External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G21690 139 / 2e-40
ATGA3OX3
ARABIDOPSIS THALIANA GIBBERELLIN 3-OXIDASE 3, gibberellin 3-oxidase 3 (.1)
AT1G15550 125 / 5e-35
ATGA3OX1, GA4
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
AT1G80340 124 / 1e-34
ATGA3OX2, GA4H
ARABIDOPSIS THALIANA GIBBERELLIN-3-OXIDASE 2, gibberellin 3-oxidase 2 (.1)
AT1G80330 107 / 4e-28
ATGA3OX4
ARABIDOPSIS THALIANA GIBBERELLIN 3-OXIDASE 4, gibberellin 3-oxidase 4 (.1)
AT4G25420 63 / 6e-12
AT2301, GA5, ATGA20OX1
GA REQUIRING 5, ARABIDOPSIS THALIANA GIBBERELLIN 20-OXIDASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G78440 63 / 6e-12
ATGA2OX1
Arabidopsis thaliana gibberellin 2-oxidase 1 (.1)
AT3G13610 61 / 2e-11
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT5G43935 61 / 3e-11
ATFLS6, FLS6
flavonol synthase 6 (.1)
AT1G17020 59 / 1e-10
ATSRG1, SRG1
senescence-related gene 1 (.1)
AT5G08640 58 / 2e-10
ATFLS1, FLS
flavonol synthase 1 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10008100
325 / 7e-113
AT4G21690 273 / 8e-90
ARABIDOPSIS THALIANA GIBBERELLIN 3-OXIDASE 3, gibberellin 3-oxidase 3 (.1)
Lus10013134
221 / 9e-72
AT4G21690 242 / 3e-77
ARABIDOPSIS THALIANA GIBBERELLIN 3-OXIDASE 3, gibberellin 3-oxidase 3 (.1)
Lus10013135
215 / 6e-70
AT1G15550 263 / 4e-86
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
Lus10011476
130 / 7e-37
AT1G15550 412 / 1e-143
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
Lus10023113
125 / 3e-36
AT1G15550 171 / 1e-51
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
Lus10008099
100 / 5e-26
AT4G21690 137 / 3e-38
ARABIDOPSIS THALIANA GIBBERELLIN 3-OXIDASE 3, gibberellin 3-oxidase 3 (.1)
Lus10023311
70 / 2e-14
AT1G30040 381 / 6e-133
GIBBERELLIN 2-OXIDASE 2, gibberellin 2-oxidase (.1.2)
Lus10025620
69 / 5e-14
AT5G08640 424 / 2e-149
flavonol synthase 1 (.1.2)
Lus10028068
64 / 2e-12
AT5G08640 429 / 1e-151
flavonol synthase 1 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G033600
182 / 5e-57
AT1G15550 328 / 2e-111
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
Potri.006G247700
181 / 1e-56
AT1G15550 328 / 2e-111
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
Potri.003G057400
135 / 6e-39
AT1G15550 451 / 2e-159
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
Potri.001G176600
135 / 2e-38
AT1G15550 452 / 1e-159
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
Potri.009G022800
72 / 4e-15
AT1G17020 302 / 6e-101
senescence-related gene 1 (.1)
Potri.011G095600
71 / 1e-14
AT1G78440 404 / 4e-141
Arabidopsis thaliana gibberellin 2-oxidase 1 (.1)
Potri.010G023550
66 / 1e-13
AT3G21420 210 / 6e-68
LATERAL BRANCHING OXIDOREDUCTASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G378400
67 / 2e-13
AT1G78440 404 / 1e-141
Arabidopsis thaliana gibberellin 2-oxidase 1 (.1)
Potri.004G139600
66 / 4e-13
AT5G08640 470 / 7e-168
flavonol synthase 1 (.1.2)
Potri.019G014454
66 / 4e-13
AT5G08640 454 / 1e-161
flavonol synthase 1 (.1.2)
PFAM info
Representative CDS sequence
>Lus10013137 pacid=23164188 polypeptide=Lus10013137 locus=Lus10013137.g ID=Lus10013137.BGIv1.0 annot-version=v1.0
ATGGTGAACGTTTCATCCAACACTATGACTACAAAGTCTACAGTCGTCTCCGACGCCTACGCACAATCCCCACTCAAATCTCACCACATCGTCCCGCTCG
ATTTCGACTCCATCCGAACCACCCTCCCGGAGACCCACGTCTGGCCTCCGGGAGCCGGTGACCAAGCCTTCCCTATTCCGGTCATCGACTTCTCGGAACC
TTCCTCCAAAAATTCGCTTATTGTCGAGGCGTGCGAGAAGTGGGGTGTGTTCCAATTGACGAACCACGAGATTCCTTTCGAACTTTTGGCTGAGGTCGAA
GCTCAGACGAGGCGGCTTTTCGCCCTCCCGGCCAGCCGGAAGGTCAAGGCTTTACGGTCTCCGAACGGGGTTAGTGGGTACGGTCAGGCTCGGATTTCTC
AGTTCTTTGAGAAGCATATGTGGAACGAAGGGTTCACTGTGGTGGGGTCTCCGGCGGATCATGCTGCTCAGCTTTGGCCGGACGATGATCGCCGCCGTCA
AGTGTTCTGGTGA
AA sequence
>Lus10013137 pacid=23164188 polypeptide=Lus10013137 locus=Lus10013137.g ID=Lus10013137.BGIv1.0 annot-version=v1.0
MVNVSSNTMTTKSTVVSDAYAQSPLKSHHIVPLDFDSIRTTLPETHVWPPGAGDQAFPIPVIDFSEPSSKNSLIVEACEKWGVFQLTNHEIPFELLAEVE
AQTRRLFALPASRKVKALRSPNGVSGYGQARISQFFEKHMWNEGFTVVGSPADHAAQLWPDDDRRRQVFW
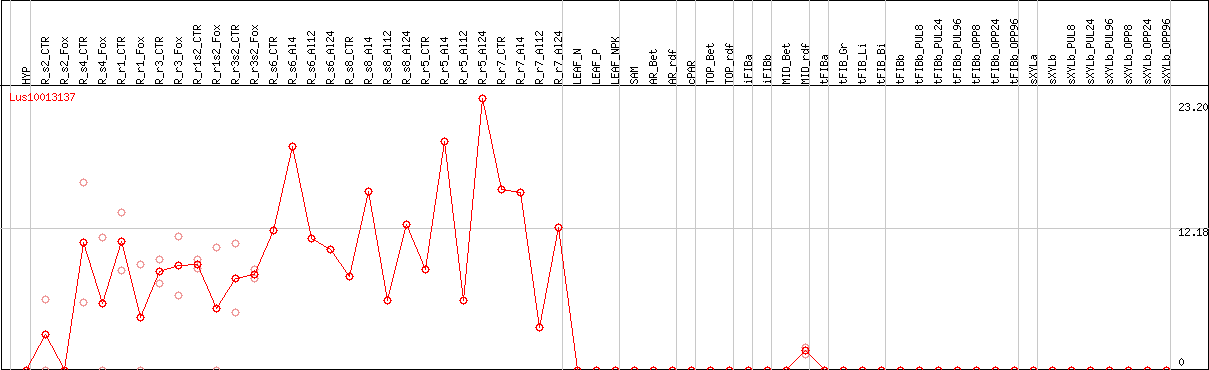
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013137 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.