRDLKYKSYQTALDAIKK
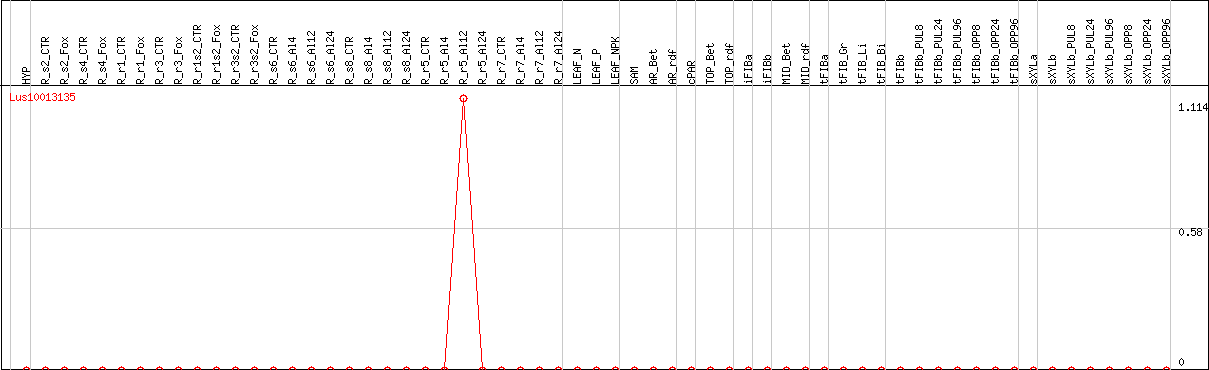
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013135 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.