Lus10013832 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10013832 pacid=23159440 polypeptide=Lus10013832 locus=Lus10013832.g ID=Lus10013832.BGIv1.0 annot-version=v1.0
ATGAGGTTTTGCTGCTGGTCTTCTTCACCCACCTCTCCTACGCCTACCTCCACTGGTCGTCTCAGTTCAGGAGGCGTGTCTCATACACCTAGCAGCAGCA
ATGCAACTTCATCAGGGAGTGGCAATACTAATTCTTCAAAGGGAAGCAATGTCTCTGGGAACAGTCAGTTTTCTGCATCCAGTGGCGATGACGCTTTTCC
CAATGGGCAGCTCTTGCCCACCCCACACTTGAGGATCTTCTCTTTCACCGAGCTTAAGCTCGCCACTAAGAACTTCAGGGCTGATACTGTGCTTGGAGAA
GGAGGGTTTGGTAAAGTTTACAAAGGTTGGCTTGATGAGAGGAAGCCTGGGAAATCTGGAAGTGGAACTGTGGTTGCTGTCAAGAAACTTAATTCTGAGA
GCTTGCAAGGATTGGAAGAATGGCAGTCAGAAATAAACTTCTTGGGAAGGCTTTCACACCCGAACCTTGTCAGGCTGTTGGGATACTGCTTGGAAGAGGA
TGAGCTTCTCCTTGTATATGAATTCATGCAGAAAGGAAGCTTGGAAAACCATCTATTTGGAAGGGGTGCAGCTGTTCAGCCACTTCCATGGGACACTCGG
TTGTCGATCGCCATAGGAGCTGCTCGAGGCCTTGCTTTCTTACATACACCGGATAAGAAAATCATTTATAGAGACTTTAAAGCCTCAAATATATTGCTGG
ATGGGTCTTACACATCCAAGATATCAGACTTCGGTTTGGCAAAGTTCGGTCCTTCGGCTAGTCAATCGCATGTGACTACTCGTGTCATGGGCACATACGG
TTATGCTGCTCCCGAGTATGTTTCCACAGGTCATCTATACGTGAAGAGCGATGTGTACGGGTTCGGAGTTGTGCTGGTCGAGATCTTGACAGGGTTAAGG
GCGCTCGACACAAGCAGACCGAGTGGGAGGCATAGCATAGCGGAATGGATCAAGCCTTATCTAAATGACAAGAGGAAGCTGAAAAGCATGATGGACACTA
ACTTGGAAGGGAAGTATCCTTCAAAAGCAGCATTCAGAATAGCACAGCTAGCTCTAGTTTGTCTGGAATCAGAGCCGAAAAATCGACCCCACATGAGGGA
AGTGGTGGAGACGTTGGAAAGGATTGCATCGAACAACGAGAGGACACTAGAGCCCCGATCACACCGTTCGAGCAACAACCGGCCTGTCGCTCACCGGAAT
GGCCAGAGACCGTTGCAGCTCCGCTCTCCACTGCATCAACAGCAAGATGCTGGTCAAGATTTGTTGCCATTGTTTTTTCTTGACAGGGAGCTGAAATCAC
TCCGCTCCTTCTCTGATTGTCAAGCGTGTGACGACACTAAGGATATTGACATTATTGCTACGTACAGTGATCCTTCTGGCAATGTGCGAGTTAGTAGGGT
TAAAATGAAAAACCTTTCTGCCTCGTGGGTGCTTGAGAACCCTGACGATGAGCATGATCGGCGTAACAGTAGTTCCAAGACATTGGAACATGGTAACTTC
CCGAAAGATAAGCCTCGGAAGCAAGTGATTCGTCAGCGGTTTGAAGGAGGCAAAACTCAGTTCCCCCTGAAATCTGTGATGAGTCCAGTGAAGCTCAAGC
GTCGGATTGCGCGGCAAAAGAGGAAAGATCTCCGAATTGCTGAGCTAATCCAACATGACAACGAAGCGGATAACCAGGCACGGTCAGCTGCAATAGAGCG
GTCTAAAGCACTGGACACGACTATCAGAGGGAAGTACAGTATATGGAGGAGAGATTTCGAAAATCCAAATTCCGATGCCATGTTGAAGCTCATGCGGGAC
CAGATGATAATGGCCAAGGCATATTCAAATATTGCCAAATCTAACAATGAAACAGGTCTTTACATTTCTCTGATGAAGAACTCCAAAGACAGTCAGCTCG
CCATTGGAGAAGCAACATCTGATGCTGAGCTTCATCCAAGTGCTCTTGATCGAGCTAAAGCTATGGGACGTGTTCTCTCTATGGCCAAGGACCAACTTTA
TGATTGTCCTACAATGTCAAGGAAGTTAAGGGCTATGCTGCAGTCTAGTGAGGAGCATTTGAATGCTGTGAGGAAAAAAAGCGCCTTCTTGATTCAGCTG
GCTGCTAAAACAGTTCCAAAGCCATTGCATTGCCTTCCTTTGCAGCTGGCTGCTGACTATTTCCTGCAGGGTCATCAGAACAAACAATATTTCAACAGTG
GAAAGCTTGAAAATCCTAAACTTTACCACTACGCCATATTCTCCGACAATGTGCTAGCCACATCCGTGGTTGTCAATTCCACTGTGCTGCATGCAAAAGA
ACCTCAGAAACACGTTTTCCATGTCGTAACAGATAAGTTGAACTTCGCGGCAATGAAAATGTGGTTCATTGTGAACCCTCCAGCACGAGCAACAATCGAG
GTGATGAACATAGATGATTTTAAGTGGTTAAATTCATCTTACTGTTCTGTCCTACGGCAACTGGAATCTTCCAGGGTTAAGGAGTACTATTTCAAGGCCA
ACCACCCGTCGTCTTTGTCAGCTGGATCTGACAATCTCAAGTACAGGAATCCAAAATACTTGTCTATGCTGAATCATCTTAGATTCTATCTTCCAGAAGT
CTATCCGAAGCTGGACAAGATCCTGTTTCTGGACGATGACATTGTCGTGCAAAAAGATTTGGCACCACTTTGGTCAGTTGATCTACGAGGAATGGTTAAT
GGCGCAGTGGAGACATGTAAGGAGAGCTTCCACAGGTTTGATAAGTATCTGAACTTCTCGAACCCGAAAATCAACGAGAACTTCCATCCAAATGCTTGCG
GGTGGGCATTCGGGATGAACATGTTCGATTTGAAGGAATGGAAGAAGCAGAACATCACTGGCATCTACCATTACTGGCAAGACCAGAATGAGGATAGAAC
TCTTTGGAAGTTGGGGACGTTGCCGCCTGGACTGATAACGTTCTATAACTTGACGTATCCGTTGGATCGAGGGTGGCATGTGTTGGGACTTGGATATGAT
CCTGCGTTGAACCAAACGGAGATAGAGAATGGAGCAGTGGTGCATTACAATGGGAACTACAAGCCGTGGTTGGATTTGGCAATTGCCAATGCAACATCAG
TGAATAAAAAACCAGCAGCCGGCCGGCCAACCACACTGCTCCCCTTATTACTGGCGGGGAAGCTCATATTCTTTGATCAGTTGAAGAAAATCTCAGGGTT
GCCGGAAGAACGAGCAGGTGGGGAAACACCGAAATTTCAACACCGGGAATCAATGGAGCCAACTGCCAAGCTCACCAGAGCACTCCTCGTCAACACCTCC
AACCCAAAGCTAGCATGGCATCTTTTCAAGCGCATTCTTACTCTACCTTCCCTTGACCATTTCCCCCATTCCGTTGAGACCATTACTCGAATCCTAGTCC
GCGCCAAACTCCTTAACGAGCTCGATGATCTTCACGTACGCCTGCATTCTAACCTGCCTGCACACATCTTACATCCTTGTCTTACTTCTTCGATTACAGT
CTCAGCTCAATCGGGCCAATTCGGTAAGGCCCTTTCTCATTTCAAAGCAATGCGTTTCCGGTTCCCAGATGAGCCGCCTTCCGTACGTCTGTATAATTGT
CTGATTAGGTCTTGCATCAAAGACGGGAGCGTCGAACACGTCGGTTGGCTGTATAAAGATATGATTGTAGCTGGGGTTTCTCCGGAGACTTTTACTTTCA
ATATTTTGATTGGTTTCCTTTGCAATGGTGGTCGTTTAGAAGAAGCACAGGAGCTGTTTGATAAAATGTCTGAGAAGGGATGTGAACCGAACGAGTTCAC
TTATGGGATTTTGGTTCGCGGGTATTGCAGAGCTGGGGACTCTACACGTGGATTGGAGCTTATGAGTGAAATGACGAGTCGAGGATTACTTCCTAACAAC
GTTGTGTATAATACTCTCATTTCTACATTCTGCAAGCAAGGTAAGAATGATGATGCCGAAAAATTGGTAGAAAGAATGAAAGAGGAAGGACTGCATCCTG
ATGTTGTGACTTTCAATTCTAGGATCTCAGCTCTTTGCGACGCTGGGAAAACTCTGGATGCTTGTAGGATTTTTAGAGATATGCAAAATGATCTACACTT
GGGGCTGCCGAGGCCCAATATCGTCACGTATAATTTGATGCTTAAGGGTTTCTTTAAGGAAGGACAGGTGGAGGATGCTAAGTCTCTGTTTGAGATTATG
AAGAACAATCGTGTGTTGGTGAATATAGAGAGTTATAATATTTGGTTGGCAGGTATGTTCAGGAACAGGAAGAATTTTGAGGCTCACCTAGTTTTTAAAG
AAATGATAGATATGGGAATCAAACCCAACATATGTTCTTGTAACATAATGATAGATGGGCTATGCAAAAATGGAATGTTCTCTGATGCGAGAACGTTGAT
GAAAGAAATGATAGTCAATCAGATTTTTCCAGATGCAGTAACTTATAGTACTTTACTTCATGGTTATTGCAGTCGAGGAATGGTCTTGCAAGCTAACAGT
GTTTTGCATGAGATGATGCAGAATGGTTGTAATCCAACGACTTACACGTGCAATATCTTACTGCACAGTCTTTGGAAGGAGGATAAAATACCTGAAGCTG
AGAAGCTGCTTCAGAAGATGCACGAGAGAGGCTATGGTGTGGATACTGTGACATGTAATATTATACTTGAAGGTCTTTGTAATCATGGGCAGCTGGATAA
GGCGATTGATGTTGTGAATGGTATGTGGGCACATGGAAGTGCTGCCCTTGGTAACTTAGGGAATTCTTTCATTGGCCTCATAGATGGTGAGAATAATGCG
AGAAAATGTGTGCCTGATTTGATCAGCTATTCAATTGTAATATCTGGATTATGCAAGGCTGGTAGAGTTGATGAGGCTAAGAAGAAGTTCCTTGAGATGA
TGGGAAAACACTTGCAGCCTGATTCGGCCATTTATGATACTTTCATTCGTAGCTTCTGTAAAGAAGGGAAGTTATCTGCTGCATTTCGAGTGCTTAAAGA
CATGGAGAAAAGAGGTTGCAATAAAACCATACATACATTCAACTCCTTGATCCTGGGTCTAGGAAGAAAAAATCAACTATTTGAAATCCACGGTTTGATG
GATGAGATGAAAGAAAGAGGGATTTCTCCAGATGTTTACACATATAACAATCTTCTCAGTTGTCTATGCGAAGGAGGGAGAACTAAGGACGCTCCATCCG
TTCTGGAGGAGATGCTGCAAAAGGGTATTTCTCCAAATGTTTCTTCTTTCAGAACATTACTTGCAGCTTTCTGCAAGGGTAGTGATTTCAATGCAGCACA
TGAGGTGTTTGAGATTGCTCTGGATGTGTGTGGCCACAAGGAAGCCCTTTATAGCTTAATGTTTAATGAGTTAGTTTCCGGTGGATTAGTTGCTGAAGCT
AGACAGCTCTTCGAGACTGTGCTAGATAGGTCTTTTGACATGGCAAACTTCCTCTACAAAGATCTCATTGATAGACTTTGTAAGAATGACAAGTTGGAAG
ATGCCAGTTGTATACTACATAAGTTGATGGATAAAGGATATAAATTCGATCCAGCTGCATTCATGCCAGTAATTGAAGGCCTAGGCCGTGTGGGAAACAA
GGGTGAAGCTGAAGAACTTTCAGAGAGAATGATGGAGATGGTTTCGGAGGATAATGTGAAAAACAGGGCTGGTGCCTCTGGAACGAGAAATAAGGATGAG
GGGACTGATTGGAAAACGATAATACAGAGGGATGACGGAAGTGGAGTTGCACTGAAAGCTCTTAAGCAGGTACAGAAGGGATGGGGTCGGGGATGCCTTT
TAGGTAAACGACCTCAGGAAGATGACTCTACCTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10013832 pacid=23159440 polypeptide=Lus10013832 locus=Lus10013832.g ID=Lus10013832.BGIv1.0 annot-version=v1.0
MRFCCWSSSPTSPTPTSTGRLSSGGVSHTPSSSNATSSGSGNTNSSKGSNVSGNSQFSASSGDDAFPNGQLLPTPHLRIFSFTELKLATKNFRADTVLGE
GGFGKVYKGWLDERKPGKSGSGTVVAVKKLNSESLQGLEEWQSEINFLGRLSHPNLVRLLGYCLEEDELLLVYEFMQKGSLENHLFGRGAAVQPLPWDTR
LSIAIGAARGLAFLHTPDKKIIYRDFKASNILLDGSYTSKISDFGLAKFGPSASQSHVTTRVMGTYGYAAPEYVSTGHLYVKSDVYGFGVVLVEILTGLR
ALDTSRPSGRHSIAEWIKPYLNDKRKLKSMMDTNLEGKYPSKAAFRIAQLALVCLESEPKNRPHMREVVETLERIASNNERTLEPRSHRSSNNRPVAHRN
GQRPLQLRSPLHQQQDAGQDLLPLFFLDRELKSLRSFSDCQACDDTKDIDIIATYSDPSGNVRVSRVKMKNLSASWVLENPDDEHDRRNSSSKTLEHGNF
PKDKPRKQVIRQRFEGGKTQFPLKSVMSPVKLKRRIARQKRKDLRIAELIQHDNEADNQARSAAIERSKALDTTIRGKYSIWRRDFENPNSDAMLKLMRD
QMIMAKAYSNIAKSNNETGLYISLMKNSKDSQLAIGEATSDAELHPSALDRAKAMGRVLSMAKDQLYDCPTMSRKLRAMLQSSEEHLNAVRKKSAFLIQL
AAKTVPKPLHCLPLQLAADYFLQGHQNKQYFNSGKLENPKLYHYAIFSDNVLATSVVVNSTVLHAKEPQKHVFHVVTDKLNFAAMKMWFIVNPPARATIE
VMNIDDFKWLNSSYCSVLRQLESSRVKEYYFKANHPSSLSAGSDNLKYRNPKYLSMLNHLRFYLPEVYPKLDKILFLDDDIVVQKDLAPLWSVDLRGMVN
GAVETCKESFHRFDKYLNFSNPKINENFHPNACGWAFGMNMFDLKEWKKQNITGIYHYWQDQNEDRTLWKLGTLPPGLITFYNLTYPLDRGWHVLGLGYD
PALNQTEIENGAVVHYNGNYKPWLDLAIANATSVNKKPAAGRPTTLLPLLLAGKLIFFDQLKKISGLPEERAGGETPKFQHRESMEPTAKLTRALLVNTS
NPKLAWHLFKRILTLPSLDHFPHSVETITRILVRAKLLNELDDLHVRLHSNLPAHILHPCLTSSITVSAQSGQFGKALSHFKAMRFRFPDEPPSVRLYNC
LIRSCIKDGSVEHVGWLYKDMIVAGVSPETFTFNILIGFLCNGGRLEEAQELFDKMSEKGCEPNEFTYGILVRGYCRAGDSTRGLELMSEMTSRGLLPNN
VVYNTLISTFCKQGKNDDAEKLVERMKEEGLHPDVVTFNSRISALCDAGKTLDACRIFRDMQNDLHLGLPRPNIVTYNLMLKGFFKEGQVEDAKSLFEIM
KNNRVLVNIESYNIWLAGMFRNRKNFEAHLVFKEMIDMGIKPNICSCNIMIDGLCKNGMFSDARTLMKEMIVNQIFPDAVTYSTLLHGYCSRGMVLQANS
VLHEMMQNGCNPTTYTCNILLHSLWKEDKIPEAEKLLQKMHERGYGVDTVTCNIILEGLCNHGQLDKAIDVVNGMWAHGSAALGNLGNSFIGLIDGENNA
RKCVPDLISYSIVISGLCKAGRVDEAKKKFLEMMGKHLQPDSAIYDTFIRSFCKEGKLSAAFRVLKDMEKRGCNKTIHTFNSLILGLGRKNQLFEIHGLM
DEMKERGISPDVYTYNNLLSCLCEGGRTKDAPSVLEEMLQKGISPNVSSFRTLLAAFCKGSDFNAAHEVFEIALDVCGHKEALYSLMFNELVSGGLVAEA
RQLFETVLDRSFDMANFLYKDLIDRLCKNDKLEDASCILHKLMDKGYKFDPAAFMPVIEGLGRVGNKGEAEELSERMMEMVSEDNVKNRAGASGTRNKDE
GTDWKTIIQRDDGSGVALKALKQVQKGWGRGCLLGKRPQEDDST
|
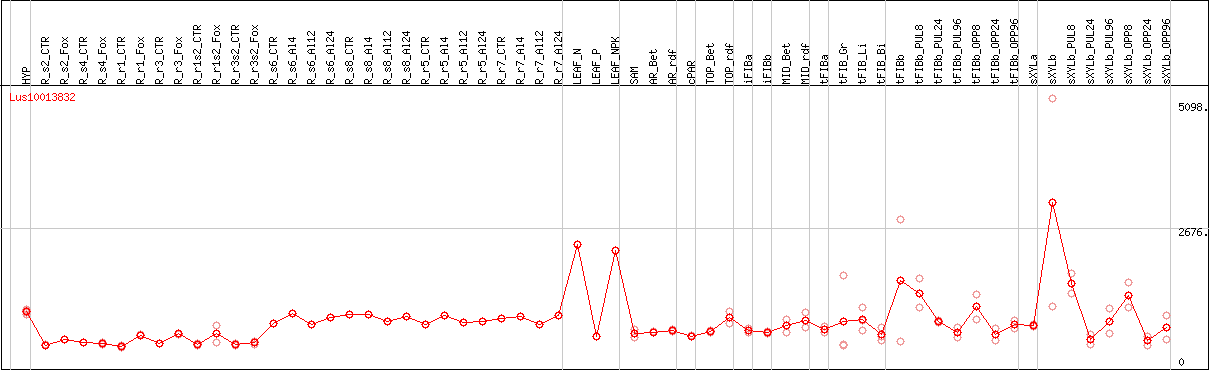
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013832 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
