External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G27000 254 / 6e-84
Protein of unknown function (DUF1664) (.1)
AT2G02730 223 / 4e-72
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2)
AT1G04960 187 / 2e-57
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2)
AT1G24265 96 / 9e-23
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2), Protein of unknown function (DUF1664) (.3)
AT1G24267 72 / 2e-14
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2)
AT1G44674 47 / 3e-07
Protein of unknown function (DUF1664) (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10036718
414 / 1e-146
AT1G27000 272 / 1e-90
Protein of unknown function (DUF1664) (.1)
Lus10037210
387 / 6e-135
AT1G27000 252 / 8e-82
Protein of unknown function (DUF1664) (.1)
Lus10030793
401 / 2e-131
AT1G69830 1173 / 0.0
alpha-amylase-like 3 (.1)
Lus10014626
191 / 1e-58
AT1G04960 318 / 3e-108
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2)
Lus10033807
136 / 1e-38
AT1G04960 218 / 2e-70
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2)
Lus10011347
96 / 1e-22
AT1G24265 226 / 2e-71
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2), Protein of unknown function (DUF1664) (.3)
Lus10003127
95 / 1e-22
AT1G24265 202 / 6e-63
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2), Protein of unknown function (DUF1664) (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G092700
327 / 3e-112
AT1G27000 331 / 1e-113
Protein of unknown function (DUF1664) (.1)
Potri.008G149000
321 / 6e-110
AT1G27000 327 / 3e-112
Protein of unknown function (DUF1664) (.1)
Potri.014G160800
200 / 9e-63
AT1G04960 309 / 6e-105
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2)
Potri.002G219300
200 / 1e-62
AT1G04960 325 / 7e-111
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2)
Potri.001G058700
110 / 7e-28
AT1G24267 211 / 1e-65
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2)
Potri.003G169200
108 / 3e-27
AT1G24267 214 / 5e-67
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF07889
DUF1664
Protein of unknown function (DUF1664)
Representative CDS sequence
>Lus10013269 pacid=23166651 polypeptide=Lus10013269 locus=Lus10013269.g ID=Lus10013269.BGIv1.0 annot-version=v1.0
ATGGCCATGCAAGCTGCACTCAGTTCCCGCCTTCTTATACTTGGCGGAGCAGGTTATTTTGGTACCGTATTGTTCAAGAAAGGTAAATTATCCGACCTGA
TCGGTGAGCTCCAGGCGTTGCTGAAGGGAATCGAGGGAGGCGATACTGATGGCGAATCTGGTCACGCTGACGCCATTGCTCAGCAGGTTAGTAGACTAGC
ACAAGAGGTTCGCCAGCTCGCTTCTCAGAGGCAGATTACTGTTGTAAATGGGAATGGCCAGTCAGGTAATTTAACTGCTCTGATCATTCCTGGAGCTGCA
ATTGGAGTAGCGGGTTATGGTTATATGAGGTGGAAGGGCATGCAATTTTCGGATCTTATGTATGTAACAAAGAGAAATATGACTGCTGCTGTTGCAAACT
TGACGAAACATCTTGAGGACGTCACTGGTGCTCTGGCAAAAGCAAAGGAACATTTAACTCAGCGCATTCAAGGTTTGGACGATAAAATGGATGTTCAGAA
GGAGATCTCAAAGGCAATTCAGAATGATGTCAATACTATCAATGAGAATATACAGCTCCTCGACTCCGACCTGTTCAATTTGCAGCGTATTGTTTCTGGT
TTGGATGGAAAGATAGGTACGTTGGAGAACAAGCAGGATCTAGCGAATCTGGGAATCACTTATCTGTGCCACTTCGCTCAGGGAAAGAAATTGGCAATGC
CTAAAGAGCTGAAGGGCCTGAAGGCGCTGATCGACGATTCTTTCTGCAAGTTGGTGACAGAAAATCTAGCTCGTACCGCAAGCGTTCCAAAGATCGATGC
TGTCCAGGTGAACAACGAAGAGCCCTCACACTCACAGACCCCTCGCGAACCTCTATTGAGGTAA
AA sequence
>Lus10013269 pacid=23166651 polypeptide=Lus10013269 locus=Lus10013269.g ID=Lus10013269.BGIv1.0 annot-version=v1.0
MAMQAALSSRLLILGGAGYFGTVLFKKGKLSDLIGELQALLKGIEGGDTDGESGHADAIAQQVSRLAQEVRQLASQRQITVVNGNGQSGNLTALIIPGAA
IGVAGYGYMRWKGMQFSDLMYVTKRNMTAAVANLTKHLEDVTGALAKAKEHLTQRIQGLDDKMDVQKEISKAIQNDVNTINENIQLLDSDLFNLQRIVSG
LDGKIGTLENKQDLANLGITYLCHFAQGKKLAMPKELKGLKALIDDSFCKLVTENLARTASVPKIDAVQVNNEEPSHSQTPREPLLR
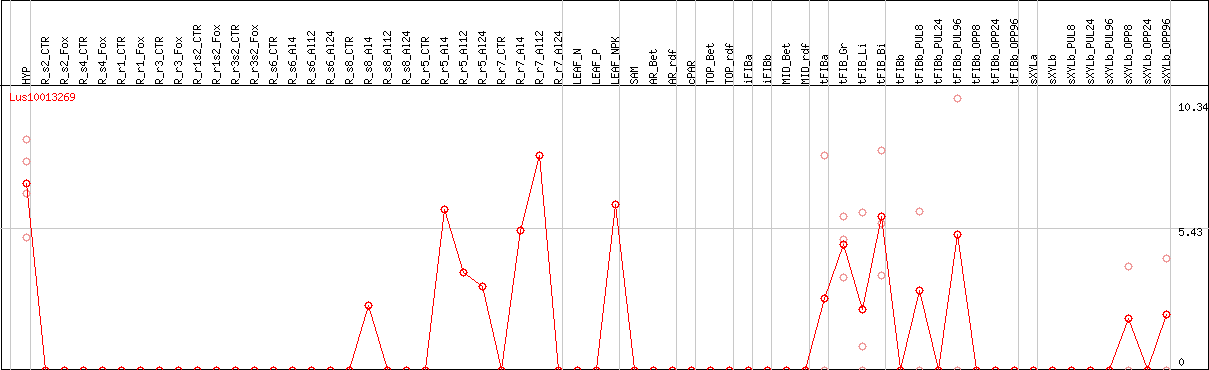
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013269 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.