External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G36870 429 / 9e-153
XTH32
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
AT3G44990 359 / 3e-125
AtXTH31, XTH31, ATXTR8, XTR8
XYLOGLUCAN ENDOTRANSGLUCOSYLASE/HYDROLASE 31, xyloglucan endo-transglycosylase-related 8 (.1)
AT1G10550 208 / 8e-66
XTH33, XET
xyloglucan:xyloglucosyl transferase 33 (.1)
AT1G32170 200 / 4e-62
XTH30, XTR4
xyloglucan endotransglycosylase 4, xyloglucan endotransglucosylase/hydrolase 30 (.1)
AT4G14130 198 / 6e-62
XTR7, XTH15
xyloglucan endotransglycosylase 7, xyloglucan endotransglucosylase/hydrolase 15 (.1)
AT1G14720 199 / 7e-62
ATXTH28, EXGT-A2, XTR2
xyloglucan endotransglycosylase related 2, ENDOXYLOGLUCAN TRANSFERASE A2, xyloglucan endotransglucosylase/hydrolase 28 (.1)
AT5G57550 195 / 8e-61
XTR3, XTH25, EXGT-A5
xyloglucan endotransglycosylase 3, xyloglucan endotransglucosylase/hydrolase 25 (.1)
AT2G01850 195 / 3e-60
ATXTH27, EXGT-A3
XYLOGLUCAN ENDOTRANSGLUCOSYLASE/HYDROLASE 27, endoxyloglucan transferase A3 (.1)
AT3G23730 191 / 3e-59
XTH16
xyloglucan endotransglucosylase/hydrolase 16 (.1)
AT3G25050 190 / 6e-59
XTH3
xyloglucan endotransglucosylase/hydrolase 3 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026535
536 / 0
AT2G36870 444 / 9e-159
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
Lus10001396
487 / 2e-175
AT2G36870 460 / 8e-165
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
Lus10023009
482 / 1e-173
AT2G36870 466 / 3e-167
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
Lus10021422
377 / 4e-132
AT2G36870 417 / 4e-148
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
Lus10016144
372 / 4e-130
AT2G36870 416 / 1e-147
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
Lus10037377
372 / 4e-130
AT2G36870 422 / 9e-150
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
Lus10041341
371 / 1e-129
AT2G36870 421 / 2e-149
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
Lus10026536
358 / 3e-124
AT2G36870 402 / 1e-141
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
Lus10013823
280 / 3e-95
AT2G36870 301 / 7e-104
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G122900
454 / 1e-162
AT2G36870 478 / 4e-172
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
Potri.016G098600
446 / 2e-159
AT2G36870 477 / 1e-171
xyloglucan endotransglucosylase/hydrolase 32 (.1.2)
Potri.009G006600
374 / 6e-131
AT3G44990 407 / 3e-144
XYLOGLUCAN ENDOTRANSGLUCOSYLASE/HYDROLASE 31, xyloglucan endo-transglycosylase-related 8 (.1)
Potri.014G115000
211 / 9e-67
AT1G10550 367 / 9e-128
xyloglucan:xyloglucosyl transferase 33 (.1)
Potri.014G146100
202 / 1e-63
AT3G23730 451 / 1e-161
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Potri.003G097300
202 / 7e-63
AT1G32170 436 / 8e-154
xyloglucan endotransglycosylase 4, xyloglucan endotransglucosylase/hydrolase 30 (.1)
Potri.009G163850
199 / 1e-62
AT2G01850 319 / 6e-109
XYLOGLUCAN ENDOTRANSGLUCOSYLASE/HYDROLASE 27, endoxyloglucan transferase A3 (.1)
Potri.011G077320
198 / 7e-62
AT3G23730 386 / 7e-136
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Potri.006G071200
196 / 2e-61
AT4G25810 379 / 3e-133
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
Potri.001G136100
197 / 6e-61
AT1G32170 469 / 6e-167
xyloglucan endotransglycosylase 4, xyloglucan endotransglucosylase/hydrolase 30 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0004
Concanavalin
PF00722
Glyco_hydro_16
Glycosyl hydrolases family 16
CL0004
Concanavalin
PF06955
XET_C
Xyloglucan endo-transglycosylase (XET) C-terminus
Representative CDS sequence
>Lus10013822 pacid=23159481 polypeptide=Lus10013822 locus=Lus10013822.g ID=Lus10013822.BGIv1.0 annot-version=v1.0
ATGGCTTCTTCTTCCTCCCTCATTCTTCTTATTCTTTTGGCTCTTTCTTCATTCATCGCCATTGTTAATGCCCGGCCGCCCCGAACCGGGTTCCGACCCA
GCTTCAGGTTTAGATCATTCCCCTTCTACAAAGGGTATAATACCCTTTGGGGCCAGTCCCACCAAAGAGTGGACAGCAATGGAGTCACCATTTGGCTTGA
TAAAACTTCAGGAAGTGGGTTCAAGTCAGTGAAGCCATTTCGGTCTGGCTATTTCGGTGCCTCGGTCAAGCTCCAACCTGGCTACACTGCTGGAGTTAAC
ACTGCCTTCTATCTTTCAAACAGTGAAGCTCACCCAGGGAGCCATGATGAGGTGGACATTGAATTCCTTGGCAAGACATTCGACGAGCCTTACACTCTAC
AGACCAACGTTTACATTAGGGGAAGCGGGGATGGCAAAATCATTGGCAGAGAGATGAAGTTCCATTTATGGTTCGATCCCACCAAGAATTTCCACCACTA
TGCCATCATCTGGAATCCCAGAGAAATCATATTCCTAGTGGACGATGTGCCAATAAGGAGGTACCAAAAGAAAAGCGCCGCAACATTCCCAATGAGGCCC
ATGTGGGTATATGGCTCGATATGGGATGCCTCGGCTTGGGCTACTGACGATGGCAAGTACAAAGCCGATTATCGGTACCAACCCTTTGTCGCTAACATGA
AGAATTTCAAGGCCACAGGCTGCACCGCCTACTCGCCGAGGTGGTGCAGGGCGCCCACCGCCTCACCTGACCGGTCCGGTAAGCTAACGAGGCAGCAATA
CAATGCCATGAATTGGGTTCAAAAGTACCATATGGTGTACAATTATTGTTCGGACAAAGCAAGAGACCATTCGAAGACCCCCGAATGTTGGTTATGA
AA sequence
>Lus10013822 pacid=23159481 polypeptide=Lus10013822 locus=Lus10013822.g ID=Lus10013822.BGIv1.0 annot-version=v1.0
MASSSSLILLILLALSSFIAIVNARPPRTGFRPSFRFRSFPFYKGYNTLWGQSHQRVDSNGVTIWLDKTSGSGFKSVKPFRSGYFGASVKLQPGYTAGVN
TAFYLSNSEAHPGSHDEVDIEFLGKTFDEPYTLQTNVYIRGSGDGKIIGREMKFHLWFDPTKNFHHYAIIWNPREIIFLVDDVPIRRYQKKSAATFPMRP
MWVYGSIWDASAWATDDGKYKADYRYQPFVANMKNFKATGCTAYSPRWCRAPTASPDRSGKLTRQQYNAMNWVQKYHMVYNYCSDKARDHSKTPECWL
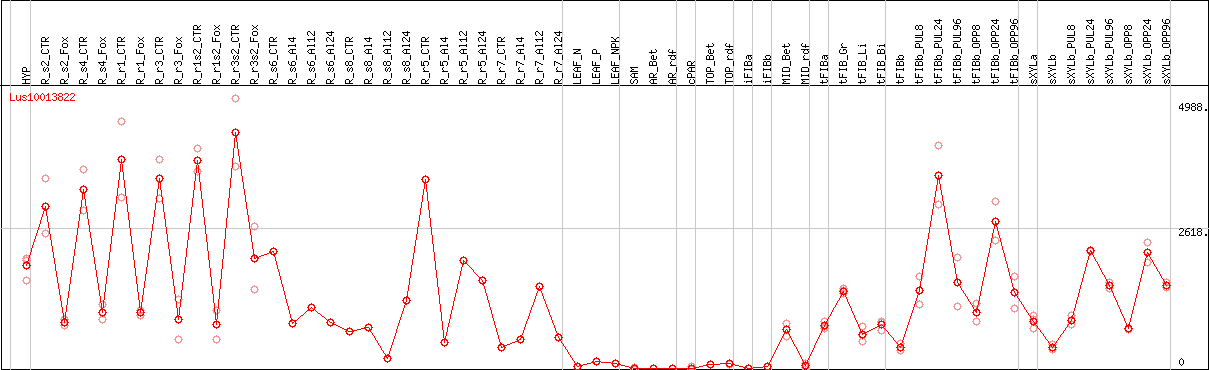
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013822 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.