External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G65480 243 / 4e-83
FT
FLOWERING LOCUS T, PEBP (phosphatidylethanolamine-binding protein) family protein (.1)
AT4G20370 236 / 2e-80
TSF
TWIN SISTER OF FT, PEBP (phosphatidylethanolamine-binding protein) family protein (.1)
AT5G62040 188 / 1e-61
BFT
brother of FT and TFL1, PEBP (phosphatidylethanolamine-binding protein) family protein (.1)
AT5G03840 176 / 1e-56
TFL-1, TFL1
TERMINAL FLOWER 1, PEBP (phosphatidylethanolamine-binding protein) family protein (.1)
AT2G27550 174 / 7e-56
ATC
centroradialis (.1)
AT1G18100 148 / 7e-46
MFT, E12A11
MOTHER OF FT AND TFL1, PEBP (phosphatidylethanolamine-binding protein) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10004452
314 / 4e-108
AT1G65480 270 / 1e-90
FLOWERING LOCUS T, PEBP (phosphatidylethanolamine-binding protein) family protein (.1)
Lus10021372
181 / 2e-58
AT5G03840 267 / 1e-92
TERMINAL FLOWER 1, PEBP (phosphatidylethanolamine-binding protein) family protein (.1)
Lus10027442
180 / 3e-58
AT5G62040 248 / 4e-85
brother of FT and TFL1, PEBP (phosphatidylethanolamine-binding protein) family protein (.1)
Lus10005753
172 / 5e-55
AT5G62040 241 / 5e-82
brother of FT and TFL1, PEBP (phosphatidylethanolamine-binding protein) family protein (.1)
Lus10020600
170 / 2e-54
AT2G27550 263 / 3e-91
centroradialis (.1)
Lus10004886
164 / 6e-52
AT2G27550 267 / 2e-92
centroradialis (.1)
Lus10043385
151 / 8e-47
AT2G27550 259 / 2e-89
centroradialis (.1)
Lus10002592
142 / 3e-43
AT1G18100 207 / 6e-69
MOTHER OF FT AND TFL1, PEBP (phosphatidylethanolamine-binding protein) family protein (.1)
Lus10018417
139 / 4e-42
AT1G18100 205 / 4e-68
MOTHER OF FT AND TFL1, PEBP (phosphatidylethanolamine-binding protein) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G179700
285 / 1e-99
AT1G65480 286 / 2e-100
FLOWERING LOCUS T, PEBP (phosphatidylethanolamine-binding protein) family protein (.1)
Potri.008G077700
281 / 3e-98
AT1G65480 257 / 8e-89
FLOWERING LOCUS T, PEBP (phosphatidylethanolamine-binding protein) family protein (.1)
Potri.004G203900
191 / 1e-62
AT2G27550 286 / 4e-100
centroradialis (.1)
Potri.009G165100
189 / 4e-62
AT2G27550 293 / 7e-103
centroradialis (.1)
Potri.015G041000
144 / 5e-44
AT1G18100 291 / 2e-102
MOTHER OF FT AND TFL1, PEBP (phosphatidylethanolamine-binding protein) family protein (.1)
Potri.015G141300
139 / 1e-42
AT5G62040 217 / 1e-73
brother of FT and TFL1, PEBP (phosphatidylethanolamine-binding protein) family protein (.1)
Potri.010G179801
132 / 2e-40
AT1G65480 133 / 3e-41
FLOWERING LOCUS T, PEBP (phosphatidylethanolamine-binding protein) family protein (.1)
Potri.010G179900
131 / 4e-40
AT1G65480 132 / 6e-41
FLOWERING LOCUS T, PEBP (phosphatidylethanolamine-binding protein) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01161
PBP
Phosphatidylethanolamine-binding protein
Representative CDS sequence
>Lus10013532 pacid=23178144 polypeptide=Lus10013532 locus=Lus10013532.g ID=Lus10013532.BGIv1.0 annot-version=v1.0
ATGCCAAGGGACAGGGATCCATTAGTAGTTGGGCGAGTGATAGGGGATGTGTTGGAACCATTCACAAGGTCTATCCCACTAAGAGTGATTTACATGAATA
ACAGAGAGATCAACAATGGCTGCGAACTCAAACCTTCTCAAGTTGTTAATCAGCCTCGTGTCGACATTGGTGGGGATGACCTCCGCACCTTTTACACTCT
GGTAATGGTCGACCCTGATGCACCAAGTCCTAGCGATCCTAGTTTGAGAGAGTATTTACATTGGCTAGTCACAGATATTCCTGCCACCACTGGGGCCAAC
TTTGGGCAAGAGGTGGTGTGTTACGAGAGCCCCAGGCCTTCAGTTGGGATACACCGTTTCATTTTCATATTGTTCAGGCAACTGGGTCGCCAAACGGTTT
ACGCCCCCGGTTGGCGCCAGAATTTCAACACCCGCGACTTTGCTGAACTCTACAACTTAGGTCCGCCTGTTGCTTCCGTCTATTTTAACTGCCAACGAGA
GAGCGGTTCTGGAGGTAGACGGCGATGA
AA sequence
>Lus10013532 pacid=23178144 polypeptide=Lus10013532 locus=Lus10013532.g ID=Lus10013532.BGIv1.0 annot-version=v1.0
MPRDRDPLVVGRVIGDVLEPFTRSIPLRVIYMNNREINNGCELKPSQVVNQPRVDIGGDDLRTFYTLVMVDPDAPSPSDPSLREYLHWLVTDIPATTGAN
FGQEVVCYESPRPSVGIHRFIFILFRQLGRQTVYAPGWRQNFNTRDFAELYNLGPPVASVYFNCQRESGSGGRRR
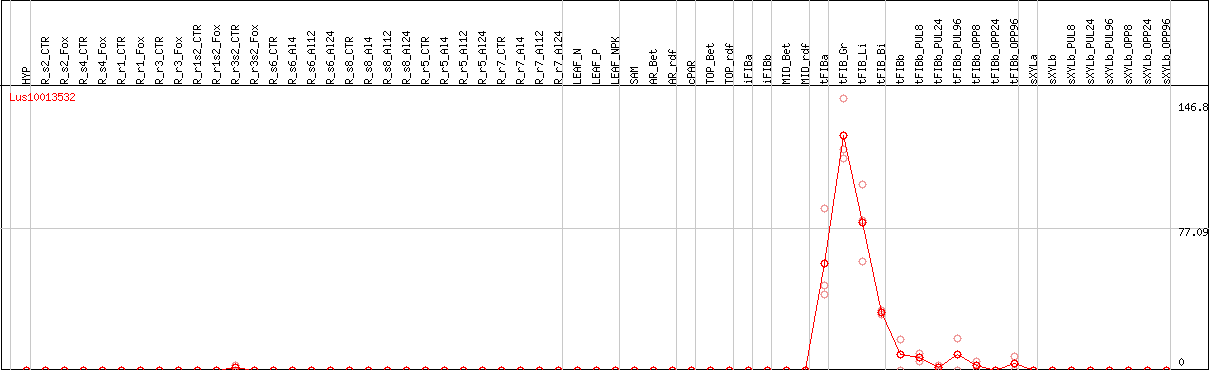
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013532 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.