Lus10014768 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10014768 pacid=23150196 polypeptide=Lus10014768 locus=Lus10014768.g ID=Lus10014768.BGIv1.0 annot-version=v1.0
ATGGGAAAGGAAGAAAGTCCAACGGTGACAAGGAGGAAAACCAGGTCATCGTGTGCTGCTGTAGCTTCAAATGATGATGAGGTTCAAACTTTGAAAACAC
CTCCACTGCCCAATCCAACAGTAAATGACCTTGCTTTCGGAGGTGAACCCCTTAGCTTTGATGATCTGCTGTCTAGTTTCCCAGCCAGGCGCTCTCAGAT
TCTTCAGCTGGTGCACTTATTAGGTCCTTCAAACTCCCCTGTGCTTCCTCTTTTTGTGTATGGAGGTGTTTCCACAGGAAAAACTACCACCATTCTCCAA
GTTTTCAGGCATCTTAATCGTCCATTTGTCTATGCAAGCTGCCGTACTTGTTATAGCCCTCGGATCTTGTTCGAATCCATTCTAAACCAGTTGTTGCTTC
ATAGAAAGAGTGCTGATAATGGTTATGCAAGTGCAAAACGATGTGAAAAGCCGTCTGATTTTGTCAACTTTCTCCGTGAAGATGTTGGAAAGATTGTCGA
CAATCTGAAGAAGGATCATTCAGGGAAGCAGAACAACTCGAATAAGTTGCCCGCACGGCCTAATGGAGATGGCCTCTACTTGATACTTGATAACTTGGAG
CTAGTGAGGGAGTGGGATAAAAGTTCGAGCTTATTGCCTTTTCTGTTTAACCTCTATGATATTCTCAAGCTGCCTAATTTCGGACTCATATTCATTAGCA
GCAGTTCACCAGATATGTATTACTCAAGTATGGGGTACATAGAGCCTATTCCTGTGTTTTTCCCTGAATATACAGAAGAAGATCTTCGCCGAATACTGAT
GAGGAATCAATCAAACCAGAAGCTATATGGTTGGTTTCTTGACATTGTACTGAGGCCATTCTGTAGAGTGACCAGGCGACTCGATGAATTAAGTCCAGTC
TTTTCATCTTTGTTTCAAAAGTATTCCGAACCACTTAGCGACACAAGTACCTTACCCAGTGATGGCATGAAGAGAAGATTGTTTATTAGCTTTCAGCCTT
GCATTTCGCCTTATTTGAATGATATATTTTGGGGGCCATCTCTATCTTTCTTAGGAGTTGATTCCGTGAAGGATACTACCAGGGAGATAGATTTCCATAT
GTCCACTGCAGCTAAATATCTTCTTATCTCAGCATTCATCGCTTCAAGAAATCCGGCTACACTGGATGCGTCTTTGTTCGATAGCACTGGTGGCTCAGAT
AATCCCAAGCGTAAGAGGAAAGCCTCCGAGAAGTCAAAGGAGCAAAAGGAAACTGCGGAACAGGAATTGCTTATGAAAGGTCCCGGAACTTTTCCATTGG
AAAGACTGCTGGCTATCTTCCAATGCATAACGTCGGTAACAGATGTATCCCTTGATGAAGACGAACATGGAGATAGTTTTTCAACAGAAGAAGGCAGCAA
TGCGCTCATGTCAAGTGTCCTACTACAGCTGTCGACTCTTTGTGATGCTAATTTCGTAGTAAAAGGCGGAAGCTGCCCGTTGGAAGGTTCGACAAGGTAT
CGATGCACCATTTCTGAAGATTTGGCTCTAAAGTCTGAAGAAAAACTAAAACACTCTCAACCATTTTCATCTACACCTCGAAAAACCATCTGGGTTCCAA
AATTTTCAATTCAATTGGTGGGTTTCACTTTCTCTGGGGAATTTCATCGAACACCTTATGGACAACATCTCGACGGCGCTGGTTCTCCTCTCAGCAGCGA
CCGCTTACCTCCTGTGGTTCACCTTCATAATCTCTCGTTCGTTGCGGGGCCCACGTGTCTGGCCGTTGCTCGGAAGCCTCCCCGGACTGATCGAGAACTG
CGGCCGCCTCCACGACTGGATCGCCGCAAACCTCCAAGCCTGCGGCGGCCCCTCCAAGCCTGCGGCGGCACTTACCAGACCTGCATATGTGCGGTGCCAT
TTTTAGCTAAAAAACAAGGTCTCGTGACCGTTACGTGCGACCCGAAGAACATCGAGCACATTTTAAAAATCCGGTTCGAAAACTACCCCAAGGGCCCCAC
CTGGCAAGCCGTATTCCACGAGCTACTCGGCGAGGGGATCTTCAATTCTGACGGTGACACATGGCGTTTCCAACGTAAGACCGCCGCACTCGAGTTCACC
ACTAGGACCCTCCGTCAGGCCATGGCTCGGTGGGTTAGTCGAACCATCAAGTTGAGGTTCTGTCCCGTCCTCAAATTGGCTCAGGAGAATTTCGAGCCGG
TCGACTTGCAGGATTTGCTACTTCGGCTCACTTTCGACAACATTTGCGGTCTGGCATTCGGGAAGGACCCGCAAACTTGCACTGCGGGACTTCCCGAAAA
CGAATTTGCATCGGCATTTGACCAAGCCACTGAAGCATCATTGCAACGTTTCATTCTTCCGAAAGTTGTGTGGAGAGTGAAGAAATTGCTCGGGCTCGGG
ATGGAAGTGAGCTTGAGCCGAAGCATTACTCGGCTCGATCGTTATCTCGCTGACATCATCAACCAACGGAAGGATGAGTTGCTGAGTCGGACAAAAGACA
AAAACAACAGCAATCTCCAACACGACGACCTAGTCACTAGGTTTATGAAGAGAAAAGAGTCCTACTCAGACTCTTTCCTCCAACATGTGGCGCTAAACTT
CATCCTAGCTGGAAGAGACACGTCATCTGTGGCGTTGAGCTGGTTCTTCTGGCTGATCATCCAAAACCCGTCAGTTGAAGATAAAATCCTACGTGAGCTA
TGCAGCGTTTTGGAGGAGACACGTGGCGCTGACCTGGCTAAGTGGCTTGACGACCCGTTGGGGTTTGAAGAGGCTGACCGCCTCATCTACCTGAAAGCGG
CTTTATCGGAGACTCTCCGGCTTTACCCGTCGGTCCCACAGGACTCCAAGCACGTCGTCGCCGACGACGTTCTACCGGACGGGACGTTCGTGCCGGCGGG
ATCGTCGATCACCTACTCCATATACGCAGCCGGGAGGATGCGGACAGCGTGGGGAGCTGATTGCATGGAGTTCCGACCGGAGCGATGGCTGTCCGCCGGC
GGTGAAAAATTCATACCGCAGGATCCGTACAAGTTCGTCGCTTTCAACGCCGGCCCGAGGGTATGCTTGGGGAAGGAGTTGGCGTATCTTCAGATGAAGT
CGGTTGCCGCGGCGGTGCTGCTCCGGCACCGGCTGACGGTGGTGGAGGGGCATAAGGTGGAGCAGAAGATGTCGTTGACTCTGTTCATGAAATACGGGCT
GAAGGTCAACGTGTTGAAGAGAGACCTTAAAGAGACGGAAGCGGCGATGATGAAGAAGCAGGCGGCAACCGTGGTCGGCGACGTTGAGGATGAAAGACAA
AATTGTAATTGGGAGAAAGCGGAGGAGCATCTGCCGTTAGCAGCCGGCGTTAGATGTAACGGTGTGTGGGAGAGTGGGAGTAAGCTTGATCAGAAAAGGA
ATTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10014768 pacid=23150196 polypeptide=Lus10014768 locus=Lus10014768.g ID=Lus10014768.BGIv1.0 annot-version=v1.0
MGKEESPTVTRRKTRSSCAAVASNDDEVQTLKTPPLPNPTVNDLAFGGEPLSFDDLLSSFPARRSQILQLVHLLGPSNSPVLPLFVYGGVSTGKTTTILQ
VFRHLNRPFVYASCRTCYSPRILFESILNQLLLHRKSADNGYASAKRCEKPSDFVNFLREDVGKIVDNLKKDHSGKQNNSNKLPARPNGDGLYLILDNLE
LVREWDKSSSLLPFLFNLYDILKLPNFGLIFISSSSPDMYYSSMGYIEPIPVFFPEYTEEDLRRILMRNQSNQKLYGWFLDIVLRPFCRVTRRLDELSPV
FSSLFQKYSEPLSDTSTLPSDGMKRRLFISFQPCISPYLNDIFWGPSLSFLGVDSVKDTTREIDFHMSTAAKYLLISAFIASRNPATLDASLFDSTGGSD
NPKRKRKASEKSKEQKETAEQELLMKGPGTFPLERLLAIFQCITSVTDVSLDEDEHGDSFSTEEGSNALMSSVLLQLSTLCDANFVVKGGSCPLEGSTRY
RCTISEDLALKSEEKLKHSQPFSSTPRKTIWVPKFSIQLVGFTFSGEFHRTPYGQHLDGAGSPLSSDRLPPVVHLHNLSFVAGPTCLAVARKPPRTDREL
RPPPRLDRRKPPSLRRPLQACGGTYQTCICAVPFLAKKQGLVTVTCDPKNIEHILKIRFENYPKGPTWQAVFHELLGEGIFNSDGDTWRFQRKTAALEFT
TRTLRQAMARWVSRTIKLRFCPVLKLAQENFEPVDLQDLLLRLTFDNICGLAFGKDPQTCTAGLPENEFASAFDQATEASLQRFILPKVVWRVKKLLGLG
MEVSLSRSITRLDRYLADIINQRKDELLSRTKDKNNSNLQHDDLVTRFMKRKESYSDSFLQHVALNFILAGRDTSSVALSWFFWLIIQNPSVEDKILREL
CSVLEETRGADLAKWLDDPLGFEEADRLIYLKAALSETLRLYPSVPQDSKHVVADDVLPDGTFVPAGSSITYSIYAAGRMRTAWGADCMEFRPERWLSAG
GEKFIPQDPYKFVAFNAGPRVCLGKELAYLQMKSVAAAVLLRHRLTVVEGHKVEQKMSLTLFMKYGLKVNVLKRDLKETEAAMMKKQAATVVGDVEDERQ
NCNWEKAEEHLPLAAGVRCNGVWESGSKLDQKRN
|
DESeq2's median of ratios [FLAX]
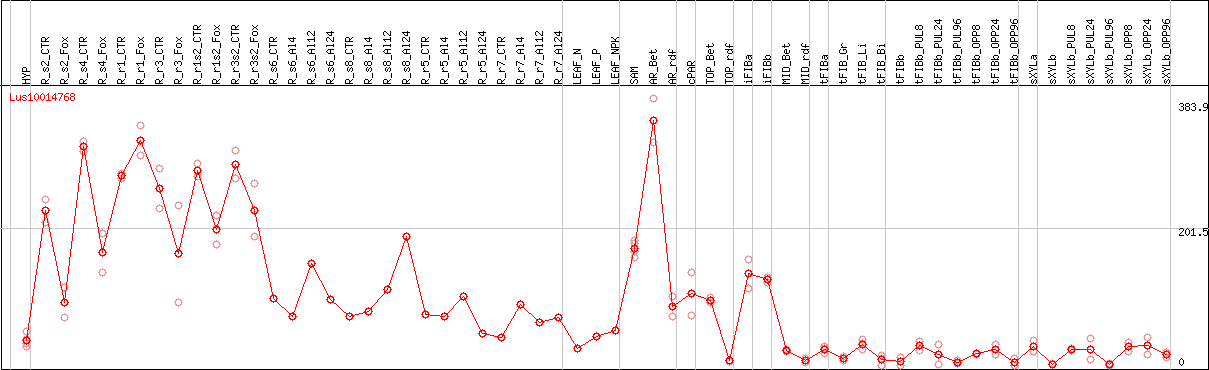
Coexpressed genes
Lus10014768 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
