Lus10013536 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10013536 pacid=23168719 polypeptide=Lus10013536 locus=Lus10013536.g ID=Lus10013536.BGIv1.0 annot-version=v1.0
ATGCACAACCCCTATCAGAAATCTCAAGACATAGCCTATACTAAGCAGCCTTACGTCGAAGATGCTGGTCCGCGAAGAATTGTCTCGTGTGCAGCAAAAG
CAATGGAATTCTCCATTATGTCACAAGCTGATGTTCTGAAGGCGGCTGAATGTCAAGTGTATCTTGGCCAGTACTATGATACCAGCGTAAAGCCAGTCCC
AGGTGGCTTGCTGGATCCCCGAATGGGTCCTGCCAAGAAAAAAGACATGCCTTGTGAAACCTGTGGCGGTAGCTTCAACGACTGCCCTGGCCACTTTGGA
TATGTGTCACTTGCGCTGCCAGTTTTCAATATTGGATACCTGAGTTTTGTTCTGGACATTCTGAAGTGCATCTGTAAAAATTGCTCTTTTATACTTCTGG
ATGAGAAGTTACGTGCCAAGTATTTAAGGCAGATGAGAAGTCCCAGGGTAGATCATTTGAAGAAGACCGCTATGATGAAGGCTGTTGTCAAGAAATGCAG
TACCATGGCAAGCGGGAAAGATATAAAATGCATCAAATGCCATTACTTGAATGGAATGGTGAAAAAGGATAAGGGTTATTTGGTTCACGACCGTTCAAAG
TCAGGTTCAAAGTCAGGTTCAACGTCAGGCGATTGCTTTACTGACGAGTGTAGATCTGCCATTTCAAACACCAAGGAGTCGAGGCCAACGATTAATCGAG
CCACTTACTACCTTAACCCTTTTGAAGTGTTGTATCTCTTTCAAAGAATTTCAGATGAGGATTGTGAATTGCTTTATCTAGGTGATAGACCTGAAAAGCT
AATCATCGCCAAGATTGCTGTGCCACCTATTGCTATTCGGCCGTCAGTTATTGTGGATGGGACACAAAGCAATGAAAATGACATTACAGTGAGGATGAAA
GATATCATTGTAGCCAGCGCTAATCTTCTAAACGAGCTTCGAGAGGGGAAAATTGCGAAAGACAAACATCTGGACAAGTGGGATCACCTTCAAGCTTTGG
TTTCACTCTATCTAAATAGTGACGCTCGAGTTCCGATAACTCGTGAATTGATGGAGCAACAGAAACAGAGGGCTAACACCTTAAGTGGTCTTACTCAGCG
CCTTAAAGGAAAACAGGGGAGGTTTCGTGGCAATCTGTCAGGGAAGCGTGTTGAATTCACTGGCCGAACTGTTATATCTCCAGACCCTAATCTAAAGATT
ACTGAGGTAGCAATTCCAATACATATGGCTCGCATTTTAACTTATCCTGAGTGTGTTAACCCTCGCAATATCGAGAAGTTGAAGCAGTGTGTCAGTAATG
GCCCACTGAAATACCCTGGTGCAAGGATGGTTAGATATCCAGATGGTTCTCAAAAGACGTTGCAAGGAAATTTCAGAAAACGCGTTGCTGAAGAGCTGAA
ATTTGGCTGTATAGTAGATCGTCATTTGGAGGATGGTGATGTCGTTCTTTTTAACCGACAACCAAGCTTGCATAGGATGTCTATCATGTCCCATAGGGCC
AGAATTATGCCCTGGAGGACATTGAGATTTAACGAATCTGTTTGTAACCCATATAATGCTGATTTTGATGGTGATGAGATGAATATGCATGTCCCGCAAA
CAGAAGAGGCTCGTACTGAGGCTCTCACACTGATGGGGGTGCAAAACAATTTGTGCACACCAAAAAATGGAGAGATTTTGGTTGCTTCTACACAAGATTT
TTTGACGACTTCATTTCTCATTACAAGGAAGGATACCTTCTATGACCGTGCAGCATTCTCACTTATGGCAACGTATATGGGGGATGGAATGGATTCAGTG
GATCTCCCCACACCTGCAATTTTGAAGGTCAGACAAGGAGAACTTCTTGTTTCTCATCGGAATGGGAACAAGGATGGACTTTACTCTGTTCTTCTCCGAG
ATTACAGTTCTTTTGGCGCTGCTGTTTGCATGAATCGACTAGCAAAGTTAAGTGCACGATGGATTGGAAATCATGGCTTCTCCATTGGAATTGATGATGT
TCAACCAGGAAAACAATTGAGTGACGAGAAGGGAAGAACAATCTCAAATGGTTATCAACATTGCAACAAAGTAATAGAGCAATACAATGCTGGAAAGCTT
GCACTCAAACCTGGTTGTGATGCAACCCAAACTTTGGAGGCTGAAGTGTGCATGAAAGAATTGCACTGGCGAAACAGTCCATTGATCATGTCGCAGTGCA
GATCTAAAGGTTCTCCTATCAATATCAGTCAGATGATTGCATGTGTTGGTCAACAGTCAGTTGGTGGCCGCCACGCTCCAGATGGATTCATGAATAGAAG
TCTTCCCCATTTTCCTAGGATGTCCAAAACCCCTGCTGCTAAAGGATTTGTCGCTAATTCTTTCTACAGCGGATTGTCTGCAACTGAGTTTTTCTTCCAT
ACGATGGCTGGGAGAGAAGGTCTAGTGGATACGGCAGTGAAAACTGCTGATACAGGATACATGTCTCGAAGGCTGATGAAAGGCTTAGAAGATTTATCTG
TTCAATACGACAATACTGTCAGAAGTGCAAATGGGGGTATAGTTCAATTTATCTATGGGGATGATGGTTTAGATCCTGCTAAAATGGAAGCAAAAGGTGG
ATTTCCATTGAATTTTGAACGATTGCTCATGAAAGTAAAGGCTACATGTTCCGTTGAAGAAGACGAGTATTTATCTCCTTCAGACATGTCGAAGGCCATT
GAAGACTTGATGCTGCAAGATCGGACTCTTGACAATCCTTGTGCCAAGGCGTTCAGAAAGAGTCTCAGAAATTTCCTTGCCAAGCATGTTGTGACATTAG
AAAGCATGCTAAACAAGGTGAATCACCATGAGGAAGGAAAAACTGATGGTATGGACGCCGAGTTTGCACAAGGAGATTGTAGTACTAGTTGCAAAGTCAT
CTACAAAATATACGGTCTTACCAAACAGCAGTTGAAGGTCTTCCTGAGAACCTGCACAGATCGTTACCTTTGGAAGAAGGTGGAACCAGGAACTGCAATT
GGAGCAATTGGAGCTCAGAGTATCGGAGAACCTGGGACGCAGATGACGCTGAAAACTTTTCACTTTGCTGGAGTTGCTAGCATGAATATTACTCAGGGAG
TTCCTCGTATTAAAGAAATCATAAATGCAGCTAAAAAGATCAGTACACCAATCATAACTGCTGAGCTTGAGAACGCTACCAATGTGAAAGCCGCTCGAAT
TGTGAAAGGACATACTCAGAGAACCCTACTAGGCCAGGTTGCGAAGAGCATCCAAATCGTGATGACTTCAAGATCAGCTTCGGTAGTGGTTACTCTTGAC
ATGGAAATAATTCAAGAGGCACAACTCGCTGTGGATATTAACGTTGTATATGAGGCAATTGTTGGGATTCCAAGATCTAAAGTGAAGTCACAGCATGTAA
AGGTTCTAGATGACAGAAAGTTGGAGGTAACCCCGCCGGGCGACAGAGATAAACTTCAGTTTGAACTTCACAACATCAAAAATATGCTCCCAAATGTTGT
AGTGAAGGGTATTCATACAGTGGAACGGGCAATTATTGCTCAGAAAAAGAATGATGGCAGTGTCAAAGATGGAGGCCCCCCAAGATACAATATGCTGATA
GAAGGCTCTTTGAACGATTTTCCAAGCGATGTAGGAGAGGTGCTTGGAATAACAAGGTTCGGTATCCAGAAGATGGACAAAAGCGTGCTGATGTTGGCCT
CGTTCGAGAAAACCACTGATCACCTGTACAGTGCTTGCGTAAATCGGAGGGAAGACAAGATTGAGGGCGTCAGCGAGTGCGTCATAATGGGGATCCCTGT
CCCGCTGGGAACTGGGATGCTCAAAATCCTCCAGAAGGTCGATAAACCGCCAGCCATGTTGCAATATGGACCTGATCCCATTGTGGGTACTAGCAGCAGC
AGCAGCAGTAAGCAGACAGACTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10013536 pacid=23168719 polypeptide=Lus10013536 locus=Lus10013536.g ID=Lus10013536.BGIv1.0 annot-version=v1.0
MHNPYQKSQDIAYTKQPYVEDAGPRRIVSCAAKAMEFSIMSQADVLKAAECQVYLGQYYDTSVKPVPGGLLDPRMGPAKKKDMPCETCGGSFNDCPGHFG
YVSLALPVFNIGYLSFVLDILKCICKNCSFILLDEKLRAKYLRQMRSPRVDHLKKTAMMKAVVKKCSTMASGKDIKCIKCHYLNGMVKKDKGYLVHDRSK
SGSKSGSTSGDCFTDECRSAISNTKESRPTINRATYYLNPFEVLYLFQRISDEDCELLYLGDRPEKLIIAKIAVPPIAIRPSVIVDGTQSNENDITVRMK
DIIVASANLLNELREGKIAKDKHLDKWDHLQALVSLYLNSDARVPITRELMEQQKQRANTLSGLTQRLKGKQGRFRGNLSGKRVEFTGRTVISPDPNLKI
TEVAIPIHMARILTYPECVNPRNIEKLKQCVSNGPLKYPGARMVRYPDGSQKTLQGNFRKRVAEELKFGCIVDRHLEDGDVVLFNRQPSLHRMSIMSHRA
RIMPWRTLRFNESVCNPYNADFDGDEMNMHVPQTEEARTEALTLMGVQNNLCTPKNGEILVASTQDFLTTSFLITRKDTFYDRAAFSLMATYMGDGMDSV
DLPTPAILKVRQGELLVSHRNGNKDGLYSVLLRDYSSFGAAVCMNRLAKLSARWIGNHGFSIGIDDVQPGKQLSDEKGRTISNGYQHCNKVIEQYNAGKL
ALKPGCDATQTLEAEVCMKELHWRNSPLIMSQCRSKGSPINISQMIACVGQQSVGGRHAPDGFMNRSLPHFPRMSKTPAAKGFVANSFYSGLSATEFFFH
TMAGREGLVDTAVKTADTGYMSRRLMKGLEDLSVQYDNTVRSANGGIVQFIYGDDGLDPAKMEAKGGFPLNFERLLMKVKATCSVEEDEYLSPSDMSKAI
EDLMLQDRTLDNPCAKAFRKSLRNFLAKHVVTLESMLNKVNHHEEGKTDGMDAEFAQGDCSTSCKVIYKIYGLTKQQLKVFLRTCTDRYLWKKVEPGTAI
GAIGAQSIGEPGTQMTLKTFHFAGVASMNITQGVPRIKEIINAAKKISTPIITAELENATNVKAARIVKGHTQRTLLGQVAKSIQIVMTSRSASVVVTLD
MEIIQEAQLAVDINVVYEAIVGIPRSKVKSQHVKVLDDRKLEVTPPGDRDKLQFELHNIKNMLPNVVVKGIHTVERAIIAQKKNDGSVKDGGPPRYNMLI
EGSLNDFPSDVGEVLGITRFGIQKMDKSVLMLASFEKTTDHLYSACVNRREDKIEGVSECVIMGIPVPLGTGMLKILQKVDKPPAMLQYGPDPIVGTSSS
SSSKQTD
|
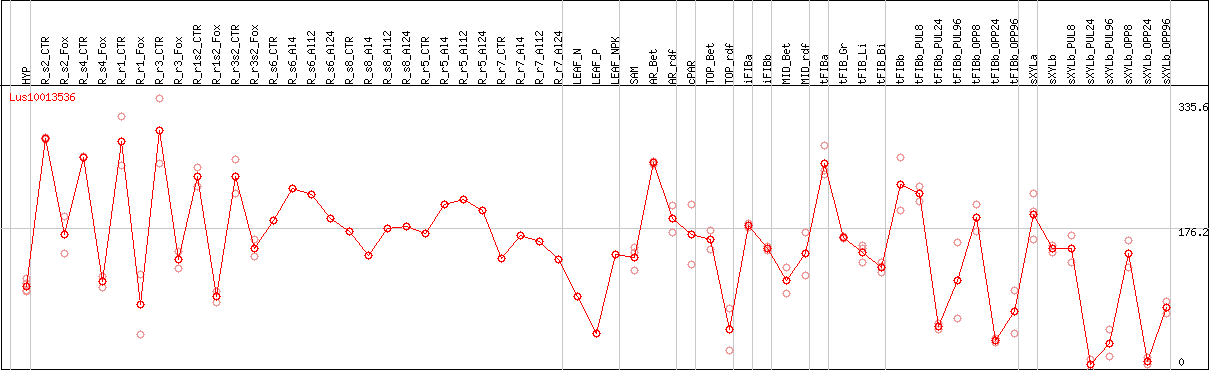
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013536 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
