External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G34050 465 / 1e-168
CCoAOMT1
caffeoyl coenzyme A O-methyltransferase 1, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
AT4G26220 288 / 4e-99
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT1G67980 258 / 4e-87
CCOAMT
caffeoyl-CoA 3-O-methyltransferase (.1.2)
AT1G67990 235 / 3e-78
ATTSM1
TAPETUM-SPECIFIC METHYLTRANSFERASE 1, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT1G24735 235 / 5e-78
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
AT3G62000 166 / 3e-50
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
AT3G61990 153 / 3e-45
OMTF3
O-MTase family 3 protein, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10019841
508 / 0
AT4G34050 466 / 8e-169
caffeoyl coenzyme A O-methyltransferase 1, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Lus10027888
485 / 1e-176
AT4G34050 461 / 9e-167
caffeoyl coenzyme A O-methyltransferase 1, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Lus10002837
485 / 2e-176
AT4G34050 462 / 2e-167
caffeoyl coenzyme A O-methyltransferase 1, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Lus10034584
244 / 2e-81
AT1G67980 299 / 2e-103
caffeoyl-CoA 3-O-methyltransferase (.1.2)
Lus10012913
238 / 7e-79
AT4G26220 277 / 1e-94
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10009484
233 / 6e-75
AT4G26220 255 / 1e-83
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10021804
161 / 1e-49
AT1G24735 233 / 3e-78
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Lus10036339
163 / 3e-49
AT3G62000 385 / 9e-136
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Lus10010276
109 / 1e-28
AT3G62000 269 / 1e-90
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.009G099800
457 / 8e-166
AT4G34050 465 / 2e-168
caffeoyl coenzyme A O-methyltransferase 1, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Potri.001G304800
453 / 5e-164
AT4G34050 455 / 1e-164
caffeoyl coenzyme A O-methyltransferase 1, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Potri.018G070300
282 / 2e-96
AT4G26220 325 / 1e-113
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.008G136600
250 / 4e-84
AT1G67980 337 / 1e-118
caffeoyl-CoA 3-O-methyltransferase (.1.2)
Potri.010G104400
243 / 3e-81
AT1G67980 335 / 6e-118
caffeoyl-CoA 3-O-methyltransferase (.1.2)
Potri.002G183600
157 / 9e-47
AT3G62000 404 / 3e-143
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Potri.008G104400
42 / 0.0002
AT3G15355 267 / 7e-84
PHO2 FAMILY UBIQUITIN CONJUGATION ENZYME 1, ubiquitin-conjugating enzyme 25 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF01596
Methyltransf_3
O-methyltransferase
Representative CDS sequence
>Lus10014074 pacid=23156775 polypeptide=Lus10014074 locus=Lus10014074.g ID=Lus10014074.BGIv1.0 annot-version=v1.0
ATGGCTGAGCAACAGCAATCAGGGGAGAATGTCAGCAGGCATCAGGAAGTTGGTCACAAGAGTCTCCTCCAGAGCGATGATCTTTACCAGTATATTCTCG
AGACGAGCGTTTACCCGAGGGAGCCGGAGTCGATGAAGGAACTCCGGGAGGTCACCGCCAAACACCCCTGGAATATCATGACTACGTCTGCCGACGAAGG
ACAGTTCCTCAACATGCTACTGAAGCTCATCAATGCTAAGAACACAATGGAGATCGGCGTCTACACCGGCTACTCCCTTCTCGCCACCGCCCTCGCTCTC
CCCGACGACGGAAAGATCTTGGCAATGGATATCAACAGAGAGAACTACGAAATCGGCCTGCCAATCATCGAGAAAGCCGGACTTGCTCACAAGATCGAGT
TCAAAGAAGGCCCTGCACTGCCCGCTCTTGACAAGATGGTCGAAGACAAAGCGAACCACGGAGCGTACGACTTCATATTTGTGGACGCGGACAAGGACAA
CTACATCAACTACCACAAGAGGCTGATCGACTTGGTGAAGGTCGGTGGAGTCATCGGGTACGACAACACCCTATGGAACGGGTCCGTGGTTGCGCCTCCT
GATGCGCCGTTGAGGAAGTACGTCAGGTATTACAGGGACTTCGTTTTGGAGCTGAACAAGGCGCTCGCTGCTGACCCGAGAATCGAGATCTGCATGCTCC
CCGTTGGAGACGGAATCACTCTCTGCCGCCGGATCAGCTAA
AA sequence
>Lus10014074 pacid=23156775 polypeptide=Lus10014074 locus=Lus10014074.g ID=Lus10014074.BGIv1.0 annot-version=v1.0
MAEQQQSGENVSRHQEVGHKSLLQSDDLYQYILETSVYPREPESMKELREVTAKHPWNIMTTSADEGQFLNMLLKLINAKNTMEIGVYTGYSLLATALAL
PDDGKILAMDINRENYEIGLPIIEKAGLAHKIEFKEGPALPALDKMVEDKANHGAYDFIFVDADKDNYINYHKRLIDLVKVGGVIGYDNTLWNGSVVAPP
DAPLRKYVRYYRDFVLELNKALAADPRIEICMLPVGDGITLCRRIS
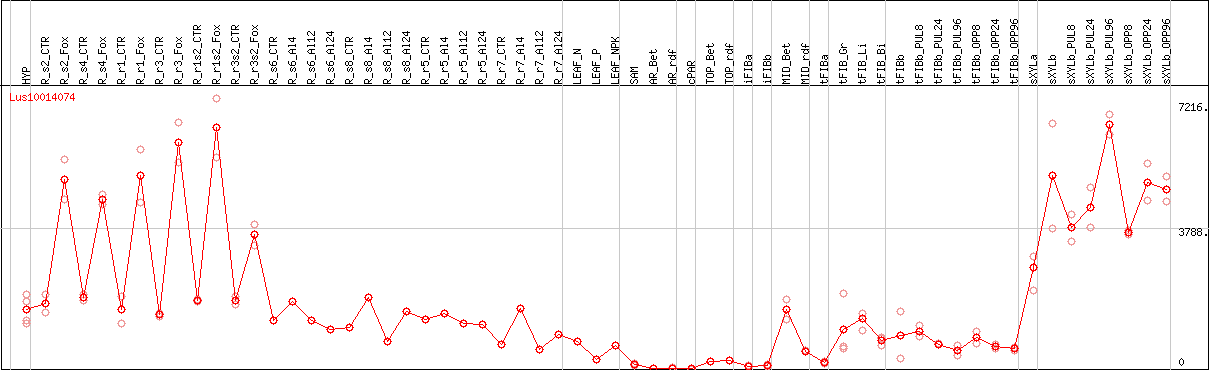
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10014074 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.