Lus10013393 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10013393 pacid=23151388 polypeptide=Lus10013393 locus=Lus10013393.g ID=Lus10013393.BGIv1.0 annot-version=v1.0
ATGGACCCAGGAAGAGACACTGGGATCAGTCAAAAACTGGTCTTTGCCATTAACGGTCGTAGGTTTGAGCTCTCTGATGTCCACCCTTCTTCTACCTTGC
TTGAGTTTTTGCGTACTCACACTCCTTTCAAGAGTGTCAAGCTCAGTTGCGGCGAAGGCGGTTGTGGAGCATGTGTTGTCCTGCTTTCCAGGTACAATGA
TGTACTTGGTCAAGCTGAAGACTTGACAGTAAGTTCATGTCTCACTCTCCTTTGCAGTTTACATGGATGTTCGATCACAACAACCGAAGGCCTCGGAAAT
AGCAAAGACGGCTTTCACCCGATTCACGAGAGGTTTGCCGGGTTCCATTCTTCTCAATGTGGCTTCTGCACTCCTGGAATGTGCATGTCCCTCTTTGGCG
CATTAGTTAATGCTAACAAAGCAAATGGCCTTGAACCACCTTCGGGGCTGACAGCTGTTGGAGCTGAAAAGGCCATTGCTGGTAATCTCTGTCGATGTAC
TGGATATCGACCTATTGCTGATGCGTGCAAAAGCTTCGCAGTGGACGTTGATGTGGAGGATTTGGGATTCAACTCTTTCGGGAAGACGAGGAAGATGCCT
CTATACAGCAATGAACATGGAATCTGTAGGTTCCCCGAGTTTTTGAAACGGGAAGTCGATACGAATCTGTCATTGGATTATGATAAACACTGTTGGCTTA
CTCCTAAGGGGTTCGGGGAGCTTCAGAACATGTTGAAAAGCGTCGAAAATGACATAGGAAACCGGGTGAAGGTGAAGCTAGTGGTTGGTAATACTAGTGT
AGGGTATTATAAGGAACGTGAAGACTTTGACACATACATTGACATCAGGCATATTCCTGGTCTTTCTAAGATTAGGCGGGATCAGGAAGGGATCGAAATA
GGGGCATCCGTGACCATTTCGAATGCAATTCGTGCTTTAAAGGGTTCTTCTATGGTTTTTAGAAAGATTGCATCTCATATGGAGAAAATAGGCACCGAGT
TTGTTCGGAATACAGGCAGTCTAGGGGGGAATCTAGTGATGGCTCAAAGGCAACGGTTTCCTTCCGACATTGCCACGATACTTCTTGGAGCTGACGCGTA
CGTGAATCTACGTACTGGAACTACTTATGAAACTGTTAGCTTAGAAGTGTTCCTTGAAAGACCACCTTTGGATTCAAGAAGTGTGCTTTCAAGTGTCAAA
ATCCCTAATTGGGAGCTGGGAAGTAAACTGTTATTTGAAACGTATCGAGCTGCACCACGGCCGAGTGGAAATGCTTTGGCATATGTAAATGCTGCTTTCT
TGGCTGAAACTTCGTTCTGTGACTCTTCTGGTGGAACTGTGTTGGATAAATGTCTGCTAGCTTTTGGTGCGTATGGAACGAAACATGCAATTCGAGCGAG
AAAAGTTGAGCAGTTATTGGCAGGAAAGGTGCTGACTATGAATCTTCTGTATGAAGCTGTTAATTTGGTAAAGGAAGAAGTGGTACCTGAAGAAGGCATC
AGCAAACCAGCCTATAGGTTAAGTGTGGCTATTGGATTTCTCTTTGAATTTCTGAGTCCCTTGGCGGATACGGCTATTGGAATCTCTAGATGTTCTTTAT
CCTCAAATGGTGTTAAAGATAAATGTGAAGAAGGTGGCCATGTTAAATTCCCTAGAATTCTGTCATCGTCGAGGCAGGTCGTTCAATTGAATAGCGATTA
CCATCCTGTTGGTGAACCGATCGTAAAATCCGGAGCTGCCCTTCAAGCTTCTGGTGAAGCGGTGTATGTTGATGACATTCCCTCCACTGTAAATTGCCTT
CATGGCGCATTCATTTGTAGCACAAAGCCATTAGCTCGAGTAAAGGGCATAGAAGTCAACCCCAAATCACAGTTAAATGGGGTTTCAGCTCTCATTACAT
ATAAGGACATTCCGAATGGAGGTAAGAACATAGGTTCTATGACCATCTTTGGACCTGAACCTTTGTTTGCAGATGAGTTTGCTCGCTGTGCTGGAGATCG
CATTGCCTTCGTGATCGCCGGTACCCAGAAAGAAGCTGATGCAGCATCACGGCTCGCCACTGTTGATTATGATCTAGAGAATCTGGAAGCCCCTATCTTA
ACCGTTGAAGAGGCAGTTGAAAGGTCTAGCTTTTTGCATGTTCCTCCTTTCGTGCTTCCCAAGGAAGTCGGTGACTTCTCTAAAGGAATGACTGAAGCTG
ATCACAAGATACTTTCCGCTGAGATAAAGCTCGACCCTCAATATTACTTCTATTTGGAAACTCAAACCGCACTTGCTGTGCCCGATGAAGACAACTGTTT
GGTAGTTTACAGCTCTTCTCAGTGCCCTGAATTTGCCCATGCTACAATTGCTCAATGTCTTGGTGTTCCTGAACATAATGTACGTGTAATTACCCGACGA
GTTGGAGGGGGCTTTGGTGGGAAGGCAATTAAGTCCATGCCAGTTGCTACAGCCTGTGCGCTTGCTGCTCATGTGTTGCGCCGCCCTGTCAGAACATACG
TCGATCGAAAAACTGATATGATGATGGCAGGTGGCAGGCATCCCATGAAAGTAACATACAGTGTCGGATACAAGTCGAACGGGAAAATTACAGCACTGAA
ACTCGACATATTAGTCGATGGTGGGATATACACGGATGTAAGTCCCATAATCCCATTGAACTTCATATCCGTGTTTAAAAAATATGACTGGGGCGCTTTA
TCTTTCGACATTAAAGTCTGCAAAACCAACACAGTAAGTAGATCAGCAATGCGAGGGCCGGGGGAGGTACAAGGATCGTACATTGCCGAAGCTATAATCG
AACACGTAGCCTCTACGCTATCAGTGGATGTCCATTGTGTTCGAAAAACCAACCTTCATACACATAAAAGCCTTCACTTGTTCTATCCTGACGGCTCAGG
TGATGCTCAGGAGTATACTTTAACTTCGGTATGGGAGAAGTTAGGGATGTCTTCTAGCCATGACCAGAGGATGGAAATAGTAAAGGCGTTTAATAAGCAT
AATGTGTGGAGAAAACGAGGAATTTCTCGTATACCGATAATTCATCCGGTTACAGTGAGACCTACCCCTGGGAAAGTGAGCATTTTGAGAGATGGTTCTG
TTGTAGTTGAAGTTGGAGGGATAGAACTTGGTCAGGGGCTCTGGACAAAGGTGAAACAAATGACAGCATTTGCTCTTGGGTTGGTAAATTGTGTTGCTGC
TGAAGAGTTATTGGATAAAGTACGGATAATACAAGCTGATACTCTAAGTATGATTCAAGGCGGGTTTACTACGGGAAGCACTACTTCAGAGTCGAGCTGT
GAAGCGGTTAGGCTTTGTTGTGAGATTTTGGTTGAGAGATTGAAACCTCTAAAGGAAAGCTTGCAGAAGGAAATGGGTCCTGTGCAGTGGGAGATGCTTA
TACAGCAGGCACATATTCGAGCTGTGAACTTATCAGCGAGCTCGTTCTTTGCCCCAGATTTTTTGTCTGACCGGTACTTAACGTATGGGGCTGCGATCAG
TGAGGTGGAGGTAAACCTACTGACTGGGGAAACAACAATTCTGAGATCAGACATTCTTTATGACTGTGGACGAAGCCTGAACCCTGCAGTGGATTTAGGA
CAGATTGAAGGGGCTTTCGTGCAAGGTATAGGGTACTTCATGCTCGAAGAGTACACAACGAACTCAGACGGACTAATAACCTCAGACAGCACATGGACGT
ACAAGATCCCTACAATAGACACCATACCGAAAGAGTTCAACGTCGAAATACTAAGCAGCGGACACCACAAGAACCGCGTGCTCTCCTCCAAAGCTTCTGG
GGAGCCACCATTGCTCTTGGCGTCTTCAGTACACTGTGCTACTAGAGCAGCAGTCAGAGCAGCCCGGGAACAGCTCCGATCGTGGGGTTGCCACGAGGGG
ATCGATTCGTGGTTTGATTTGAAAGTGCCTGCTGTGATGCCTATTGTGAAGGAGTATTGTGGACTGGACATTGTAGAGAGGTACTTGAAATGGAAGATGA
GGAGTGAAAAAGAGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10013393 pacid=23151388 polypeptide=Lus10013393 locus=Lus10013393.g ID=Lus10013393.BGIv1.0 annot-version=v1.0
MDPGRDTGISQKLVFAINGRRFELSDVHPSSTLLEFLRTHTPFKSVKLSCGEGGCGACVVLLSRYNDVLGQAEDLTVSSCLTLLCSLHGCSITTTEGLGN
SKDGFHPIHERFAGFHSSQCGFCTPGMCMSLFGALVNANKANGLEPPSGLTAVGAEKAIAGNLCRCTGYRPIADACKSFAVDVDVEDLGFNSFGKTRKMP
LYSNEHGICRFPEFLKREVDTNLSLDYDKHCWLTPKGFGELQNMLKSVENDIGNRVKVKLVVGNTSVGYYKEREDFDTYIDIRHIPGLSKIRRDQEGIEI
GASVTISNAIRALKGSSMVFRKIASHMEKIGTEFVRNTGSLGGNLVMAQRQRFPSDIATILLGADAYVNLRTGTTYETVSLEVFLERPPLDSRSVLSSVK
IPNWELGSKLLFETYRAAPRPSGNALAYVNAAFLAETSFCDSSGGTVLDKCLLAFGAYGTKHAIRARKVEQLLAGKVLTMNLLYEAVNLVKEEVVPEEGI
SKPAYRLSVAIGFLFEFLSPLADTAIGISRCSLSSNGVKDKCEEGGHVKFPRILSSSRQVVQLNSDYHPVGEPIVKSGAALQASGEAVYVDDIPSTVNCL
HGAFICSTKPLARVKGIEVNPKSQLNGVSALITYKDIPNGGKNIGSMTIFGPEPLFADEFARCAGDRIAFVIAGTQKEADAASRLATVDYDLENLEAPIL
TVEEAVERSSFLHVPPFVLPKEVGDFSKGMTEADHKILSAEIKLDPQYYFYLETQTALAVPDEDNCLVVYSSSQCPEFAHATIAQCLGVPEHNVRVITRR
VGGGFGGKAIKSMPVATACALAAHVLRRPVRTYVDRKTDMMMAGGRHPMKVTYSVGYKSNGKITALKLDILVDGGIYTDVSPIIPLNFISVFKKYDWGAL
SFDIKVCKTNTVSRSAMRGPGEVQGSYIAEAIIEHVASTLSVDVHCVRKTNLHTHKSLHLFYPDGSGDAQEYTLTSVWEKLGMSSSHDQRMEIVKAFNKH
NVWRKRGISRIPIIHPVTVRPTPGKVSILRDGSVVVEVGGIELGQGLWTKVKQMTAFALGLVNCVAAEELLDKVRIIQADTLSMIQGGFTTGSTTSESSC
EAVRLCCEILVERLKPLKESLQKEMGPVQWEMLIQQAHIRAVNLSASSFFAPDFLSDRYLTYGAAISEVEVNLLTGETTILRSDILYDCGRSLNPAVDLG
QIEGAFVQGIGYFMLEEYTTNSDGLITSDSTWTYKIPTIDTIPKEFNVEILSSGHHKNRVLSSKASGEPPLLLASSVHCATRAAVRAAREQLRSWGCHEG
IDSWFDLKVPAVMPIVKEYCGLDIVERYLKWKMRSEKE
|
DESeq2's median of ratios [FLAX]
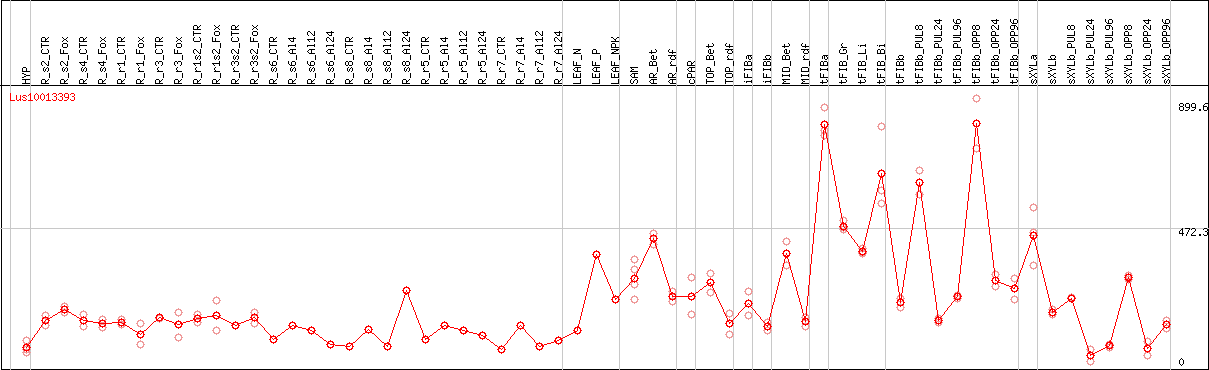
Coexpressed genes
Lus10013393 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
