Lus10013904 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10013904 pacid=23170071 polypeptide=Lus10013904 locus=Lus10013904.g ID=Lus10013904.BGIv1.0 annot-version=v1.0
ATGCCGCTGGTTAGGTTTGAGGTCAGGAACCAGTACGCTTTGGGCCATCCGGAGCTCTACAAGGAGGCTGACGTGGATGATCCCAAGGCCGTTCTCGACG
GTGTTGCCGTCGCCGGTCTCGTCGGGATCTTGCGCCAATTAGGCGATCTCGCTGAATTTGCAGCAGAGGTGTTTCACGGATTGCAGGAGCAGCTAACGAC
GACAGCTTCTCGGAGTCAGAAGTTGAAGCTTCGAGTGCAGAACATTGAAGCTGCGTTGCCTCCTATTGAGAAGGCTATATTGGCACAGAATAGCCACATC
CATTTTGCTTACACTGCTGGTTCTGAGTGGCATCCTAGAATCCGGAGTGAGCAGAATCACTTCATCTACAATGACTTGCCCCGTTTCATTATGGATTCTT
ATGAAGAATGTCGTGATCCGCCACGCTTTCATATACTTGACAAATTTGACACTGGTGGTCCAGGGTCGTGTTTGAAGAGGTATTCAGATCCAACCTTCTT
TCGAAGAGCGTCAGGTGGCTTCAAGGAACATGATGCTGAGAAAGTTCAAAGAGATAAAAAGGCTCGCAAAATCAAGAAAAAGAGATCACTTCGAAGGAAA
GGTAGCTCACTGCAGAATGGGTCAATCTCAAGTCACAGCGAGAGATCCCAGTTTTTTCAGCCAACAGTTAATGGACGGAGTTCTTCTTCACATACGGCCT
CTAATATCGACTTGCCATTGAAATCTGAGATTGGTGATCGTTCAAATTCTATTGATTCAGGAAATGGGTCAAATCATATTGAATGTGTTTTCCGTCCTAG
TTCTTCTGCACAACCTGAGGAACAGAAGTCTAATGGATTCTCGTCAAGGTCATTCCAGGGTGATGATTCCCGTGATTCTGGCTTTTCGGAGGGACAGTCC
AGGGTTGCTCATGATAGTTTTCTGCATTGTTCCTCACCAAAGCAGATTCACCATAGGTCTCCTGGTGTTACTTGGGTTGAAAAGGAAGAATGCGAAGAGC
CTGGAGTTGCTGATGATAGCTTTCTGCATTGTTCCTCACCAAAGCAGATTCACCATCGGTCCCCTGGTGTTACTTGGGTTGGAAAGGAAGAACACGAAGA
GCCTGGAGTTGCTGATGATAGTTTTCTGCACGTTTCCTCACCGGAGCTGATTCCCATTAGTTCCTCTTGTCGTACCTGGGAAGAAAAGGCAGAAGGTGCA
CAGCCTGAAGCTCATAAAGGTTCTCTACCTAGGTCCTCACCAGAGCGGCTCCCCCCTGGTTTCTCCTGTGTCTCTTGGGATGAAAAAGAAGAATTATCAG
AGGCTGAAAGCCAACACTATGACGGAGATGAAGCTCCAGATGTTTTGACTGTAGACTCTGTCGTAGATATGCAAGAATCAGGGACCAATACTGAAATTCA
TGATCCAACAGATCTCAGATTTGAGAATGAGCATGTTCCAGGGCCAAGTTTCACTGCTGCCACCCTGCTTGAAGATGTTGAAAGTGAAACTGACAATTTC
ATGGACGCTCTAAATTCAATAGATACAGAATCTGAGAGCGAACTTGATTATCAGACGAAACATGAAGTGAAAGAAGTTGAATGTGCTAACAGGGATAATG
AGGGCTTAGCAGTTACTTCAGGTCAGACAAACCATGAAGTGAAACAAGTTGAATGTGCAATTGAGGAACGAAGGACAGCAGATCGGCAGGACGAGGTCTC
TGCAGCTACTTCAGGTCATCATCCAAAGCATGATAGCTACTCCTTGCCTGATTTTTCTTCAAACCATGGAATGGCTTCTGATGTACTCATCCCAGTTCCT
TCGGTCAGTCCTGATGAGGAACAAACTCATTCTTCCGAGTCATCATCTACTGCTTTCGGTTCTGAATCAGATATACTTGCTTCTACAACTGCGGATGCCC
CTGATGCCTCAAAAGTGGAATCTCCTGTTCTTAACCGGTCATCTGTGATTGTAGTTAACGGAGTAAAGGAACCTTTGGCTGGTGCAGTTCTAAAGGATTC
TGGGAAGCCTCAAGAATCTTCTGCCGCTGAGTTATCCCAAGTAGATGGACCATCTGCGATTGCAGTTAAAAAAGTCGAGGAACCATTTGCTGATGATATT
GTCCTAGCTGATCCCGGAAAGCCTCAAGAATCTTCTATCACAGAGTTCTCCCATTTGGAACCAATCAAGCTCTGGACTAATGGGGGCCTCTTAGGACTTC
AACCATCAAGGCCTCCTGATTTCAACGTGTCAAATGCTGGTAGTCAAGGTTCAGAAACAAATACAGTTAAGGTGGTAAACATCCCAATGCCGAGCAACAA
CGACGATAATGTTTCCAGCGCTAAAGATTCAATCGATCCTAACTACAGAAGTAGATTCCATCCGTATGACAGAGGATTGAATGCCAATAATCCAGTGACT
CCTGGAAATGGATTGCCGCTTGGTTCATTCATCAAGTTTACTACATCCGGAGGAGCCAGTCATGAAAGCGACGGATACCCATCTCAAAATATGTTTGGGC
TTGGGCAAAGATTACTTGCTAACGGCTTTGGCAGAAGGATATCACTTCCACCCAGTAGTGAATCTGAGCTGGCTAGTTCATCACCCCAACCTGCCGAGTT
GGAGAAAAAGAGTAGGCAGGCCTTGGTTAAATTTCAACCGAGTCAACCAGACGATCTCAACATACAGTTTGGACACGAGCCTTCAGATGCAGACTACCTT
ACATCTTCACCACCACTTGAGCACATGAAATTATCTTTCCAACCTACTGCCGGTGTCGAGGCTTCAAAACTGCAGCTGAAATTTCCTGATGGTAACTACT
GCAACGAAAGTACTACAGATATGTTTCCATCGTTTCAGTTGGTTCCGGAGGCTACAACTGGACGAAACCATGCAATCTCCGACTCGGATGATGACACATT
CTGCAGATCATCTCCTTACTTGTCAGATGATTGTCTTAGCCATCACTCCGATACGGATTCAGATCAGTGGGAAGCTGGTCAGTCCCCCGAAAGCGTCCAT
AACAGCGACCACGACGCTTCATGTGCAATCGAATCCGTGAAGCTCGAGGAGATGGAGAGCAATGGCCTTTGTACAAGTCCTGCAGACGAAACTGAACATG
CTGATGGTAGTACAGTTTCATCATGCCTCTCTGCTTCGGTACTCGATCTTCCTAGTTTCGACTCTGTCAAGCCGCCTCTTGAGGAAGTAAGACACAAAGA
AGATCACGGGTCAAGTCCATTGCCTCCACCTCTCCCCCCAGCTCAATGGCGGATATCAAAAGAATCGAAACCCGTGTCCGAAACGATAGAAAGGAAACCA
CAAGCTATACCTGATGCTCCCCAGACTGCGTTCGTTCCGAAACTCGTCGCAACTCCTACAACTCAGCAACCTAAACCAGCTCCAGCCGCTGAAAACCAGG
CTCATCCGGTCATCGTTCCATTCACCCCAAAGAGAAAGGATGAACCGAAGTTGAATCGGTGGAAAGAACCTAATCCACCCACAAATGGGAAGGTGATGCA
CGAACAAGAAGATTTCTTGCAGCAAATTAGATCAAAGTCATTCTCCCTGAGACCCACTGCACCAGCCAAGACGAAGGTGGCCGGTGCAACTCCAATCGAC
ACAGTGACCGAAATCTTGAAGAAGGCCAGTGCAATTCGCCAGGTAGTTGCGAGCGATGTCGATGACGACGATGCATGGAGTGACCCTTGA
|
||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10013904 pacid=23170071 polypeptide=Lus10013904 locus=Lus10013904.g ID=Lus10013904.BGIv1.0 annot-version=v1.0
MPLVRFEVRNQYALGHPELYKEADVDDPKAVLDGVAVAGLVGILRQLGDLAEFAAEVFHGLQEQLTTTASRSQKLKLRVQNIEAALPPIEKAILAQNSHI
HFAYTAGSEWHPRIRSEQNHFIYNDLPRFIMDSYEECRDPPRFHILDKFDTGGPGSCLKRYSDPTFFRRASGGFKEHDAEKVQRDKKARKIKKKRSLRRK
GSSLQNGSISSHSERSQFFQPTVNGRSSSSHTASNIDLPLKSEIGDRSNSIDSGNGSNHIECVFRPSSSAQPEEQKSNGFSSRSFQGDDSRDSGFSEGQS
RVAHDSFLHCSSPKQIHHRSPGVTWVEKEECEEPGVADDSFLHCSSPKQIHHRSPGVTWVGKEEHEEPGVADDSFLHVSSPELIPISSSCRTWEEKAEGA
QPEAHKGSLPRSSPERLPPGFSCVSWDEKEELSEAESQHYDGDEAPDVLTVDSVVDMQESGTNTEIHDPTDLRFENEHVPGPSFTAATLLEDVESETDNF
MDALNSIDTESESELDYQTKHEVKEVECANRDNEGLAVTSGQTNHEVKQVECAIEERRTADRQDEVSAATSGHHPKHDSYSLPDFSSNHGMASDVLIPVP
SVSPDEEQTHSSESSSTAFGSESDILASTTADAPDASKVESPVLNRSSVIVVNGVKEPLAGAVLKDSGKPQESSAAELSQVDGPSAIAVKKVEEPFADDI
VLADPGKPQESSITEFSHLEPIKLWTNGGLLGLQPSRPPDFNVSNAGSQGSETNTVKVVNIPMPSNNDDNVSSAKDSIDPNYRSRFHPYDRGLNANNPVT
PGNGLPLGSFIKFTTSGGASHESDGYPSQNMFGLGQRLLANGFGRRISLPPSSESELASSSPQPAELEKKSRQALVKFQPSQPDDLNIQFGHEPSDADYL
TSSPPLEHMKLSFQPTAGVEASKLQLKFPDGNYCNESTTDMFPSFQLVPEATTGRNHAISDSDDDTFCRSSPYLSDDCLSHHSDTDSDQWEAGQSPESVH
NSDHDASCAIESVKLEEMESNGLCTSPADETEHADGSTVSSCLSASVLDLPSFDSVKPPLEEVRHKEDHGSSPLPPPLPPAQWRISKESKPVSETIERKP
QAIPDAPQTAFVPKLVATPTTQQPKPAPAAENQAHPVIVPFTPKRKDEPKLNRWKEPNPPTNGKVMHEQEDFLQQIRSKSFSLRPTAPAKTKVAGATPID
TVTEILKKASAIRQVVASDVDDDDAWSDP
|
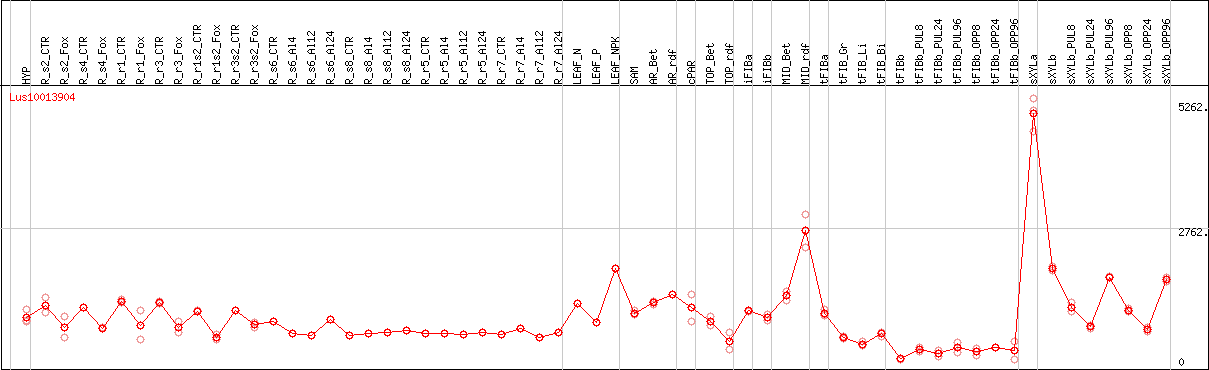
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10013904 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
