External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G25220 367 / 9e-129
WEI7, TRP4, ASB1
WEAK ETHYLENE INSENSITIVE7, TRYPTOPHAN BIOSYNTHESIS 4, anthranilate synthase beta subunit 1 (.1.2)
AT5G57890 362 / 3e-127
Glutamine amidotransferase type 1 family protein (.1)
AT1G25155 358 / 2e-126
Glutamine amidotransferase type 1 family protein (.1)
AT1G24909 358 / 2e-126
Glutamine amidotransferase type 1 family protein (.1)
AT1G25083 358 / 2e-126
Glutamine amidotransferase type 1 family protein (.1)
AT1G24807 349 / 2e-122
Glutamine amidotransferase type 1 family protein (.1)
AT2G28880 101 / 6e-24
ADCS, EMB1997
embryo defective 1997, aminodeoxychorismate synthase, para-aminobenzoate (PABA) synthase family protein (.1)
AT3G27740 69 / 4e-13
VEN6, CARA
VENOSA 6, carbamoyl phosphate synthetase A (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10030761
509 / 0
AT1G25220 363 / 9e-127
WEAK ETHYLENE INSENSITIVE7, TRYPTOPHAN BIOSYNTHESIS 4, anthranilate synthase beta subunit 1 (.1.2)
Lus10036525
104 / 5e-25
AT2G28880 1026 / 0.0
embryo defective 1997, aminodeoxychorismate synthase, para-aminobenzoate (PABA) synthase family protein (.1)
Lus10041403
103 / 1e-24
AT2G28880 1078 / 0.0
embryo defective 1997, aminodeoxychorismate synthase, para-aminobenzoate (PABA) synthase family protein (.1)
Lus10018639
67 / 2e-12
AT3G27740 676 / 0.0
VENOSA 6, carbamoyl phosphate synthetase A (.1.2)
Lus10039874
66 / 8e-12
AT3G27740 678 / 0.0
VENOSA 6, carbamoyl phosphate synthetase A (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G138800
407 / 6e-145
AT1G25220 404 / 2e-143
WEAK ETHYLENE INSENSITIVE7, TRYPTOPHAN BIOSYNTHESIS 4, anthranilate synthase beta subunit 1 (.1.2)
Potri.010G102200
403 / 4e-143
AT1G25220 401 / 1e-142
WEAK ETHYLENE INSENSITIVE7, TRYPTOPHAN BIOSYNTHESIS 4, anthranilate synthase beta subunit 1 (.1.2)
Potri.010G221500
100 / 8e-24
AT2G28880 1016 / 0.0
embryo defective 1997, aminodeoxychorismate synthase, para-aminobenzoate (PABA) synthase family protein (.1)
Potri.003G080900
71 / 7e-14
AT3G27740 647 / 0.0
VENOSA 6, carbamoyl phosphate synthetase A (.1.2)
Potri.001G153400
70 / 2e-13
AT3G27740 676 / 0.0
VENOSA 6, carbamoyl phosphate synthetase A (.1.2)
Potri.003G128200
42 / 0.0003
AT1G63660 910 / 0.0
GMP synthase (glutamine-hydrolyzing), putative / glutamine amidotransferase, putative (.1), GMP synthase (glutamine-hydrolyzing), putative / glutamine amidotransferase, putative (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0014
Glutaminase_I
PF00117
GATase
Glutamine amidotransferase class-I
Representative CDS sequence
>Lus10013241 pacid=23166675 polypeptide=Lus10013241 locus=Lus10013241.g ID=Lus10013241.BGIv1.0 annot-version=v1.0
ATGGCTGCAAATTTGGTCTCACGAGCTTCGTTAATCCAGTCCAAACCCTCAATATCATTCAAAAACCAGCAGACTCATAGTCTCCTCCTCCCTCAGCAGC
TCTCTGTGCCGTCAAATTTTGGGACAAAGGGAAATGGATTGGCGGTGAGATGTTCGATTGCTGTCCCTGAAGCTCCTTCAAGGATTGGTGATTCAAATGG
GAAAGGGGCGAAATCCCCTGTCGTTGTGATTGACAATTATGACAGTTTCACTTACAATCTTTGTCAGTATATGGGAGAGTTGGGATGCGAATTTGAGGTC
TACCGGAATGATGAGTTAACCGTTGAAGAGTTGAAAAGGAAAAAGCCTAGAGGTGTGCTGATATCTCCAGGACCAGGGGCGCCCAAGGATTCTGGAATAT
CGTTGCAGACAGTTTTGGAGCTTGGACCAATTGTGCCCTTGTTTGGTGTGTGTATGGGTTTGCAGTGCATTGGCGAGGCATTTGGTGGAAAGATCGTGCG
TTCTCCATTTGGGGTCATGCATGGGAAAAGTTCACCGGTGTATTATGATGAGAAAGGAAAAGGCTTGTTCACTGGGTTACCGAACCCATTCAAAGCTGGT
AGATATCACAGCCTTGTAATTGACCGAGACACCTTCCTGATTGATGAACTAGAGATCACTGCATGGACTGAGGATGGGTTGATTATGGCTGCCCGTCACA
AGAAGTACAAATATCTACAGGGAGTGCAGTTTCATCCTGAAAGCATTATAACCTCCGAAGGAAAGATCATCGTCCAGAACTTCCTCAAAATGGTCGAGGA
GAAAGAGGCAGCCGACTCGAACAACAGCTGA
AA sequence
>Lus10013241 pacid=23166675 polypeptide=Lus10013241 locus=Lus10013241.g ID=Lus10013241.BGIv1.0 annot-version=v1.0
MAANLVSRASLIQSKPSISFKNQQTHSLLLPQQLSVPSNFGTKGNGLAVRCSIAVPEAPSRIGDSNGKGAKSPVVVIDNYDSFTYNLCQYMGELGCEFEV
YRNDELTVEELKRKKPRGVLISPGPGAPKDSGISLQTVLELGPIVPLFGVCMGLQCIGEAFGGKIVRSPFGVMHGKSSPVYYDEKGKGLFTGLPNPFKAG
RYHSLVIDRDTFLIDELEITAWTEDGLIMAARHKKYKYLQGVQFHPESIITSEGKIIVQNFLKMVEEKEAADSNNS
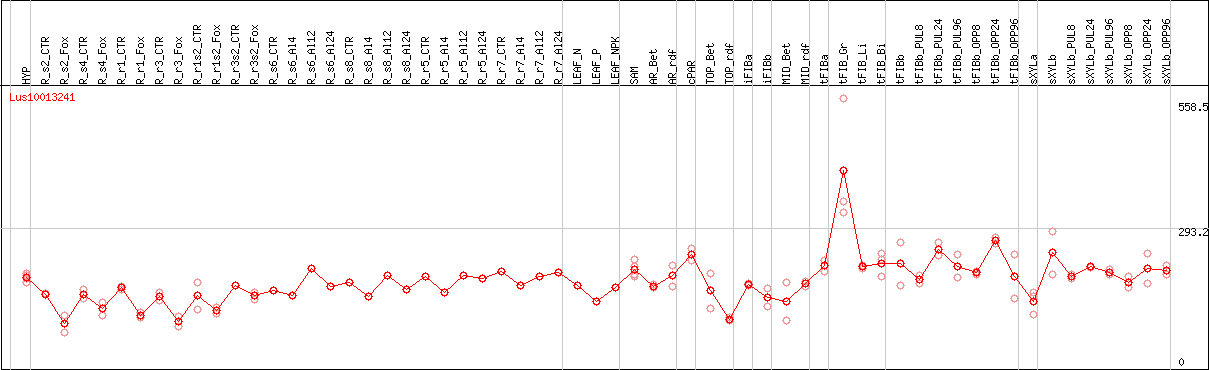
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013241 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.