External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G55960 372 / 5e-130
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT1G29780 92 / 4e-22
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT1G29770 87 / 7e-20
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT5G45700 79 / 5e-17
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT5G11860 72 / 5e-14
SSP5
SCP1-like small phosphatase 5 (.1.2.3.4)
AT5G46410 65 / 1e-11
SSP4
SCP1-like small phosphatase 4 (.1.2)
AT1G55900 65 / 1e-11
TIM50, EMB1860
embryo defective 1860, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
AT4G18140 64 / 2e-11
SSP4b
SCP1-like small phosphatase 4b (.1.2.3)
AT5G23470 48 / 3e-06
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT3G19595 46 / 1e-05
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10009890
502 / 0
AT3G55960 445 / 9e-159
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10028284
379 / 9e-133
AT3G55960 383 / 2e-134
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10002484
301 / 6e-102
AT3G55960 333 / 2e-114
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10004810
271 / 7e-90
AT3G55960 295 / 3e-99
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10007321
97 / 3e-23
AT5G45700 206 / 3e-65
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10029273
94 / 6e-22
AT5G45700 211 / 4e-67
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10040202
81 / 4e-19
AT3G55960 77 / 3e-18
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10004596
75 / 8e-15
AT5G46410 398 / 3e-135
SCP1-like small phosphatase 4 (.1.2)
Lus10025376
74 / 2e-14
AT5G46410 343 / 2e-114
SCP1-like small phosphatase 4 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G188500
407 / 4e-144
AT3G55960 406 / 8e-144
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.008G068800
390 / 2e-137
AT3G55960 342 / 3e-118
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.006G100800
321 / 9e-110
AT3G55960 304 / 6e-103
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.016G116700
303 / 5e-103
AT3G55960 248 / 1e-81
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.004G062900
88 / 5e-20
AT5G45700 194 / 1e-60
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.011G072000
87 / 1e-19
AT5G45700 230 / 5e-75
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.001G353700
79 / 3e-16
AT5G46410 409 / 2e-138
SCP1-like small phosphatase 4 (.1.2)
Potri.011G078300
72 / 4e-14
AT5G46410 419 / 1e-143
SCP1-like small phosphatase 4 (.1.2)
Potri.001G364500
67 / 2e-12
AT1G55900 405 / 3e-141
embryo defective 1860, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
Potri.011G093000
66 / 5e-12
AT1G55900 415 / 5e-145
embryo defective 1860, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0137
HAD
PF03031
NIF
NLI interacting factor-like phosphatase
Representative CDS sequence
>Lus10014844 pacid=23150212 polypeptide=Lus10014844 locus=Lus10014844.g ID=Lus10014844.BGIv1.0 annot-version=v1.0
ATGGCTGAGTTGACTCAGTCCGATGTGGTCTACTCTCCTCGGTCTCTCCAGCTATGGATGACGCTCTGGAACTGGCTGGCTTTCTTCTTCCAGATCTTCC
TTCAGATCCTCAGAACTCTTGGACACCTTCCTCTTCTCTCCTCCTCCTCGACGACTTCTCACTCTTTCAAGCCCTTGCCCGACGTCGAATTGCCGGAGAT
CGATTCTGCTCCTTCTCCTTCCACTTTGGAGATCGCCACCGGTGACCTCGACTCCGGTCGGATAATTCAGCCCCTCCAAAGACTCACGGTTGTTCTCGAC
TTGGACGAGACTCTGATTTGTGCATACGAGACCTCCAGTTTGCCTCCGGCCCTCAGGAGCCAGGCCACTGACGCCGGATTGAAGTGGTTCGAGTTGGAGT
GCGTGTCGTCAGACAAGGACAGTGACGGGAAACCTAAGATCAATTATGTTACGGTGTTTGAGCGTCCAGGGTTGCATGAATTCCTTAAAAAGCTCAGTGA
ATTTGCTGACCTTGTGCTCTTCACTGCTGGCTTGGAAGGTTATGCTAGACCCCTTGTTGACAGAATAGATACAGAAAAGCTGTTTAGCCTTCGCCTTTAT
CGGCCATCGACGACTAGCACGGAATACCGTGAGCATGTGAAGGATCTATCTCGCATATCAAACGATGGCAGCCACATTGTTATTGTGGACAACAACCCAT
TCAGTTTCTTGTTACAACCCTTGAATGGAATCCCATGCATTTCATTTTCTGCTGGCCAACCACAGGACACACAGCTACTAGATATAATCCTACCTCTCCT
GAAGCACCTATCTCAGCAGATGGACGTGAGGCCAGTGCTGTATGAGAGATTCCACATGCCTGAATGGTTTCAGAAACATGGAATCCCTGCTTCTGTCTGG
ACATGA
AA sequence
>Lus10014844 pacid=23150212 polypeptide=Lus10014844 locus=Lus10014844.g ID=Lus10014844.BGIv1.0 annot-version=v1.0
MAELTQSDVVYSPRSLQLWMTLWNWLAFFFQIFLQILRTLGHLPLLSSSSTTSHSFKPLPDVELPEIDSAPSPSTLEIATGDLDSGRIIQPLQRLTVVLD
LDETLICAYETSSLPPALRSQATDAGLKWFELECVSSDKDSDGKPKINYVTVFERPGLHEFLKKLSEFADLVLFTAGLEGYARPLVDRIDTEKLFSLRLY
RPSTTSTEYREHVKDLSRISNDGSHIVIVDNNPFSFLLQPLNGIPCISFSAGQPQDTQLLDIILPLLKHLSQQMDVRPVLYERFHMPEWFQKHGIPASVW
T
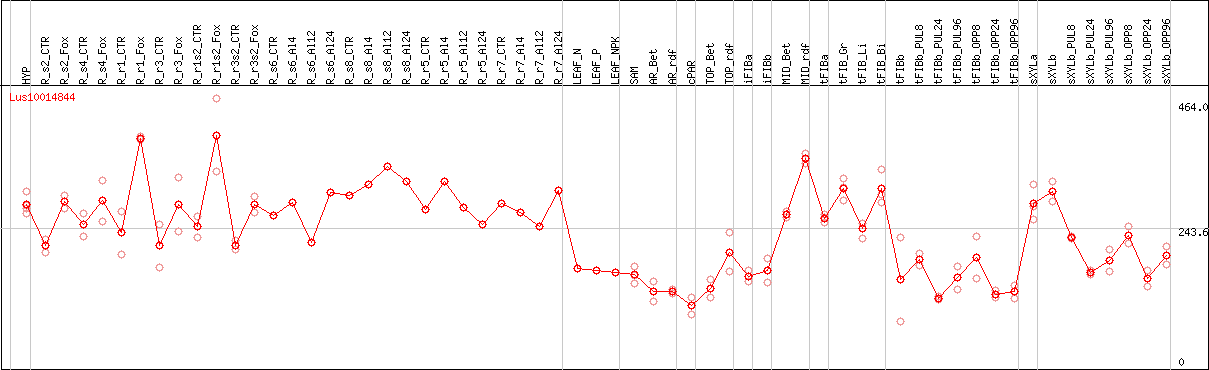
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10014844 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.