Lus10014759 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10014759 pacid=23150198 polypeptide=Lus10014759 locus=Lus10014759.g ID=Lus10014759.BGIv1.0 annot-version=v1.0
ATGAGCGTGATCGATTTGATAACCAGAGTCGACTCCATTTGCCGCAAGTTCGATAAGTACGATGTGGATAAGCAGACTGAGCTCAACGTCACCGGCGAAG
ATGCCTACGCGCGCCTCTACTCCACGATCGAGGCCGAGCTAGACTCCACTATCCAGAAAGCGGATTCCGTTGCTTCGGCGAAGAATAGGGCTGCCGTTGT
TGCTATTAATGCTGAGATTCGGAGGACTAAGGCTCGGTTGCTTGACGACATTCCAAAATTGCAGAGACTTGCTATGAAAAAGGTCAGAGGGCTTTCAATT
GAAGAGATGGAGGCTCGCAGTGATTTAGTTTCTGCACTCAAAGACAGGATTGCTTCCGTACCAGATGGATCCTCCACTGCAGCTCAACAAAGTAGCAATG
GTGGAGGAGCTTCAGCATCTTCTACAGGAATTAAGTTTAATTCTAACTTTGGTAATCAATTGATGAATTTCCTATGTGATACTGATCAAGGTTTAGATTT
TATAGCTGAAGGGCTGGACACTCTAAAGGACATGGCGCACGATATGAATGAGGAACTGGATAGACAAGTACCTTTGATAGATGAAATCGATGACAAGGTT
GATAAAGCAGCGTCTGACCTCAGAAGTACAAATGTCAGACTGAAAGATACTGTTAACCAGCTAAGGTCCAGTCGCAACTTTTGCATCGACATCGTCCTCT
TGTGCATCGTTCTGGGCATTGCTGCCTATTTGTACAAATTTCCTCGTGATCCAGCGAGCATCTCTCGTTTGTACCTCCGTAGCAGCGCGTCGCCGTCGGC
GGGGATTAGGAAGAAGATGACTTATTCCGATGAGAAGGCGGCGCAAATGCAGTTGCGCCAGAATTACAGGAATCTCTGGCACACCGATCTCTGCGGTGCC
GTCGGCGCCGATGCTCCCTTTTGCTGTTTTGCGACGGTCTGCGGGCCATGCGCTTCCTACATCCTGCGGAAACGAGCTCTTTACAATGACTTATCAAGGT
ACACTTGCTGCGCTGGATACATGCCTTGCAGTGGCAACTGTGGAGAGAGAAACTGCCCTGAGTTCTGTCTTTGCTCCGAGGTGTTGTGCTGCTTTTCGAA
TTCGGTAGCCTCGACCCGATTCCTGTTGCAAGATGAATTCAACATCCAGACATCAACCTGTGACAACTGCATCATTGGGTTCATGTTCTGCCTACAGCAG
TTGGCCTGCATATGTTCATGCGTAGCCTGCTTAACTGGGATTGATGAAATAGCTGACCTTGCTGACCTCTTGACTTGCCTCTCTGATCTCGTCTTCTGCT
CCGTTTGTCCATGTCTTCAGACTCAACACAAAGTTGAGATGGATCAAAGGGACATGAAGTATGGACCTCCACCAACGATGGGGGTTCCTCCAATGCAACA
GATGTCAAGAATTGACCAGCCTATCCCTCCTTCTGCAAACAACTATTACTCACCATATGAAGAGGATCTCAAGGCCAAGAGTGGTCCAATGAGTCAACCT
CCTCCACCCTATCCGGGCGCAGGATATCCGTATGGAGGATATCCTTCTCCGGGATCGGGATACGGTCCTCCTCCTCCAGGAACGGGATATGGTCCTCCTC
CGCCTGGATATCCTTATCCCGGATCTTACCAGGGATATCCTCCGCCGGGATACATTAATCAATCCCCGGGATACCCTCCTTCGTATCCGCCACCTCCTTC
TGCTTATCCTCCGCCTAATTACACTAACAACAACCACGACGACAACAAAAAGTGA
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10014759 pacid=23150198 polypeptide=Lus10014759 locus=Lus10014759.g ID=Lus10014759.BGIv1.0 annot-version=v1.0
MSVIDLITRVDSICRKFDKYDVDKQTELNVTGEDAYARLYSTIEAELDSTIQKADSVASAKNRAAVVAINAEIRRTKARLLDDIPKLQRLAMKKVRGLSI
EEMEARSDLVSALKDRIASVPDGSSTAAQQSSNGGGASASSTGIKFNSNFGNQLMNFLCDTDQGLDFIAEGLDTLKDMAHDMNEELDRQVPLIDEIDDKV
DKAASDLRSTNVRLKDTVNQLRSSRNFCIDIVLLCIVLGIAAYLYKFPRDPASISRLYLRSSASPSAGIRKKMTYSDEKAAQMQLRQNYRNLWHTDLCGA
VGADAPFCCFATVCGPCASYILRKRALYNDLSRYTCCAGYMPCSGNCGERNCPEFCLCSEVLCCFSNSVASTRFLLQDEFNIQTSTCDNCIIGFMFCLQQ
LACICSCVACLTGIDEIADLADLLTCLSDLVFCSVCPCLQTQHKVEMDQRDMKYGPPPTMGVPPMQQMSRIDQPIPPSANNYYSPYEEDLKAKSGPMSQP
PPPYPGAGYPYGGYPSPGSGYGPPPPGTGYGPPPPGYPYPGSYQGYPPPGYINQSPGYPPSYPPPPSAYPPPNYTNNNHDDNKK
|
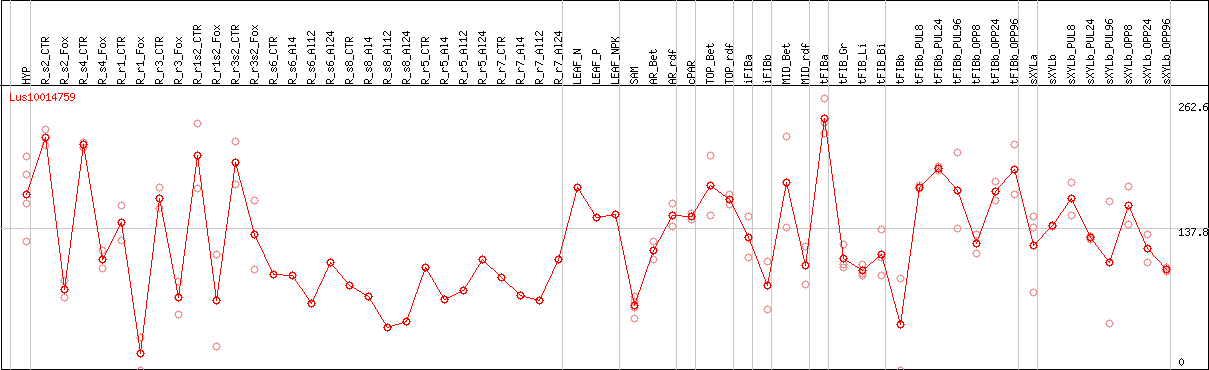
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10014759 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
