External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G22070 163 / 5e-48
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT3G52060 142 / 7e-40
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
AT5G25330 119 / 4e-31
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT4G32290 107 / 1e-26
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT3G21310 83 / 5e-18
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT1G68380 81 / 4e-17
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT1G68390 80 / 7e-17
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT1G51770 79 / 2e-16
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
AT4G31350 79 / 2e-16
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
AT5G25970 75 / 4e-15
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10004106
316 / 9e-107
AT5G22070 486 / 3e-172
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10008954
133 / 2e-36
AT3G52060 400 / 2e-139
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
Lus10028866
130 / 4e-35
AT3G52060 393 / 1e-136
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
Lus10001503
121 / 7e-32
AT4G32290 382 / 7e-132
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10002919
118 / 9e-31
AT5G25330 389 / 4e-135
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10035828
88 / 2e-19
AT1G68390 401 / 7e-138
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10036610
87 / 2e-19
AT1G68390 404 / 3e-139
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10036774
84 / 3e-18
AT5G16170 350 / 1e-118
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10037154
82 / 1e-17
AT5G16170 253 / 1e-81
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G215400
196 / 5e-60
AT5G22070 421 / 1e-146
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.009G017300
173 / 1e-51
AT5G22070 418 / 7e-146
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.008G018600
156 / 1e-45
AT3G52060 407 / 2e-142
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
Potri.010G242900
145 / 4e-41
AT3G52060 407 / 9e-143
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
Potri.015G045500
92 / 6e-21
AT5G16170 394 / 2e-135
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.004G097700
86 / 4e-19
AT5G16170 452 / 6e-159
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.008G123500
83 / 3e-18
AT1G68380 254 / 1e-82
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.010G121800
82 / 1e-17
AT1G68390 399 / 1e-137
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.001G195900
81 / 2e-17
AT3G21310 504 / 8e-180
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.006G233400
79 / 2e-16
AT5G11730 557 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0110
GT-A
PF02485
Branch
Core-2/I-Branching enzyme
Representative CDS sequence
>Lus10013361 pacid=23151335 polypeptide=Lus10013361 locus=Lus10013361.g ID=Lus10013361.BGIv1.0 annot-version=v1.0
ATGCCAGCCAACAACAGTCAACCATCCACGGCGGGAACATTCCCCTCCGCCGCCTCCTCCTCCTCCTCTTCCTCCACCCTCCACCACTCCCAATCCTTCA
AACTCCCATCCTCCGCCTCCATGTTCTCCACCCCTTTCCTCCTCACTTTCTCCTTAATCCTCTCCCTCCCGATCCTCTTCCTCCTCGCCCCACGCATCCT
CCCTCCCCGCCCTTCCTCTGTCCCCATCCCTCCTTCCGACGAGCTCGAAGACCTCTCCCTCTTCCGCCGCGCTTCCTCCACTCCCATCTCGAAACCCTCC
TCATTCTCCCATCTTTCATCCTCGAAGCCGAAACTCAAAATCGCTTTTTTGTTCCTAACCAACACCGATCTCCATTTCGCACCTCTCTGGGAGCGGTTCT
TCCGTCCTGTCTCCGGAAAAGACCTCTACAACGTTTACGTCCACGCGGATCCGACCTCCAATGTCACCAACCCCGGAGGCGTGTTCTTCGGCAGGTTCAT
CGACCACGCCAAGAAGACCTACCGCGCTTCCCCGACTCTCATATCCGCCACGCGCCGCCTCCTTGCTACGGCGATCCTAGACGATCCTTCCAACGTTTTC
TTCGCCGTTATATCGCAGTACTGTATCCCTCTGCATTCATTTCAGTACGTCTACCACTCGCTGATCAAGTCCAGATCGTTCGATCTGTCCGCCGGATCTG
ACGATCCGGCCGACGGCGATCTGTCTACCCGGCCGACGAGTCTTCCTTCGGGATCCGGCTACAGTACAGGAGCTTCATCGAGGTCGTGTCCAATGAGCCT
CGGTTGTGGAAGAGGTACAATGCACGCGGGAGACACACGATGA
AA sequence
>Lus10013361 pacid=23151335 polypeptide=Lus10013361 locus=Lus10013361.g ID=Lus10013361.BGIv1.0 annot-version=v1.0
MPANNSQPSTAGTFPSAASSSSSSSTLHHSQSFKLPSSASMFSTPFLLTFSLILSLPILFLLAPRILPPRPSSVPIPPSDELEDLSLFRRASSTPISKPS
SFSHLSSSKPKLKIAFLFLTNTDLHFAPLWERFFRPVSGKDLYNVYVHADPTSNVTNPGGVFFGRFIDHAKKTYRASPTLISATRRLLATAILDDPSNVF
FAVISQYCIPLHSFQYVYHSLIKSRSFDLSAGSDDPADGDLSTRPTSLPSGSGYSTGASSRSCPMSLGCGRGTMHAGDTR
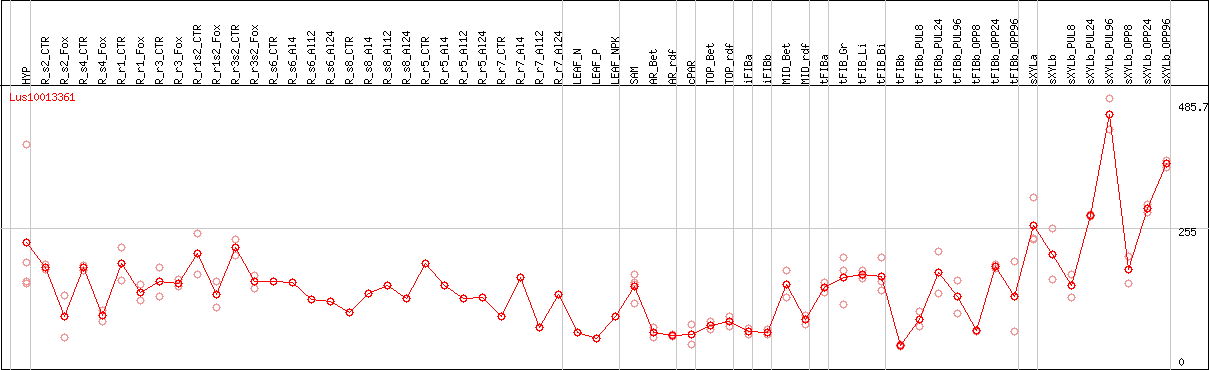
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013361 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.