External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G56570 257 / 1e-82
PGN
PENTATRICOPEPTIDE REPEAT PROTEIN FOR GERMINATION ON NaCl, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G56310 194 / 5e-59
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT4G16835 194 / 6e-58
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G49170 194 / 4e-57
EMB2261
embryo defective 2261, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G22070 188 / 4e-55
pentatricopeptide (PPR) repeat-containing protein (.1)
AT3G57430 188 / 7e-55
OTP84
ORGANELLE TRANSCRIPT PROCESSING 84, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G20230 185 / 3e-54
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G49142 184 / 3e-54
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G39350 183 / 5e-54
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G13650 185 / 7e-54
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10019891
461 / 5e-162
AT1G56570 666 / 0.0
PENTATRICOPEPTIDE REPEAT PROTEIN FOR GERMINATION ON NaCl, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10010998
192 / 4e-57
AT4G16835 778 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10014211
190 / 1e-56
AT3G11460 791 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10022702
185 / 2e-55
AT3G11460 641 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10000213
187 / 3e-55
AT2G13600 435 / 8e-144
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10013540
186 / 2e-54
AT4G02750 1075 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10016425
186 / 3e-54
AT2G22070 1002 / 0.0
pentatricopeptide (PPR) repeat-containing protein (.1)
Lus10000597
179 / 3e-54
AT4G16835 545 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10001220
185 / 6e-54
AT3G57430 1110 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 84, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G006700
304 / 1e-100
AT1G56570 732 / 0.0
PENTATRICOPEPTIDE REPEAT PROTEIN FOR GERMINATION ON NaCl, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G010900
256 / 2e-82
AT1G56570 666 / 0.0
PENTATRICOPEPTIDE REPEAT PROTEIN FOR GERMINATION ON NaCl, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.016G019900
192 / 6e-57
AT4G02750 542 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.004G047800
192 / 2e-56
AT4G13650 654 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.007G021700
189 / 4e-56
AT3G47840 796 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.003G223900
186 / 5e-56
AT5G56310 508 / 1e-176
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.016G082200
186 / 6e-56
AT2G37320 602 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.016G051300
186 / 2e-55
AT3G13770 908 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.004G184800
189 / 3e-55
AT2G27610 1061 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.015G047600
189 / 3e-55
AT1G18485 1077 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10014026 pacid=23156858 polypeptide=Lus10014026 locus=Lus10014026.g ID=Lus10014026.BGIv1.0 annot-version=v1.0
ATGATGATTGGATATGGGGCTCATGGATATGGTATAGAGGCTGTAGCGCTGTTTGAGGAGATGGTGAGGTCAGATGTCAGGCCTGATCAGATTGTGTTTA
TGGCGGTTTTGAGTTCTTGTAGTCATGCTGGACTAGTCGACCAGGGCCTGAGGTATTTCAATACTATGATTGGTCACTACAAAATTAGGCCAAATCAGGA
AATTTATGGGTGTGTTGTGGATTTGTTGGGCAGAGCTGGGAGAGTTGAGGAGGCCTATCAGCTGATACACAGCATGCCATTTAAAGCTGACGAATCTGTT
TGGGGTGCGCTCCTTAGCGCCTGTAAAACCCATAACCTTCTAGATTTGGGTAAATTAGCTGCCTTGAGGGTGTTGGATTTGCAGCCAGATATGGTCAGTG
CTTATGTCATGCTGTCAAATATTTATGCAGCCGAAGGTAAATGGGACGAGTTTGCGAGGACAAGGAGATTGATGAAGGAAATTGTGAGCAAGAAGGTTTC
GGGGAAGAGTTGGATTGAAGTAAGAAACGAAGTATACAGTTTTGTCGTGGATGATACAGTTGGCATTCATATGGAGTTGGCACATCGAACTCTAGGTCTG
TTATGGCAGCACATGAAAGAAAAAGATAATATGGATGAAGACGACTGCTTAATAGATGCCATTGCAAATGGGGCTTGA
AA sequence
>Lus10014026 pacid=23156858 polypeptide=Lus10014026 locus=Lus10014026.g ID=Lus10014026.BGIv1.0 annot-version=v1.0
MMIGYGAHGYGIEAVALFEEMVRSDVRPDQIVFMAVLSSCSHAGLVDQGLRYFNTMIGHYKIRPNQEIYGCVVDLLGRAGRVEEAYQLIHSMPFKADESV
WGALLSACKTHNLLDLGKLAALRVLDLQPDMVSAYVMLSNIYAAEGKWDEFARTRRLMKEIVSKKVSGKSWIEVRNEVYSFVVDDTVGIHMELAHRTLGL
LWQHMKEKDNMDEDDCLIDAIANGA
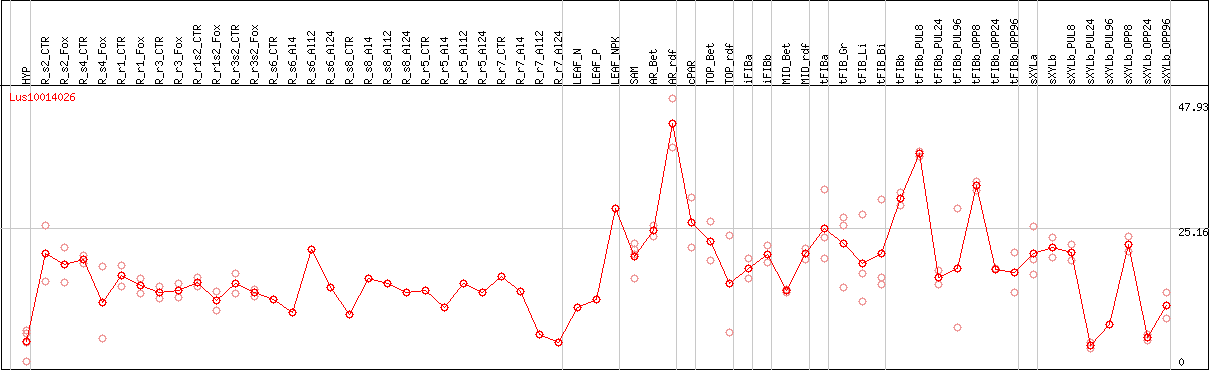
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10014026 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.