Lus10013768 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10013768 pacid=23169113 polypeptide=Lus10013768 locus=Lus10013768.g ID=Lus10013768.BGIv1.0 annot-version=v1.0
ATGGCGATGAAGCAGGTAGCTACCGCGAAAAAGGGTGTTATCCATGAAGAACTAGCCAGAGTTTTGGTACAAGTTGAAAACGAACTTGAAAAGTTTCATG
ATTTACCTGATGTCGACAGCATTCTGTTTGAGAAAGCTAAATGGAAGGTTTATCTGAAGGATTCGTTTAAAAGTGTTCATTATCGAATGCTTTATGTTTT
GTATCTTCTCATCTTTATGCTGTTTTTCCTGAACAACAACTTCAGAACGTCATTTCAGAAGCTTGACTCATCTGCCTTGAAAAAGCTGGTTATGAACTTG
TTGCTGAGGCTTTCTTATGGATTGAGGCCGAGGAGAAGGAAAACAGACGTAGAATGTGAGAACTCAAAGCAGATTGTCAAGCATTTCAGGGTTAGGCATC
TGTTAGTTATATGCTCGGTGGATATTTTAAAGGACTCAGGCTTCCACTGGCAAGTCTTGAAGATTTGGGATATAGTGTCTGCTATGAACACGCCATTTCT
GATTAAACGTCTTGATTCCATCTTTGCATTGCATTCTGATGGTTACATCAATCGATGCAAGGTGAAGTCCTTTGAAGGCATACTAGAGGTTCCAATGACT
TGGAAACATGACATACCTCGCTACAAGAGCACTTGCAGTGCTGAAACTGGAAATGCATCTCCTGAAAGATGCATGACAAGCGATGGTTTGATGCTTATGA
AATTCTATTCTTTATCTTCTGGCATCAACAGGAACCTGCTATCTGGTTGTGATGGCCAGGAGGTTGATTTTCCATTAGAGCTGTCTGATCAAGAGAAGGA
AATTGTTCAATTTGATAAGAGCTCTTTCGTACTTGGTAGATCATGCAGTGGGAAAACCACTGTGTTGGTTATGAAGTTATTTCAAAGGGAGCAGCTTCAT
CACATAGCTTATCAAGGATTTCATGAGCAAGGTGACGATGTAGAAACTGAAGGCGGTAGTGTCTTGAAACAAATGTTTGTGACGGTAAATCCCAGATTAT
GCCATGTTGTTAAGCAGCACATTTCTCGTTTGAAAAGATCCACCTGCGGATGGAACTTCGTATCTGAAACCGCTGAACTTGAATTGGATGAAATTGATGA
GACATCCCTTTCTCTGGACTTCCCAAATTCATTTGTTGAAATCCCATTAATATGCTTCCCTCTTGTTATAACTTTCAAGAAGTTTTTGCTTATGCTGGAT
GCCACGGTCGGGGTCTCTTTCTTTGAGAGGTTCCCTGAGGCCTTTCATCAAGAAAAAGTTAAGAGTTCCAAGTCCTTGACCTTGCAAACTTTTATCAGAA
CCAGGGAGGTTGGTTATGAGAAATTTAGTTCATTGTACTGGCGCCACATGAACCAGAAAATATTAACCAAGAGTGTTGATCCGTTTCTGGTTTTCACCGA
GATAACTTCTCATATCAAGGGCGGATTCCAAACTTTAGAGGCCAGTGGTGGTCATCTAAGCCGTGAGGATTATGTGTCCCAATCTGAACGTCGTCTATCT
AATTTGACAGAGCTGGAAAGGGGCAGAATTTATGACATCTACCTTGAGTATGAAAAGAAAAAGGCTAAAAACGGAGAGTTTGATTTAGCAGATTTTGTGA
CCGACATACTTCTTCGGTTAAGAGTTGAAGGATTAATCAAAGCACCAATGGACTTTGTTTACGTTGATGAAGTTCAAGATCTTACAATGAAGCAGATTGC
TCTTTTCAAGCATGTCTGTTCAAATGTTGAAGAAGGTTTTCTTTTCACTGGCGACACAGCACAAGCGATTTCCAAAGGCGTTGAGTTCAGGTTCAAGGAT
ATAAGGTCCCTATTCTATGAAACGTTTATACTTAATTCAGGACGTTTAGGACCTTCCCGTGGCACTGCAAAGCAACAACTTTCTGAGGTTTTTCAGCTGA
GTCAGAACTTCCGAACACATGCTGGGGTTCTTAAGCTAGCTCAAACTGTTATTGACTTGATTTATTATTTCTTCCCTCGCTCGATTGATGAACTGAATCC
GGAGACGAGTCATCTTTATGGGGAGATGCCCGTGCTGCTTCATTCTTTGAAGCATAAGAGCATGTTCAACTGCATGTCTGACATTTACTTTGGTGCTGAC
AAGAAAAGTTGGGAGTTTGGGGCTGAGCAGGTAATATTAGTAAGAGATCAAGAAGTAAAAAACAATGTCGTATGCCAAGTGGGCAAGCAAGCTCTTGTTC
TCACGATATTGGAGTGCAAAGGCTTAGAGTTTGAGGATGTCTTGCTGTATGATTTTTTTGGTTCCTCTCCTGTGCCAGATCAGTGGGAGGTACTAAATAT
ATACATGAAGGCACATAAATTGTCTCCTAGTGATGAATGTCTGATGCCAAACTTAAATGAGGGAAAGCTGGTTTCTTTGTGTCACGAGCTCAAGTTACTG
TACGTAGCTATAACTCGTACTAGGCAAAGGTTCTGGATTTTGGATGAAATCTGGAATCCTATGCTGGAATTGTGGAAGCAGAAGGGTATTGTCAAGCTTC
AACAGTTCGATAGTGAAGTCTTACAAATTATGCAAGCCCCTTCTACTCGGGAAGCATGGAAGTCACGGGGGAAGAAGTTCTTCCATTTGCAGAACTATGA
AACTGCTAGAATGTGCTTCAAGAAAGCAGGGGAATCATACTGGGAAAAGTATTGCGAGGCGTCCGAATTGAGATCAAATGCTGATCAAATGAGGAGCTCT
GATCCTGTAGTATCTCGTCGTCATCTGTCAGATGCTGCTCTAATCTATGCCAAAATTGGGAAGGATGCACTTGCAGCAGAATGCTACTTCGAGTTGCAGG
ATTATGTCAAAGCAGCTGAAATTTATTTGGAGAAGTTTGGAGCCTCAAAACTACAAGAGGCTGGTGAATGTTTCTGTCTTGCCAAGTGCTACAGAAGAGC
TGCTGAAGTCTATTATAAGTGCAACGCTTATGTGAAGTGTTTGTCTGCCTGTATCGACGGCAAATGTTACGAAACCGGATATGACTACATTCAACAATGG
AGGAAGGATGGCGTTTTTAATTTTACGGAATTCGTGGAATTCCTTAAAAAAGGAGCACTTCATTTTCATAGGCTGGAAAATGACGAATTCATGATCATGT
TTGTGCGAGCTCTTAATTCCAAGGACCAAATACGTAGCTTCTTAAGAAGTAGACACTTGCTTGATTATCTTCTCCGCTTGGAGGAAGAGTGGGCAAATTA
TTTAGAAGCAGCGGCTATTGCAATGGATATCGGTCTGGAAACTGTTGCTGCTAATAATCACGAGAGGGGAGGACAATATCAAGAGGCTTGCCATGTTATC
CTGCGTCACGTGCTTGGTGAGTCCCTTTGGGGAGACGCAAATAAAGGCTGGCCGGTCAGAGAATTCGAGGGGAAGACGAAGCTACTGAACCAAGCTAAAA
CCTGTGCAGAGCATGTTTCAGCAGCTTACCATGGATTTGTTTGCCAGGAGACATTTATGTTATCTTATGCAAATGGTAATAGTTTGTGGAAGCTGAAAAA
TAGTTTTGAGTATTTTCAGCGGCAGGGCAATATTAGGAATGAAATATTGTCCGCTCGAAGTGTCATCGATATTTTCATTGATTCAAGTGCCTCGGGCAAC
TTCAGGGATGATGTTTTAGTTGCCGATATATTGAACTATACAGAAGATTGGAGGATGCCCTGGAAAAAGCTTTCCTTCTTAAATCTGTATGACTATTGGA
AGTCATGGAAGGGGCATATTGACAAAATAGTTGCATATTGTGTTGAGAAAGGACACTGGGGTTGTCATGGCGAGTTTTGCATGGACTACTTGGGTGTAAG
GGAGCAGCAGAGCGATAAAGAAACTGAATACAATGTACTTTATCCTGATGCAGCATGGATCAATTCTATGATTAGCTCGAAAGTTGTTCAGAACAAAAGG
AAGCGAGTCAAGGTTAACGCTGACCAGTTCTGGAAAGCTGCTAAAGACTATTGGTTCACAGAAAGACGTGTGGTTGGGGAGAAGGTGCTCCATAAACTTG
AAAACGTTTATGAATATTGCGTTCGGAATGGTTTATCAGTATTCTGCAGAAGCATGCTACTGGTAAAGATTTTTGAGCTGTCCCGTTCTCTTTTCTCATT
CAAATTGGGGAATCCCGTTTATCAGTATACTGCAGAAGCATGCTACTGGACAGATCTAGTACGAAGGTTTATCGAGCACTATACAGAGAACATCCTGAGC
ATTCATTTCCTCGAGCCGAACGAGTCATTCAGAAAGCATATGATCATCGCAAGGACATTGGAAGGTTTCAAGGTCCCTTTTGAGGAAGCAGTTTGTTATT
ACATAGAGAATTTACCACACGGGGTAACATATCGTCAAATTGCAGAGGTTACGGCAGTTGCTCTCTCGGGAAAACTTCATCACGAAACGCTACAGATGCT
GGTCCAAACCTTGAAGAAGAAGGCGGTTTGCAGCTCGTGCATTGAGGAAACCATGGGACCTTCGAAAGAAGCTACAGTGAGGAAGCTCCACGAAGTTGTA
GAACACATGTACTATGCCGAGTTTGAGAAAGCAGATGAGACTATGTCACCAAATTGTCTAGTGTATCTGCTAGACCGGTTGGTAATATTATTGTTGGCTG
TGAAGGGAGGTTTCTACTGTCCGAGATCATCCGTGGTTGAGTGGTTGATGCACAATGGATGGCATTACACCTGCTCAACTTCTACTTCTGATCGTTATGA
GCAACCTTTTGTGGAAGAAGCTCTAGATTCGGTATCCTTCATAGTCAAAGACATCTTCTGCAACGAGCGTGACTGCAAAACACCGAGATGGACAAAGTCA
TTCAATGAGAAATTGGAGTCTTATCCATTTGTGGTGCTGAGGTTAGTAGCCATAGTTTGCATTCTTAACATCAACACTGGAAAGTGCTCTGATCTGCTCT
CCAACATCTTGAGTAGAAGAGAAATCACGTCTCTGTCACCGCTGGAGTTCTACAGTTGTCTGACCCGTACGAGGTGTCAGGATGAAACGAACCATGCAGT
TGTGTCTTTTTCAAAAGCACTGGCTGAGATTAATAACCCTCTGGTTTATGTGGACTTGAAGGGAAGCGGTTCCACCAGATTTAAACCTCCAGGAGCCCCC
ATCATCTTTTTGGACACCAAGGTTAGTGGAAGACAAGACATACTGAATCGGGTGTGCCCGAGAAACATGTCGGCATCTGCAAATAGTGTCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10013768 pacid=23169113 polypeptide=Lus10013768 locus=Lus10013768.g ID=Lus10013768.BGIv1.0 annot-version=v1.0
MAMKQVATAKKGVIHEELARVLVQVENELEKFHDLPDVDSILFEKAKWKVYLKDSFKSVHYRMLYVLYLLIFMLFFLNNNFRTSFQKLDSSALKKLVMNL
LLRLSYGLRPRRRKTDVECENSKQIVKHFRVRHLLVICSVDILKDSGFHWQVLKIWDIVSAMNTPFLIKRLDSIFALHSDGYINRCKVKSFEGILEVPMT
WKHDIPRYKSTCSAETGNASPERCMTSDGLMLMKFYSLSSGINRNLLSGCDGQEVDFPLELSDQEKEIVQFDKSSFVLGRSCSGKTTVLVMKLFQREQLH
HIAYQGFHEQGDDVETEGGSVLKQMFVTVNPRLCHVVKQHISRLKRSTCGWNFVSETAELELDEIDETSLSLDFPNSFVEIPLICFPLVITFKKFLLMLD
ATVGVSFFERFPEAFHQEKVKSSKSLTLQTFIRTREVGYEKFSSLYWRHMNQKILTKSVDPFLVFTEITSHIKGGFQTLEASGGHLSREDYVSQSERRLS
NLTELERGRIYDIYLEYEKKKAKNGEFDLADFVTDILLRLRVEGLIKAPMDFVYVDEVQDLTMKQIALFKHVCSNVEEGFLFTGDTAQAISKGVEFRFKD
IRSLFYETFILNSGRLGPSRGTAKQQLSEVFQLSQNFRTHAGVLKLAQTVIDLIYYFFPRSIDELNPETSHLYGEMPVLLHSLKHKSMFNCMSDIYFGAD
KKSWEFGAEQVILVRDQEVKNNVVCQVGKQALVLTILECKGLEFEDVLLYDFFGSSPVPDQWEVLNIYMKAHKLSPSDECLMPNLNEGKLVSLCHELKLL
YVAITRTRQRFWILDEIWNPMLELWKQKGIVKLQQFDSEVLQIMQAPSTREAWKSRGKKFFHLQNYETARMCFKKAGESYWEKYCEASELRSNADQMRSS
DPVVSRRHLSDAALIYAKIGKDALAAECYFELQDYVKAAEIYLEKFGASKLQEAGECFCLAKCYRRAAEVYYKCNAYVKCLSACIDGKCYETGYDYIQQW
RKDGVFNFTEFVEFLKKGALHFHRLENDEFMIMFVRALNSKDQIRSFLRSRHLLDYLLRLEEEWANYLEAAAIAMDIGLETVAANNHERGGQYQEACHVI
LRHVLGESLWGDANKGWPVREFEGKTKLLNQAKTCAEHVSAAYHGFVCQETFMLSYANGNSLWKLKNSFEYFQRQGNIRNEILSARSVIDIFIDSSASGN
FRDDVLVADILNYTEDWRMPWKKLSFLNLYDYWKSWKGHIDKIVAYCVEKGHWGCHGEFCMDYLGVREQQSDKETEYNVLYPDAAWINSMISSKVVQNKR
KRVKVNADQFWKAAKDYWFTERRVVGEKVLHKLENVYEYCVRNGLSVFCRSMLLVKIFELSRSLFSFKLGNPVYQYTAEACYWTDLVRRFIEHYTENILS
IHFLEPNESFRKHMIIARTLEGFKVPFEEAVCYYIENLPHGVTYRQIAEVTAVALSGKLHHETLQMLVQTLKKKAVCSSCIEETMGPSKEATVRKLHEVV
EHMYYAEFEKADETMSPNCLVYLLDRLVILLLAVKGGFYCPRSSVVEWLMHNGWHYTCSTSTSDRYEQPFVEEALDSVSFIVKDIFCNERDCKTPRWTKS
FNEKLESYPFVVLRLVAIVCILNINTGKCSDLLSNILSRREITSLSPLEFYSCLTRTRCQDETNHAVVSFSKALAEINNPLVYVDLKGSGSTRFKPPGAP
IIFLDTKVSGRQDILNRVCPRNMSASANSV
|
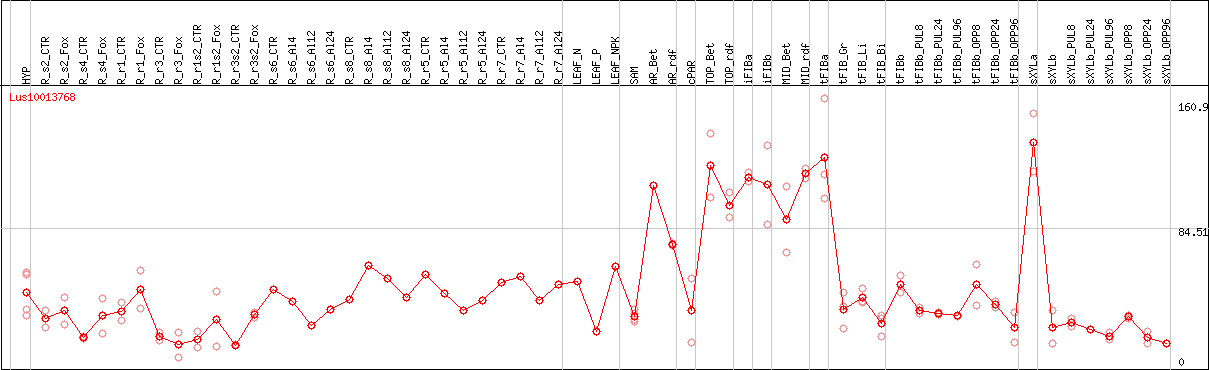
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013768 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
