External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G19440 420 / 2e-148
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G51410 408 / 2e-143
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G09510 370 / 8e-129
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G09480 370 / 8e-128
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G09490 367 / 2e-127
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT1G66800 352 / 2e-121
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G09500 345 / 1e-118
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3)
AT4G35420 275 / 3e-91
TKPR1, DRL1
tetraketide alpha-pyrone reductase 1, dihydroflavonol 4-reductase-like1 (.1)
AT1G15950 256 / 2e-83
IRX4, ATCCR1, CCR1
IRREGULAR XYLEM 4, cinnamoyl coa reductase 1 (.1.2)
AT1G80820 248 / 9e-81
CCR2, ATCCR2
cinnamoyl coa reductase (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026070
654 / 0
AT5G19440 419 / 4e-148
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10002300
581 / 0
AT5G19440 423 / 2e-149
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10002302
552 / 0
AT5G19440 413 / 2e-145
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10026072
479 / 2e-167
AT5G19440 339 / 1e-112
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10039595
423 / 2e-149
AT5G19440 462 / 6e-165
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10009955
417 / 4e-147
AT1G51410 548 / 0.0
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10008668
394 / 8e-138
AT5G19440 388 / 9e-136
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10024130
369 / 6e-121
AT5G09860 933 / 0.0
nuclear matrix protein-related (.1.2)
Lus10019732
329 / 1e-112
AT5G19440 349 / 3e-120
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.009G057500
440 / 2e-156
AT5G19440 444 / 1e-157
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.009G057600
439 / 5e-156
AT5G19440 474 / 7e-170
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.009G052000
428 / 2e-151
AT1G51410 563 / 0.0
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.001G256400
424 / 8e-150
AT5G19440 552 / 0.0
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.009G057800
404 / 1e-141
AT5G19440 435 / 5e-154
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.008G120200
269 / 6e-89
AT1G68540 505 / 0.0
tetraketide alpha-pyrone reductase 2, cinnamoyl coA reductase-like 6, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.008G138600
268 / 2e-88
AT4G35420 535 / 0.0
tetraketide alpha-pyrone reductase 1, dihydroflavonol 4-reductase-like1 (.1)
Potri.001G045000
261 / 7e-86
AT1G15950 484 / 3e-173
IRREGULAR XYLEM 4, cinnamoyl coa reductase 1 (.1.2)
Potri.001G046400
260 / 3e-85
AT1G15950 510 / 0.0
IRREGULAR XYLEM 4, cinnamoyl coa reductase 1 (.1.2)
Potri.001G046100
260 / 3e-85
AT1G15950 511 / 0.0
IRREGULAR XYLEM 4, cinnamoyl coa reductase 1 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF01370
Epimerase
NAD dependent epimerase/dehydratase family
Representative CDS sequence
>Lus10014363 pacid=23164689 polypeptide=Lus10014363 locus=Lus10014363.g ID=Lus10014363.BGIv1.0 annot-version=v1.0
ATGAGCGGCGAAGGAAGAGTAGTATGTGTGACGGGCGGGTCGGGTTACATAGCCTCATGGATCATCAAGCTTCTGTTGGAACGCGGCTATACTGTCAAAA
CCACTGTTCGTGACCCAACTGATAAAACTAAGACCCAGCACTTGCTTTCCCTTGATGGGGCGGGAGATAGGCTAAAGCTGTACAAAGCAGATCTTAATGA
AGATGGGTGTTTTGATTCTGCTGTGGAGGGATGCGAAGGAGTCTTCCATACTGCCTCCCCTGTTTTCTTTCAATCTACCAACCCACAGGCAGATCTAATT
GAACCTGCAGTAAAGGGCACACTCAGCGTTCTTAAAGCATGTGCAAAAGTTCCTTCTATCAAGAGAGTTGTTCTAACCGCGTCGATGGGTTCAGTGCTCT
GGAATGGGAAGCCTCGTTCTCCTGGTGTTGTGGTCGATGAGACTTGGTTCTCTGATCCACAATTTTGTGAGAAGAAACAGTTTTGGTACATGCTTTCGAA
AACTTTGGCCGAGGAGGCTTCCTGGAAATTTTCGAAAGAGAATGGCATTGATCTTGTTGCGATTAATCCAGGGTTCGTAATCGGTCCTTACCTGCAGCAA
TCCATTAACTTAACGGTGGAGGAGGTTCTGAAGCTTGTAAATGGAACTAGAGAATTTCCAGGTGATATCTATAGATTTGTTGATGTACGAGATGTGGCGA
ACTCGCATATCCTAGCATTCGAGGTCCAATCAGCTAGTGGCAGATATTGTATAGTCGGAAGAACAGCACACTATAGCGAGGCTTTGGAGATTTTGCATGA
GCACTACCCTGATCTACCAGTATTTGCAAAATGGGAGAATGACAAGACATCTCATCCTACATATGAGATTTCTCAAAAGAAGGCAAAGAGCTTGGGTCTC
GAGTTTACTCCTCTGGAAGTGACTCTTAGGGATACTATAGAATGTTTCAAGGAGAAGGGTCTACTCAGTTTCTGA
AA sequence
>Lus10014363 pacid=23164689 polypeptide=Lus10014363 locus=Lus10014363.g ID=Lus10014363.BGIv1.0 annot-version=v1.0
MSGEGRVVCVTGGSGYIASWIIKLLLERGYTVKTTVRDPTDKTKTQHLLSLDGAGDRLKLYKADLNEDGCFDSAVEGCEGVFHTASPVFFQSTNPQADLI
EPAVKGTLSVLKACAKVPSIKRVVLTASMGSVLWNGKPRSPGVVVDETWFSDPQFCEKKQFWYMLSKTLAEEASWKFSKENGIDLVAINPGFVIGPYLQQ
SINLTVEEVLKLVNGTREFPGDIYRFVDVRDVANSHILAFEVQSASGRYCIVGRTAHYSEALEILHEHYPDLPVFAKWENDKTSHPTYEISQKKAKSLGL
EFTPLEVTLRDTIECFKEKGLLSF
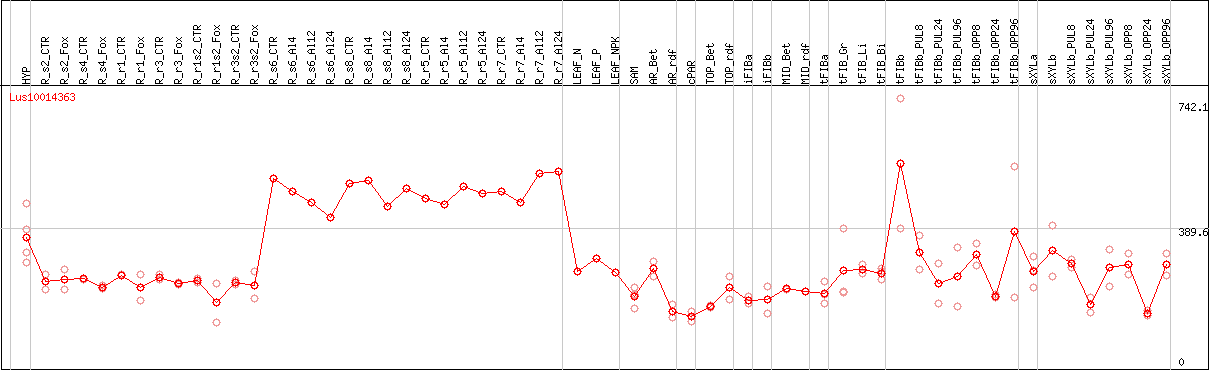
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10014363 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.