Lus10013245 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10013245 pacid=23166684 polypeptide=Lus10013245 locus=Lus10013245.g ID=Lus10013245.BGIv1.0 annot-version=v1.0
ATGGATGCACCAAGCTTCATTCTGCTTCCTCTTATTCTTACTCTTCTCTTCCATTCTAAAATCTATGCATCCTCAATTAATGACACTTTACACGCTAACA
GATCAATCAAGGATGGGGAGCTTATCATCTCCAAAGGTGGCAACTTTGCTCTTGGATTTTTCAGCCCTGGTAGTTCAACCTACAGAAGATACCTTGGTAT
CTGGTTTAATAACAAGCCAGGCCAAACTGTGGTGTGGGTAGCTAACAGGAACACCCCCATCAACGGCACCTCAGGAATCCTCATTAACGATGGATATGGA
AACCTTTTGCTCTACAGCAGCATCCGAAACCAAGAGCTTCCCATATGGTCAACCAACGTTTCCAGAACTTTTCTATGTTCAGCTCAACTATTGGATTCAG
GAAATCTGGTGTTGGTCCAGCAAGGTACCAATATCAAAACATCACTACTTCTGTGGCAAAGCTTTGATCATCCTACGGATACCTTGTTACCTGGTATGAA
AATCGGCATCAATCTAGAAACAGGTTTGCACGTGTTCCTGACATCTTGGAGATCAGAAGATGATCCAGGAACCGGCGATTTCTCGTTGAAGCTCAATCCG
AATGGCTTGCCTCAAATCTTTTTGTACCAGGGTGTGAATCCCCGTTGGCAGAGTGTCCCATGGCCATGGAAAGTAGACCATCGTTTATTCATTGATTCTT
TTGTTAATAATGAGGTTGGGGTGTGGTTCTGGTTTGTTCCTCGTGATGCTTCGACTATGCACATGTCTGTGATTCTCAATTCAGGAACAGTGAGGGCCAT
GATGTGGGATGGAGCAGGTGGGATATGGAAGGAGTTCAATACGATTTCGCATCAGAAGTGTGACATATACGGCCATTGTGGTGCTTATGGAATGTGTGAC
TCTAGCTCCTCTTTTCTTTCTGAATGTATGTGCTTACCTGGTTATGAACCAAAGTCCCAAAGCAAGGTGCTTTTCCAAGATGGAAATGGCGGGTGTGTTA
GGAAGCGATCGGAATCCTCTGTTTGTGGGAAGGGAGAAGGTTTTGTGAGAGTTAGAAATGTGAAGATTCCTGATACATCAGAAGCAGTATGGATTGACCA
AGGGTTGAACAAAGTGAACTGTGAAGAAGAGTGTAGAAAGAATTGCTCCTGCTGTGCATATACTACTGCTAATATTGTTGGGAAAGGGACTGGTTGTTTG
ACTTGGTATGGAGAACTGATGGATACAAGACACCTCCCGATACCTGATCAAGAAATCTACGTTCGTGTCGATACTCTCGAACTAGCTGAGTATGCAGTGC
AATCTGCTGAGTTTGATGAGATTAGTTTTAAGCTAAATGTTCTCGCTCCATCAGTTTCTTCGGTTTTGATAGTCTGCATCTTGTTTGCCTACACACTGTT
CAAGAGGTGGAGAAAGTGGAAAGCAAGGAAAGCAAAAGAAAGGCGGATCAGAAACTTGTTTCGTCCGAGCGAAGATTCGAGTTACTTGCAGAATTCTATA
GTGGAAAAGGAAACGGAACGAAGTAAAGCGTACCCAGAATTACCATTTTTCAGCCTCGACGTGATACGTGCTGCAACCAACAACTTTTCATCTGATAACA
AACTTGGAGAAGGGGGTTTTGGTTCTGTGTACAAGGGGAAATTTCCATCCGGGGAAATGGTAGCTGTAAAAAGAGCGCCAAGAAATTCAAGGCAGGGGCT
AGAACAGTTTAAAAGTGAAGTGTTCTTAATTTCAAAATTGCAGCATAGGAACCTTGTGAAGCTCCACGGATGTTGCATTGAGGAAGTAGAGCAGATGTTG
ATTTATGAATACTTACCTCATAAGAGTTTGGATTCACTTCTTTTTGTTGAGCATCAGACTGATAGCCCATTCTTAGACTGGGAAAAGCGATTTAATATCA
TAGTTGGTATTGCTCGAGGGATCTTGTATCTTCATCAAGACTCGAGATTCAGAATAATCCACAGAGATCTAAAAATCAGCAATGTTCTCTTGGATGCGGA
GCTTAAACCTAAAATATCCGATTTTGGAATGGCCAGAGTACTGGAGGGAAACCAAGTTAAAGGGATGACTAGTAAAATTGGTGGAACATATTTCGGGGCG
ATGCTGCTGGAAATTGTGACTGGGAAGAAGATCAACAGTGGGTTTACTACTCACGAGGAGGATTGCGCTGCCTTAAACTTGATTGGATATGTATGGGAGC
TATGGAAAGCAGGGAGAGTGGGCGATATAGTGGATCTATTACTTCAAGTGTCGACGAGTACCCTGAGATGCATCCACATCGGGCTGTCATGTCTCGAGGA
AGATCCGGTAGATCGACCAGATATGCAGATAGTAGTTGTCATGTTGAACAGCGAAACAATCCCACTTCCTCCACTGAAACGTCCTGCATTCGTCTGTGGT
GCAAACTCTACAGTTGAGGGTGGACATCCTTTTCCAATTTTGATTAGCTTGGAGGGTAGAAAGAGCTCCAGTCAAAGACTAATTTCAACTAAATATCAAA
TGGATGCACAAAGGCTCATTCTGCTTCCGCTGCTTCTTACTCTTCTCTTCCATGATAAAGCCTCTGCATCATCAGATGACACTCTACATGCTAACAGATC
AATCAACGATGGCGAGCTTATCATCTCCAAAGGAGGTAACTTTGCTCTTGGGTTCTTCAGCCCTGGTAATTCAACCTACAGAAGATATCTTGGTATTTGG
TTTCATAACAAGCCAGGCCAAACTGTTGTGTGGGTGGCTAACAGGAACACCCCCATCAACGACACTTCAGGATTCCTCGTCAACGACGGATATGGAAACA
TTTTGCTCTACAGCAGCACCCGAAACCAAGAGGTCCCGATATGGTCTACTAACGTCTCGAGAGCCTTTCCTTGTTTGGCTCAACTGTTGGATTCAGGAAA
TCTAGTGTTGGTCCTACAAGGTAGTACATCAAGTAACAATACATCACTACTTGTGTGGCAAAGCTTTGATCATCCTACGGATACCTTTTTACCCGGTATG
AAACTCGGCATCAATCTAGAAACAGGTTTGGACATGTTCCTGACATCTTGGAGATCAGAAGACGATCCAGGAACCGGCGATTTCTCGTTTAAGCTCAACC
CGAATGGCTCGCCTCAGTTCTTTCTGTACCAGGGTATGAATCCCCGTTGGCGGAGTATCCCATGGCCATGGACAGTAGACCATCCTTTATACGTTGATTC
TTTTGTTAATAATGAAGTTGGGATGTGGTACTCGTATGCTCCTCGTGATGCTTCGACTGTGTTCATGTATGTGATCCATAATTCGGGAACAGTGAAAGCA
GTGGTATGGGACGAAGGCGGTGGGATATGGAAGGAGTTCTATACGATTTTGCATCAAAAGTGTGATATATACGGCTATTGTGGTGCTTACGGAAAGTGTG
ACCCTGGCTCCTCTTTTCTGTCAGAATGCATTTGCTTACCTGGTTATGAACATCCAATCTCCCAAAGCAAGGTGCTTTCCGAAGATGGAAATCGCGGGTG
TGTCAGGAAGCGATCAGAGTCCTCACTTTGTGCGAAGGGAGAGGGTTTTGTGAGAGTTAGAAATGTGAAGATTCCGGATGCATCAAAAGCAGTTTGGGTA
GACCAAGGGTTGAACGAAGTGAATTGTGAAGATGAGTGTAGAAAGGATTGCTCCTGCACTGCATATACTACTCCTAATATTGTTGCGAAAGGGACTGGTT
GTTTGACATGGCATGGAAAATTGATGGATACGGTAGACTTCCCGACACCTGATCAAGAAATCTATGTTCGTGTTGATGCTCTAGAACTAGCTGAGTATGC
AGTGCAATCTGATGAGCTTAGTTTCAAGCTAAGCGTTCTCGTTCCATCAGTTACTTCCGTTTGCATTGTCTGCATCTTGTTTGCCTACACACTGTTGAAG
AGGTGGAGAAAGCGGAAAGCAAGGAAAGCAACAGAAAGGCGGATCAGAAACTTGTTTCGTCCGAGCGAGGATTCGAGTTACTTGCAGGATTCTATAGTGA
CAAAGGAAATGGAGCGAAGTAAAATGTATCCAGAATTACCAGTTTTTAGCCTCGACGTGATACGTGCTGCAACTAACAACTTTTCGTCTGATAACAAACT
TGGACAAGGGGGTTTTGGCTCTGTGTACAAGGGGAAATTTCCAACTGGGGAAGAGATTGCTGTAAAAAGACCGTCAAAAAATTCAAGACAGGGGGTAGAA
CAGTTCAAAAGTGAAGTATTCTTGATCTCGAAATTGCAGCATAGGAACTTGGTGAAGCTCTATGGATGCTGCATTGAGAAAGAAGAGCAGATGTTGATTT
ATGAGTACTTACCTTATAAGAGTTTGGATTCACTTCTTTTCGTTGAGCATCAGCTTGGAAGCAACCTATTCTTAGATTGGGGAAAGCGGTTTAATATCAT
TATTGGTATTGCACGTGGGATCTTATATCTTCATCAAGATTCGAGGTTCAGAATAATCCACAGAGATTTAAAAATCAGCAACGTTCTCTTAGATGCGGAG
TTTAATCCCAAAATATCTGATTTTGGGATGGCTAGAATACTGGAGGGGAGCCAAGTCGAAGGGATGACTCGTAGAATTGGTGGAACATATGGCTATATGG
CTCCAGAGTATGCTTATTGTGGAAAGTTTTCAGTGAAATCGGACGTTTATAGTTTTGGAGTGATGTTGTTTGAAATTGTGACCGGGAAGAAGATCAAAAG
TGGGTTTGCTACTCACGAGGAGGATCATGCTGCCTTAAGCTTAATTGGATACGTATGGGAGCTATGGAAAGCAGAGAGAGTGGGGGATATAGTGGATGCA
TCACTTCAACTATCAATTAGTACCATGAGATGCATCCACATCGGGCTGTTATGTCTCGAGGAAGAACCAGTAGAGCGACCAGATATGCAGACAGTGGTTG
TGATGTTGAATAGCGAAACAACCCCGCTTCCTCTGCCGAAACGTCCTGCATTTGTCTGTGGAGCAAAGTCCGGAGTTGAGGTGAGGGTGGAAGAAGAATG
CTCAATAAACGAGTTTACATTTTCAGACACTCTTACTCGATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10013245 pacid=23166684 polypeptide=Lus10013245 locus=Lus10013245.g ID=Lus10013245.BGIv1.0 annot-version=v1.0
MDAPSFILLPLILTLLFHSKIYASSINDTLHANRSIKDGELIISKGGNFALGFFSPGSSTYRRYLGIWFNNKPGQTVVWVANRNTPINGTSGILINDGYG
NLLLYSSIRNQELPIWSTNVSRTFLCSAQLLDSGNLVLVQQGTNIKTSLLLWQSFDHPTDTLLPGMKIGINLETGLHVFLTSWRSEDDPGTGDFSLKLNP
NGLPQIFLYQGVNPRWQSVPWPWKVDHRLFIDSFVNNEVGVWFWFVPRDASTMHMSVILNSGTVRAMMWDGAGGIWKEFNTISHQKCDIYGHCGAYGMCD
SSSSFLSECMCLPGYEPKSQSKVLFQDGNGGCVRKRSESSVCGKGEGFVRVRNVKIPDTSEAVWIDQGLNKVNCEEECRKNCSCCAYTTANIVGKGTGCL
TWYGELMDTRHLPIPDQEIYVRVDTLELAEYAVQSAEFDEISFKLNVLAPSVSSVLIVCILFAYTLFKRWRKWKARKAKERRIRNLFRPSEDSSYLQNSI
VEKETERSKAYPELPFFSLDVIRAATNNFSSDNKLGEGGFGSVYKGKFPSGEMVAVKRAPRNSRQGLEQFKSEVFLISKLQHRNLVKLHGCCIEEVEQML
IYEYLPHKSLDSLLFVEHQTDSPFLDWEKRFNIIVGIARGILYLHQDSRFRIIHRDLKISNVLLDAELKPKISDFGMARVLEGNQVKGMTSKIGGTYFGA
MLLEIVTGKKINSGFTTHEEDCAALNLIGYVWELWKAGRVGDIVDLLLQVSTSTLRCIHIGLSCLEEDPVDRPDMQIVVVMLNSETIPLPPLKRPAFVCG
ANSTVEGGHPFPILISLEGRKSSSQRLISTKYQMDAQRLILLPLLLTLLFHDKASASSDDTLHANRSINDGELIISKGGNFALGFFSPGNSTYRRYLGIW
FHNKPGQTVVWVANRNTPINDTSGFLVNDGYGNILLYSSTRNQEVPIWSTNVSRAFPCLAQLLDSGNLVLVLQGSTSSNNTSLLVWQSFDHPTDTFLPGM
KLGINLETGLDMFLTSWRSEDDPGTGDFSFKLNPNGSPQFFLYQGMNPRWRSIPWPWTVDHPLYVDSFVNNEVGMWYSYAPRDASTVFMYVIHNSGTVKA
VVWDEGGGIWKEFYTILHQKCDIYGYCGAYGKCDPGSSFLSECICLPGYEHPISQSKVLSEDGNRGCVRKRSESSLCAKGEGFVRVRNVKIPDASKAVWV
DQGLNEVNCEDECRKDCSCTAYTTPNIVAKGTGCLTWHGKLMDTVDFPTPDQEIYVRVDALELAEYAVQSDELSFKLSVLVPSVTSVCIVCILFAYTLLK
RWRKRKARKATERRIRNLFRPSEDSSYLQDSIVTKEMERSKMYPELPVFSLDVIRAATNNFSSDNKLGQGGFGSVYKGKFPTGEEIAVKRPSKNSRQGVE
QFKSEVFLISKLQHRNLVKLYGCCIEKEEQMLIYEYLPYKSLDSLLFVEHQLGSNLFLDWGKRFNIIIGIARGILYLHQDSRFRIIHRDLKISNVLLDAE
FNPKISDFGMARILEGSQVEGMTRRIGGTYGYMAPEYAYCGKFSVKSDVYSFGVMLFEIVTGKKIKSGFATHEEDHAALSLIGYVWELWKAERVGDIVDA
SLQLSISTMRCIHIGLLCLEEEPVERPDMQTVVVMLNSETTPLPLPKRPAFVCGAKSGVEVRVEEECSINEFTFSDTLTR
|
DESeq2's median of ratios [FLAX]
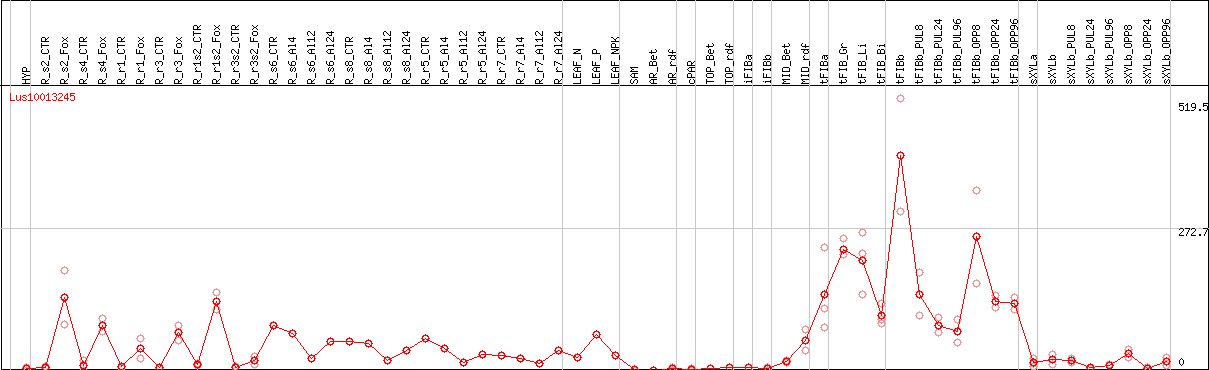
Coexpressed genes
Lus10013245 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
