External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G38160 446 / 3e-158
PDE191
pigment defective 191, Mitochondrial transcription termination factor family protein (.1.2.3)
AT1G78930 131 / 4e-34
Mitochondrial transcription termination factor family protein (.1)
AT2G21710 118 / 2e-29
EMB2219
embryo defective 2219, Mitochondrial transcription termination factor family protein (.1)
AT2G44020 115 / 1e-28
Mitochondrial transcription termination factor family protein (.1)
AT4G02990 112 / 1e-27
RUG2, BSM
RUGOSA 2, BELAYA SMERT, Mitochondrial transcription termination factor family protein (.1)
AT5G55580 107 / 1e-25
Mitochondrial transcription termination factor family protein (.1)
AT3G18870 99 / 6e-24
Mitochondrial transcription termination factor family protein (.1)
AT4G14605 98 / 1e-22
Mitochondrial transcription termination factor family protein (.1)
AT5G54180 82 / 3e-17
PTAC15
plastid transcriptionally active 15 (.1)
AT2G36000 78 / 3e-16
EMB3114
EMBRYO DEFECTIVE 3114, Mitochondrial transcription termination factor family protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026571
397 / 6e-140
AT4G38160 252 / 4e-83
pigment defective 191, Mitochondrial transcription termination factor family protein (.1.2.3)
Lus10035714
136 / 6e-36
AT1G78930 582 / 0.0
Mitochondrial transcription termination factor family protein (.1)
Lus10014877
128 / 4e-33
AT4G02990 665 / 0.0
RUGOSA 2, BELAYA SMERT, Mitochondrial transcription termination factor family protein (.1)
Lus10035227
125 / 6e-32
AT4G14605 518 / 0.0
Mitochondrial transcription termination factor family protein (.1)
Lus10032061
123 / 2e-31
AT4G14605 521 / 0.0
Mitochondrial transcription termination factor family protein (.1)
Lus10003775
122 / 6e-31
AT5G55580 546 / 0.0
Mitochondrial transcription termination factor family protein (.1)
Lus10022321
119 / 5e-30
AT4G02990 540 / 0.0
RUGOSA 2, BELAYA SMERT, Mitochondrial transcription termination factor family protein (.1)
Lus10004614
117 / 3e-29
AT2G44020 707 / 0.0
Mitochondrial transcription termination factor family protein (.1)
Lus10042322
117 / 7e-29
AT2G21710 720 / 0.0
embryo defective 2219, Mitochondrial transcription termination factor family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G209400
526 / 0
AT4G38160 481 / 3e-172
pigment defective 191, Mitochondrial transcription termination factor family protein (.1.2.3)
Potri.009G170300
518 / 0
AT4G38160 479 / 8e-170
pigment defective 191, Mitochondrial transcription termination factor family protein (.1.2.3)
Potri.007G001800
137 / 5e-36
AT1G78930 633 / 0.0
Mitochondrial transcription termination factor family protein (.1)
Potri.014G137400
120 / 2e-30
AT4G02990 669 / 0.0
RUGOSA 2, BELAYA SMERT, Mitochondrial transcription termination factor family protein (.1)
Potri.009G116200
119 / 1e-29
AT2G21710 748 / 0.0
embryo defective 2219, Mitochondrial transcription termination factor family protein (.1)
Potri.001G361800
115 / 2e-28
AT5G55580 584 / 0.0
Mitochondrial transcription termination factor family protein (.1)
Potri.013G116700
114 / 4e-28
AT2G44020 764 / 0.0
Mitochondrial transcription termination factor family protein (.1)
Potri.017G067600
103 / 1e-24
AT4G14605 571 / 0.0
Mitochondrial transcription termination factor family protein (.1)
Potri.004G150600
98 / 2e-23
AT3G18870 311 / 1e-106
Mitochondrial transcription termination factor family protein (.1)
Potri.016G072200
83 / 9e-18
AT2G36000 280 / 1e-92
EMBRYO DEFECTIVE 3114, Mitochondrial transcription termination factor family protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02536
mTERF
mTERF
Representative CDS sequence
>Lus10013854 pacid=23159448 polypeptide=Lus10013854 locus=Lus10013854.g ID=Lus10013854.BGIv1.0 annot-version=v1.0
ATGGCGTGTAATCAAAACCCTACCAGCGTGATGTGGTTCTTTCGGGACAGAGGCTTCGACGAGAAAAGCATCAACGACATGTTCACAAAGTGCAAGCGTC
TTGAAGGTGCTCACAAGGATAGAGCATCTGAGAATTGGGATTTCTTGAAAAGCATTGGCATTCAAGAAAGGAAGCTCCCTTCCATAATCTCCAAGTGCCC
CAAACTCCTAACCCTCGGCCTCCACGAGAGGCTCGTCCCTATGGTGGAGTGTCTCGCCACGCTCGGTACGAAACCACGCGAGGTTGCATCGACAATCACC
AAGTTCCCTCACATACTCTCTCATAGCGTTCAAGAGAAGCTATGCCCTCTATTGGCATTCTTTCAAGCACTGGGGATTTCTGATAAACAAGTTGGGAAGC
TGATAATGCTCAATCCTAGGCTGGTTAGCTACAGCATTGATACCAAGATGAAAGAGCTTGTTGACTTCCTTGTTGGTTTGGGGCTCACCAGAGATGGAAT
GATCGGTAGAGTTCTTGTGAAGCACCCTTTCATATTGGGGTACAATATAGAGAAAAGGCTGCGGCCAACCTGCGAGTTCTTAAGATCGGTTGGCCTCTCA
GCTTCTGATCTTCAGACTGTTGTTTTGAAGTTCCCTGATGTTGTGTGTAGAGATGTGGAGAAGACTTTGAAACCGAATTTTGCTTATCTGAGGGGATGTG
GGTTTGGAGATGGACAGTTGGTTGCTTTGGTGACTGGTTACCCTCCCATTCTGATCAAGAGCATCCAGAATTCTTTAGAACCTAGGATGAAGTTCTTGGT
TGATGTTATGGGGAGACAGCTTGATGAAGTTGTGGAGTACCCTAACTTCTTCCAGTGTGGGTTGAAGAAGACGCTGGAGTCGAGGCACAGATTTTTGAGG
CAGAGGAATGTAGAATGTAGCTTGAGCGATATGTTGAGCTGTAATCATAAGAAGTTCTTCATGAAATATGGGTCCGGTTGA
AA sequence
>Lus10013854 pacid=23159448 polypeptide=Lus10013854 locus=Lus10013854.g ID=Lus10013854.BGIv1.0 annot-version=v1.0
MACNQNPTSVMWFFRDRGFDEKSINDMFTKCKRLEGAHKDRASENWDFLKSIGIQERKLPSIISKCPKLLTLGLHERLVPMVECLATLGTKPREVASTIT
KFPHILSHSVQEKLCPLLAFFQALGISDKQVGKLIMLNPRLVSYSIDTKMKELVDFLVGLGLTRDGMIGRVLVKHPFILGYNIEKRLRPTCEFLRSVGLS
ASDLQTVVLKFPDVVCRDVEKTLKPNFAYLRGCGFGDGQLVALVTGYPPILIKSIQNSLEPRMKFLVDVMGRQLDEVVEYPNFFQCGLKKTLESRHRFLR
QRNVECSLSDMLSCNHKKFFMKYGSG
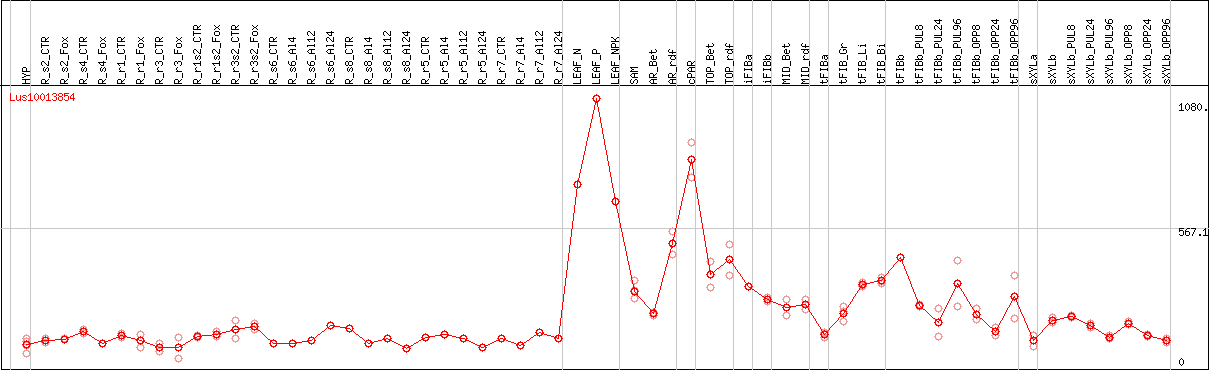
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013854 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.