Lus10013744 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10013744 pacid=23169081 polypeptide=Lus10013744 locus=Lus10013744.g ID=Lus10013744.BGIv1.0 annot-version=v1.0
ATGGCGTCTTCGTCTAGGATGGGTTCGGATCAGGCCACACCTCCGCAGCCTCAGCAGCGGAGGATCATGAGGACGCAGACGGCTGGGAATCTTGGAGAGT
CCATATTTGACAGTGAGGTTGTGCCTTCTTCTCTGGTGGAGATTGCGCCTATTCTTCGTGTGGCTAATGAAGTGGAGTCCAGTAACCCCAGGGTTGCCTA
TCTCTGCCGATTTTATGCATTTGAGAAAGCTCATAGGTTGGATCCCACTTCCAGTGGAAGAGGTGTTCGTCAATTTAAAACTGCACTACTCCAACGTCTT
GAAAGGGAAAATGATCCCACCTTGAAGGGTAGGGTTAAGAAAAGTGATGCGAGGGAAATGCAGAGCTTTTATCAACACTATTATAAAAAGTATATCCAAG
CTTTGCAAAATGCTGCTGATAAAGCCGATCGGGCACAGCTTACCAAGGCCTACCAAACTGCTAATGTGCTTTTTGAGGTTTTGAAGGCTGTAAATATGAC
ACAATCAATTGAAGTTGACAGAGAGATTTTAGAAGCTCAAGACAAGGTTGCGGAGAAGACTCAACTATACGTTGCTTACAATATACTCCCGTTGGATCCT
GATAGTGCGAACCAGGCGATCATGAGATATCCTGAGATCCAAGCTGCTGTTGTTGCTCTTCGCAACACAAGGGGGCTTCCATGGCCCAAGGATCATAAGA
AGAAGGATGAAGATATATTAGACTGGCTTCAGGCGATGTTTGGGTTCCAGAAAGGTAACGTTGCAAATCAAAGGGAGCATTTAATCCTGTTGTTGGCGAA
TGTGCACATAAGGCAGTTTCCGAAGATGGATCAACAACCCAAGTTGGATGACCGTGCATTAACTGATGTCATGAAGAAGCTTTTTAAGAATTACAAGAAG
TGGTGCAAATATTTGGACAGGAAAAGCAGTCTCTGGATGCCAACCATACAACAAGAAGTGCAGCAGCGCAAGCTGCTATACATGGGTCTTTATCTTCTGA
TATGGGGTGAAGCTGCAAACTTGAGATTTATGCCAGAGTGCTTATGCTACATCTACCACCATATGGCATTTGAGCTTTATGGCATGCTAGCAGGAAATGT
GAGTCCCATGACAGGAGAAAATGTAAAGCCTGCCTATGGTGGTGAAGAAGAGGCTTTCCTCACAAAAGTTGTTACACCTATCTACAATGTCATTGCAAAG
GAAGCTGAAAGGAGCAGAAAAGGGAAGTCAAAGCATTCTCAGTGGAGGAACTATGATGATCTGAATGAATATTTCTGGTCAGTCGATTGCTTCCGATTAG
GTTGGCCAATGCGTGCAGATGCCGATTTCTTTTCTCTACCGGCTGAACAGCATCATTTTGAGAAAGATGGGGACAACAACAAGCCAGCTTATAAAGATCG
ATGGGTGGGAAAGGTCAATTTTGTTGAGATTCGTACATTCTGGCACATCTTCAGGAGCTTTGATAGGATGTGGAGTTTTTTTATCCTATGCTTGCAGGCT
ATGATTATCATTGCTTGGAATGATTCTGCTGGCCGTCCTATCTTCACTGGTGATGTAGTCAAGAAAGTATTGAGTGTTTTTATAACAGCCGCCATATTGA
AACTAGGCCAAGCTGTGCTATTTCTCTTTCCCTTCGTGCGGAGGTTCCTAGAGCAGTCCCACTATAGAATAGTGATGCTCATGATGTGGTGGTCTCAGCC
TCCACTGTATGTTGCAAGGGGAATGCATGAAAGCACACTTGCACTGTTTAATTATACAGTGTTCTGGGTTCTTCTCATTATTACAAAGCTGGCCTTCAGT
TACTTTATTGAGATAAGGCCTTTAGTTGGTCCAACAAAAGCTATAATGAGTGTCCATATAAGTACTTTCCAATGGCATGAATTCTTCCCTCAAGCTAGGA
ACAATATCGGAGTGGTGATTGCTCTTTGGGCACCAATTATTCTTGTCTATTTCATGGATGCCCAGATTTGGTATGCCATATTTTCCACAATTTTCGGTGG
TATATATGGTGCATTTCGACGACTTGGAGAGATTCGAACTTTGGGAATGCTTCGCTCACGTTTTCAGTCATTGCCGGGTGCTTTCAATGCCCGCTTAATG
CCAGAAGAAAGGAGTGAACCAAAGAAGAAAGGATTAAGAGCTACCTTATCTCGCAATTTTGCAACTATACCGTCCAACAAAGAGAAAGAGGCTGCAAGAT
TTGCTCAGTTGTGGAATAAAATTATAACTAGTTTTAGAGAGGAAGATCTTATCAGTAACAGGGAAATGGACTTGTTGCTTGTGCCATATTGGGCTGACCG
TGATCTAGATCTCATACAATGGCCTCCATTTCTATTGGCTAGCATGATACCAATAGCATTGGACATGGCCAAGGATAGTAATGGGAAGGACAAAGAGCTG
AAAAAGAGAATTGAAGCAGAGAGCTATATGTCATGTGCTGTTCGTGAATGTTATGCTTCATTTAGGAACATTATCAAGTTCTTAGTGCAAGGCGACCGTG
AAAAAGAGGTAATTGAATATATATTTGAAGAAGTGGACAAGCACATTGAAGCAGGTGATCTTATCAGCGAGTACAAGATGAGCGCGCTTCCCAACCTCTA
TGAGCATTTTGTGCATATATCTAGTCTGGTTGATTCAATTCATGGCGGACCTGGCCATGAGGGGATGGTTCCTCTTGAGCAACAATATCAACTGTTCGCA
TCTTCTGGCGCTATCAAATTTCCCATTCAACCTGTAACAGAAGCATGGAAAGAAAAGATTAAGCGTCTTGATCTTTTGCTTACCACCAAAGAATCAGCAA
TGGATGTACCATCCAACCTGGAAGCCAGGAGGCGGATTTCTTTCTTTTCGAATTCGTTATTCATGGATATGCCCACTGCACCAAAAGTTCGCAATATGCT
TTCCTTCTCTGTTTTGACTCCCTACTACACAGAAGAAGTTTTATTCTCCTTGCGGGAATTGGAAGTACCAAATGAAGATGGGGTTTCAATCCTATTTTAC
CTGCAGAAGATCTATCCTGATGAATGGAATAACTTCCTTGAGCGAGTGAATTGCACCGGTGAAGAAGAACTCAAGGGAATAGATGACCTTGAAGAAGAAC
TTCGTCTATGGGCATCATACAGGGGCCAAACCCTGACGAGAACTGTGAGAGGAATGATGTACTACCGTAAGGCGTTGGAGCTTCAAGCATTTCTTGATAT
GGCTAGAGATGAAGATCTTATGGAGGGGTATAAGGCCATAGAACTGAATACTGAACTTCATTTGAGAGGAGAAAGATCATTGTTGGCCCAATGTCAAGCC
ATTGCTGATATGAAATTCACATATGTGGTGTCCTGCCAGCAATATGGTATCCACAAGCGGTCTGCAGACCCTCGAGCACAAGACACTCTGAGGCTCATGA
CTGCGTACCCGTCACTTCGTGTGGCGTACATTGATGAGGTTGAAGAGCCTAATAAAGATAAATCAAAAAAGGTCAACCAAAAAGTTTATTACTCAGTCTT
AGTGAAGGCTGCCTCACCTAAGGCCATCGATTCTTCAGAGCCTGTTCAGCACTTGGATGAGGTTATTTATCGAATCAAACTACCTGGGCCAGCCATCTTG
GGCGAGGGGAAACCAGAAAACCAGAATCATGCCATCATTTTTACACGAGGAGAGGGTTTACAGACTATAGACATGAACCAGGATAACTACATGGAGGAAG
CCTTGAAAATGAGAAATCTGTTGCAAGAATTTCTCAAAAAGCATGATGGTGTGAGATATCCTACCATACTTGGTCTTAGGGAGCACATATTCACTGGAAG
TGTTTCTTCTCTTGCATGGTTTATGTCAAATCAGGAGACTAGTTTTGTGACTATTGGTCAGAGACTACTAGCTAACCCGCTCAAGGTTCGGTTTCACTAT
GGTCATCCAGATGTGTTTGATAGGCTCTTCCACCTGACCAGAGGGGGAGTCAGTAAAGCCTCCAAGGTTATCAATCTCAGTGAGGATATATTTGCTGGTT
TCAATTCCACTCTTCGTGAAGGGAATGTTACCCATCACGAATACATACAGGTTGGGAAAGGAAGAGATGTCGGTCTCAATCAAATCTCTATGTTTGAAGC
CAAAATTGCTAATGGCAATGGGGAGCAGACTCTGAGTAGAGATATATACCGTCTTGGACACCGATTTGATTTTTTCCGCATGTTGTCTTGCTATTTCACC
ACAGTTGGTTTCTACTTCAGTACTCTGATAACTGTGCTAATTGTTTACGTATTCCTTTATGGTCGGCTTTATCTGGTTCTGAGTGGACTGGAAGAAGGCC
TGAAAAATCAGCGAGCTATTCGAGATAACAAGCCTCTGCAGGTGGCTCTTGCTTCTCAGTCGTTTGTTCAAATTGGATTTCTCATGGCATTGCCCATGCT
GATGGAAATCGGGCTGGAGAGAGGTTTCCGCACTGCCCTTAGTGAATTTGTACTGATGCAACTTCAGCTGGCACCGGTGTTTTTCACATTCTCTCTCGGA
ACCAAGACTCATTACTACGGACGGACTTTACTTCACGGAGGTGCCAAATACAGGCCTACAGGCCGAGGTTTTGTTGTGTTCCATGCCAAGTTTGCAGACA
ATTACAGGCTTTATTCACGTAGTCACTTTGTCAAGGGGATTGAGATGATTATTTTACTTGTCGTCTATGAAATCTTCGGGCAAACCTATAGAAGTGCTGT
TGCCTATGTCTTGATCACCATTTCAATGTGGTTCATGGTTGGTACCTGGCTGTTTGCTCCGTTCCTCTTCAATCCTTCTGGATTCGAGTGGCAGAAGATT
GTTGATGATTGGACTGATTGGAACAAGTGGATAAGTAACAGAGGTGGCATAGGTGTGCCACCAGAAAAGAGCTGGGAATCATGGTGGGAGGAGGAACAAG
AACATTTACATCACTCCGGGAAGCGTGGTATCGTTGCGGAGATACTTTTATCTTTGCGGTTCTTTATATACCAGTATGGACTGGTATACCACTTATCCAT
CACAAAGAAAGCCAAGACCAAAAGTTTTCTTGTTTATGGAATATCTTGGCTCGTCATTTTCTTGGTATTGTTTGTGATGAAGACGGTGTCTGTTGGCCGG
CGAAAATTCAGTGCAAACTTCCAGCTTGTCTTCCGGCTGATCAAGGGATTGATCTTCATTACTTTCATTTCCATTTTGGTCGTTCTCATAGCATTACCTC
ACATGACGATCCAGGATATCATTGTTTGCATTCTTGCATTCATGCCTACCGGCTGGGGAATGCTTCTGATTGCACAAGCTTGTAAACCAGTTGTTCATCG
AGCTGGGTTCTGGGGATCAGTTCGAACGCTAGCTCGTGGGTACGAGATTCTTATGGGCCTACTTCTTTTCACTCCGGTTGCATTCCTTGCTTGGTTCCCA
TTCGTGTCCGAGTTCCAGACTCGCATGCTTTTCAACCAAGCATTCAGCAGAGGTCTACAGATTTCACGTATTCTCGGCGGGCAGAGGAAAGATCGATCTT
CGCGCAACAAGGAGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10013744 pacid=23169081 polypeptide=Lus10013744 locus=Lus10013744.g ID=Lus10013744.BGIv1.0 annot-version=v1.0
MASSSRMGSDQATPPQPQQRRIMRTQTAGNLGESIFDSEVVPSSLVEIAPILRVANEVESSNPRVAYLCRFYAFEKAHRLDPTSSGRGVRQFKTALLQRL
ERENDPTLKGRVKKSDAREMQSFYQHYYKKYIQALQNAADKADRAQLTKAYQTANVLFEVLKAVNMTQSIEVDREILEAQDKVAEKTQLYVAYNILPLDP
DSANQAIMRYPEIQAAVVALRNTRGLPWPKDHKKKDEDILDWLQAMFGFQKGNVANQREHLILLLANVHIRQFPKMDQQPKLDDRALTDVMKKLFKNYKK
WCKYLDRKSSLWMPTIQQEVQQRKLLYMGLYLLIWGEAANLRFMPECLCYIYHHMAFELYGMLAGNVSPMTGENVKPAYGGEEEAFLTKVVTPIYNVIAK
EAERSRKGKSKHSQWRNYDDLNEYFWSVDCFRLGWPMRADADFFSLPAEQHHFEKDGDNNKPAYKDRWVGKVNFVEIRTFWHIFRSFDRMWSFFILCLQA
MIIIAWNDSAGRPIFTGDVVKKVLSVFITAAILKLGQAVLFLFPFVRRFLEQSHYRIVMLMMWWSQPPLYVARGMHESTLALFNYTVFWVLLIITKLAFS
YFIEIRPLVGPTKAIMSVHISTFQWHEFFPQARNNIGVVIALWAPIILVYFMDAQIWYAIFSTIFGGIYGAFRRLGEIRTLGMLRSRFQSLPGAFNARLM
PEERSEPKKKGLRATLSRNFATIPSNKEKEAARFAQLWNKIITSFREEDLISNREMDLLLVPYWADRDLDLIQWPPFLLASMIPIALDMAKDSNGKDKEL
KKRIEAESYMSCAVRECYASFRNIIKFLVQGDREKEVIEYIFEEVDKHIEAGDLISEYKMSALPNLYEHFVHISSLVDSIHGGPGHEGMVPLEQQYQLFA
SSGAIKFPIQPVTEAWKEKIKRLDLLLTTKESAMDVPSNLEARRRISFFSNSLFMDMPTAPKVRNMLSFSVLTPYYTEEVLFSLRELEVPNEDGVSILFY
LQKIYPDEWNNFLERVNCTGEEELKGIDDLEEELRLWASYRGQTLTRTVRGMMYYRKALELQAFLDMARDEDLMEGYKAIELNTELHLRGERSLLAQCQA
IADMKFTYVVSCQQYGIHKRSADPRAQDTLRLMTAYPSLRVAYIDEVEEPNKDKSKKVNQKVYYSVLVKAASPKAIDSSEPVQHLDEVIYRIKLPGPAIL
GEGKPENQNHAIIFTRGEGLQTIDMNQDNYMEEALKMRNLLQEFLKKHDGVRYPTILGLREHIFTGSVSSLAWFMSNQETSFVTIGQRLLANPLKVRFHY
GHPDVFDRLFHLTRGGVSKASKVINLSEDIFAGFNSTLREGNVTHHEYIQVGKGRDVGLNQISMFEAKIANGNGEQTLSRDIYRLGHRFDFFRMLSCYFT
TVGFYFSTLITVLIVYVFLYGRLYLVLSGLEEGLKNQRAIRDNKPLQVALASQSFVQIGFLMALPMLMEIGLERGFRTALSEFVLMQLQLAPVFFTFSLG
TKTHYYGRTLLHGGAKYRPTGRGFVVFHAKFADNYRLYSRSHFVKGIEMIILLVVYEIFGQTYRSAVAYVLITISMWFMVGTWLFAPFLFNPSGFEWQKI
VDDWTDWNKWISNRGGIGVPPEKSWESWWEEEQEHLHHSGKRGIVAEILLSLRFFIYQYGLVYHLSITKKAKTKSFLVYGISWLVIFLVLFVMKTVSVGR
RKFSANFQLVFRLIKGLIFITFISILVVLIALPHMTIQDIIVCILAFMPTGWGMLLIAQACKPVVHRAGFWGSVRTLARGYEILMGLLLFTPVAFLAWFP
FVSEFQTRMLFNQAFSRGLQISRILGGQRKDRSSRNKE
|
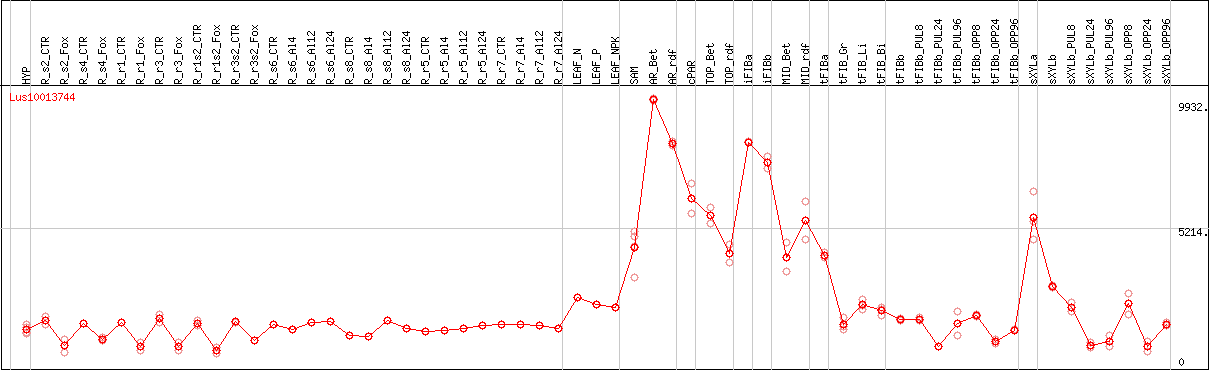
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013744 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
