External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G38310 249 / 3e-84
RCAR10, PYL4
regulatory components of ABA receptor 10, PYR1-like 4 (.1)
AT2G40330 231 / 4e-77
RCAR9, PYL6
regulatory components of ABA receptor 9, PYR1-like 6 (.1)
AT5G05440 218 / 4e-72
RCAR8, PYL5
regulatory component of ABA receptor 8, PYRABACTIN RESISTANCE 1-LIKE 5, Polyketide cyclase/dehydrase and lipid transport superfamily protein (.1)
AT2G26040 187 / 2e-60
RCAR14, PYL2
regulatory components of ABA receptor 14, PYR1-like 2 (.1)
AT5G53160 185 / 2e-59
RCAR3, PYL8
PYR1-like 8, regulatory components of ABA receptor 3 (.1.2)
AT1G01360 184 / 3e-59
PYL9, RCAR1
PYRABACTIN RESISTANCE 1-LIKE 9, regulatory component of ABA receptor 1 (.1)
AT4G17870 184 / 6e-59
RCAR11, PYR1
regulatory component of ABA receptor 11, PYRABACTIN RESISTANCE 1, Polyketide cyclase/dehydrase and lipid transport superfamily protein (.1)
AT4G27920 179 / 2e-57
RCAR4, PYL10
regulatory components of ABA receptor 4, PYR1-like 10 (.1)
AT4G01026 174 / 4e-55
RCAR2, PYL7
regulatory components of ABA receptor 2, PYR1-like 7 (.1)
AT1G73000 161 / 6e-50
RCAR13, PYL3
regulatory components of ABA receptor 13, PYR1-like 3 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10022675
373 / 4e-132
AT2G38310 249 / 1e-83
regulatory components of ABA receptor 10, PYR1-like 4 (.1)
Lus10012231
265 / 2e-90
AT2G38310 234 / 1e-78
regulatory components of ABA receptor 10, PYR1-like 4 (.1)
Lus10007530
212 / 8e-71
AT2G38310 198 / 1e-65
regulatory components of ABA receptor 10, PYR1-like 4 (.1)
Lus10002861
207 / 4e-68
AT2G38310 176 / 7e-56
regulatory components of ABA receptor 10, PYR1-like 4 (.1)
Lus10007275
196 / 7e-64
AT2G40330 183 / 1e-58
regulatory components of ABA receptor 9, PYR1-like 6 (.1)
Lus10029222
194 / 5e-63
AT2G40330 182 / 2e-58
regulatory components of ABA receptor 9, PYR1-like 6 (.1)
Lus10001059
187 / 3e-60
AT5G53160 306 / 1e-107
PYR1-like 8, regulatory components of ABA receptor 3 (.1.2)
Lus10014929
178 / 4e-56
AT5G53160 289 / 2e-100
PYR1-like 8, regulatory components of ABA receptor 3 (.1.2)
Lus10040916
176 / 4e-56
AT1G01360 265 / 1e-91
PYRABACTIN RESISTANCE 1-LIKE 9, regulatory component of ABA receptor 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.016G125400
311 / 1e-108
AT2G38310 256 / 3e-87
regulatory components of ABA receptor 10, PYR1-like 4 (.1)
Potri.006G104100
306 / 5e-107
AT2G38310 249 / 2e-84
regulatory components of ABA receptor 10, PYR1-like 4 (.1)
Potri.010G183900
233 / 4e-78
AT2G40330 243 / 8e-82
regulatory components of ABA receptor 9, PYR1-like 6 (.1)
Potri.008G073400
229 / 1e-76
AT2G38310 229 / 2e-76
regulatory components of ABA receptor 10, PYR1-like 4 (.1)
Potri.001G142500
201 / 1e-65
AT4G17870 286 / 1e-99
regulatory component of ABA receptor 11, PYRABACTIN RESISTANCE 1, Polyketide cyclase/dehydrase and lipid transport superfamily protein (.1)
Potri.003G091700
194 / 5e-63
AT4G17870 273 / 3e-94
regulatory component of ABA receptor 11, PYRABACTIN RESISTANCE 1, Polyketide cyclase/dehydrase and lipid transport superfamily protein (.1)
Potri.018G054400
189 / 6e-61
AT2G26040 273 / 1e-94
regulatory components of ABA receptor 14, PYR1-like 2 (.1)
Potri.006G230600
188 / 8e-61
AT2G26040 266 / 6e-92
regulatory components of ABA receptor 14, PYR1-like 2 (.1)
Potri.012G000800
186 / 5e-60
AT5G53160 303 / 1e-106
PYR1-like 8, regulatory components of ABA receptor 3 (.1.2)
Potri.003G139200
184 / 2e-59
AT5G53160 297 / 3e-104
PYR1-like 8, regulatory components of ABA receptor 3 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0209
Bet_v_1_like
PF10604
Polyketide_cyc2
Polyketide cyclase / dehydrase and lipid transport
Representative CDS sequence
>Lus10014239 pacid=23152927 polypeptide=Lus10014239 locus=Lus10014239.g ID=Lus10014239.BGIv1.0 annot-version=v1.0
ATGCCTTCAAATCCTCCCAAATCCTCACTCCTCCTCCACCGCATCACCGACGCCGCCGCCGCCGCCACGGCGTGCCAGAAGCAGCGTCGGTTTCCGCTTC
CCTCCGCCGTCCCGGATGCGGCAGTAGCCCGGTACCACGACCACGCGTTGGGGCCCAACCAGTGCAGCTCCGCCGTTGTGCAGCAGATCGCCGCCCCTAT
CTCCACCGTCTGGTCCGTGTTGAGGAGGTTCGATAACCCGCAAGCGTACAAGCACTTCGTCAAGAGCTGCCACGTCATCCTCGGCACCGGCGACGTCGGG
TCCCTCCGTGAGGTCCACGTCATCTCGGGCCTTCCGGCTGCCAACAGCACCGAGAGGCTTGAGATCCTCGACGAGGAGAGACACGTCATCAGCTTCAGCG
TCGTGGGTGGGGACCACAGGCTGGCCAATTACCGTTCCGTCACCACGCTACACGGCGGCGGCGGAGAGACGGTGGCGGTAGAGTCTTACGTGGTCGACGT
TCCACCGGGGAACACGAAGGAGGACACATGCGTGTTCGTCGACACGATCGTCCAGTGCAACCTTCAGTCGCTGGCGCAGATCGCGGAAAATTCAGCTCGC
CGCAGTAAAGCAGTGTCGCCGTCGACGGCTCCGACCAAATCGTGA
AA sequence
>Lus10014239 pacid=23152927 polypeptide=Lus10014239 locus=Lus10014239.g ID=Lus10014239.BGIv1.0 annot-version=v1.0
MPSNPPKSSLLLHRITDAAAAATACQKQRRFPLPSAVPDAAVARYHDHALGPNQCSSAVVQQIAAPISTVWSVLRRFDNPQAYKHFVKSCHVILGTGDVG
SLREVHVISGLPAANSTERLEILDEERHVISFSVVGGDHRLANYRSVTTLHGGGGETVAVESYVVDVPPGNTKEDTCVFVDTIVQCNLQSLAQIAENSAR
RSKAVSPSTAPTKS
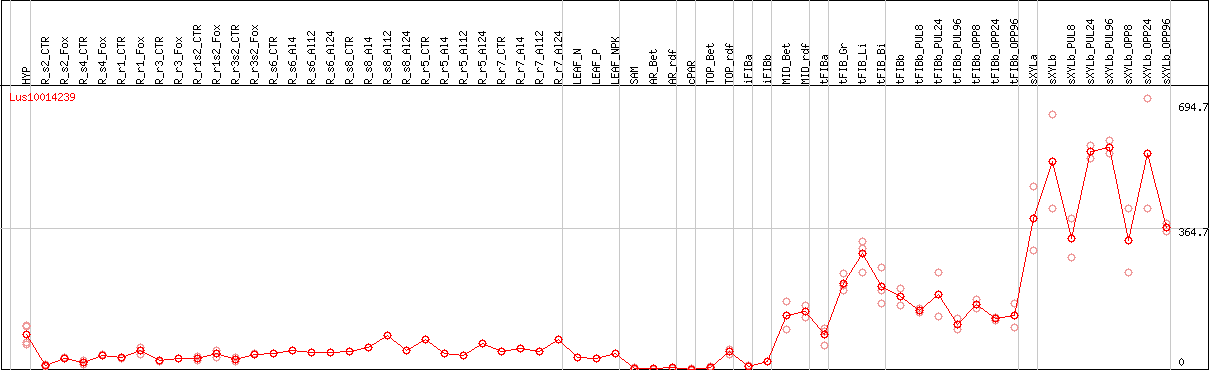
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10014239 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.