External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G35070 93 / 6e-22
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
AT1G45976 62 / 6e-11
SBP1
S-ribonuclease binding protein 1 (.1)
AT3G12920 60 / 4e-10
BRG3
BOI-related gene 3, SBP (S-ribonuclease binding protein) family protein (.1)
AT1G32740 57 / 5e-09
SBP (S-ribonuclease binding protein) family protein (.1)
AT1G60610 55 / 2e-08
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2), SBP (S-ribonuclease binding protein) family protein (.3)
AT1G10650 54 / 3e-08
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
AT4G19700 53 / 7e-08
BOI, RING
Botrytis Susceptible1 Interactor, SBP (S-ribonuclease binding protein) family protein (.1)
AT1G79110 52 / 2e-07
BRG2
BOI-related gene 2, zinc ion binding (.1.2)
AT5G45100 49 / 2e-06
BRG1
BOI-related gene 1, SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
AT5G47050 47 / 6e-06
SBP (S-ribonuclease binding protein) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017276
260 / 4e-87
AT4G35070 122 / 2e-34
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Lus10014118
127 / 1e-35
AT4G35070 171 / 7e-54
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Lus10019790
125 / 7e-35
AT4G35070 170 / 1e-53
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Lus10016867
113 / 2e-31
ND /
Lus10002049
106 / 1e-27
ND /
Lus10010849
104 / 1e-27
ND /
Lus10005768
103 / 8e-26
AT5G52140 90 / 7e-21
RING/U-box superfamily protein (.1)
Lus10021853
100 / 2e-25
ND /
Lus10015574
100 / 4e-25
ND /
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G019000
139 / 5e-39
AT4G35070 148 / 6e-43
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Potri.005G242600
124 / 2e-33
AT4G35070 135 / 3e-38
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Potri.004G175500
114 / 8e-30
AT4G35070 142 / 1e-41
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Potri.009G135400
98 / 8e-24
AT4G35070 135 / 6e-39
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Potri.014G027000
60 / 6e-10
AT1G45976 380 / 2e-132
S-ribonuclease binding protein 1 (.1)
Potri.012G082200
59 / 1e-09
AT1G10650 266 / 2e-87
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Potri.002G124700
58 / 2e-09
AT1G45976 378 / 2e-131
S-ribonuclease binding protein 1 (.1)
Potri.001G148600
55 / 2e-08
AT1G32740 221 / 4e-70
SBP (S-ribonuclease binding protein) family protein (.1)
Potri.010G042200
53 / 1e-07
AT1G10650 393 / 2e-137
SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
Potri.015G118000
51 / 4e-07
AT5G45100 259 / 2e-85
BOI-related gene 1, SBP (S-ribonuclease binding protein) family protein (.1), SBP (S-ribonuclease binding protein) family protein (.2)
PFAM info
Representative CDS sequence
>Lus10013569 pacid=23168720 polypeptide=Lus10013569 locus=Lus10013569.g ID=Lus10013569.BGIv1.0 annot-version=v1.0
ATGGCTGTTCAAGCCCAGCCCTTCCTCTTCCCCATGTGTGCTTCCGACCTCGATCATCACCTCCCACCCTGTTTCCCTATTTTCCATGACTCCCATCCAC
ACCTATCACCACCCAACAACCTCTTCCTCTCTCAACCGCCGGCGGAAGCTTCTACCTCCACCTCCCTCCACCAACACCTCCAGATTCAGCGGCAGCAGCT
CGACGCCGTTCTCCAACTCCAGAACGAGAGGCTAAGAATGGCGTTACAAGAACAGAGGAAGCAACAAATAGGGGTTCTATTGAAAGTTATCGAAGCCCGT
GCCACGTCGATGCTGAGGCGGAAGCAGGATGAAATGGCATTAGCGGCGAGGAAAAAGGCTGAGCTGGAGGCTTGCTTAAGTAGAGCCGAGGCGGAGAAGG
ACACGTGGCAAAGGCTGGCTATGGAGAAGGAAGCGGCGGTGGCGGATCTTAAACGCCAGCTGGAGCGAATGAGAGGAGGAAACGACGCCGCTGCAGAGGA
TGCAGAGTCTTCCGCCGGAGATGATGAGTGGGATGGAAATGGAAATGGAAATGGCGGTGGTGGTTGCAAGAGGTGCGGGATTGGAGAGGCGTGTATGGTG
TTTCTGCCGTGTAGGCATGTGTGCTCCTGCAAGGACTGTGAAGGGTTTCTGGCCATTTGCCCTGTCTGTACATCTCTCAAGGAAGCCACCCTCGAGGGAG
GTTCTTGTGATGATGATGTTGAAGAACAAATGAATTATGGTGGTTATTTCTTAAACGGATTGTTTTACGACACACAAGATTATGCGTGTGATGCGGCGAA
GGACATTGCTTTAAGGTTAGGGTTTCAATTGATTCGCGACCCACATAAGAACAACCCGGGTGAAGAAGAAAAATGTTTGTACTTGTATTGCTCGCATGGT
GGTAGGAGCAAATGA
AA sequence
>Lus10013569 pacid=23168720 polypeptide=Lus10013569 locus=Lus10013569.g ID=Lus10013569.BGIv1.0 annot-version=v1.0
MAVQAQPFLFPMCASDLDHHLPPCFPIFHDSHPHLSPPNNLFLSQPPAEASTSTSLHQHLQIQRQQLDAVLQLQNERLRMALQEQRKQQIGVLLKVIEAR
ATSMLRRKQDEMALAARKKAELEACLSRAEAEKDTWQRLAMEKEAAVADLKRQLERMRGGNDAAAEDAESSAGDDEWDGNGNGNGGGGCKRCGIGEACMV
FLPCRHVCSCKDCEGFLAICPVCTSLKEATLEGGSCDDDVEEQMNYGGYFLNGLFYDTQDYACDAAKDIALRLGFQLIRDPHKNNPGEEEKCLYLYCSHG
GRSK
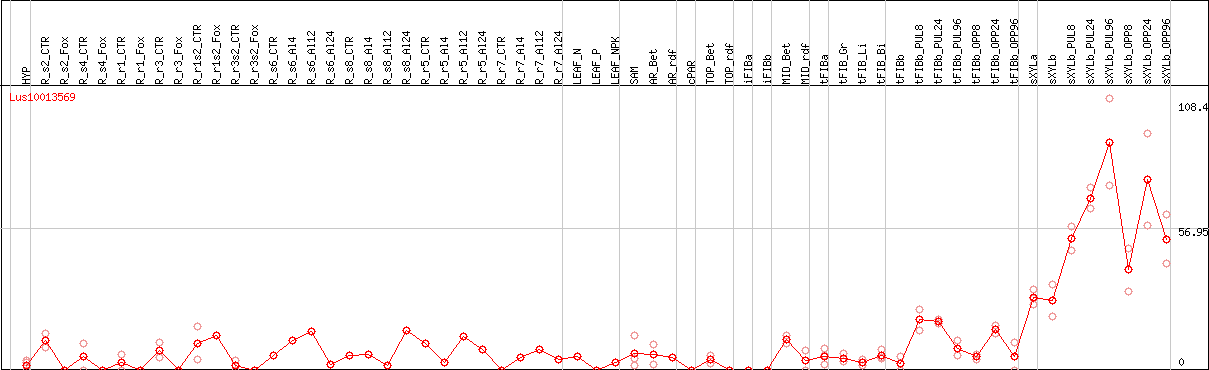
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013569 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.