External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G26910 209 / 3e-70
RPL10B
ribosomal protein L10 B, Ribosomal protein L16p/L10e family protein (.1)
AT1G14320 209 / 5e-70
RPL10A, RPL10, SAC52
SUPPRESSOR OF ACAULIS 52, ribosomal protein L10 A, ribosomal protein L10, Ribosomal protein L16p/L10e family protein (.1.2)
AT1G66580 197 / 3e-65
RPL10C, SAG24
ribosomal protein L10 C, senescence associated gene 24 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031568
248 / 1e-86
AT1G26910 209 / 5e-70
ribosomal protein L10 B, Ribosomal protein L16p/L10e family protein (.1)
Lus10012113
242 / 2e-84
AT1G26910 209 / 3e-70
ribosomal protein L10 B, Ribosomal protein L16p/L10e family protein (.1)
Lus10010429
242 / 2e-84
AT1G26910 209 / 3e-70
ribosomal protein L10 B, Ribosomal protein L16p/L10e family protein (.1)
Lus10031002
239 / 4e-83
AT1G26910 206 / 5e-69
ribosomal protein L10 B, Ribosomal protein L16p/L10e family protein (.1)
Lus10035401
238 / 4e-83
AT1G26910 206 / 5e-69
ribosomal protein L10 B, Ribosomal protein L16p/L10e family protein (.1)
Lus10034092
225 / 7e-78
AT1G26910 216 / 7e-73
ribosomal protein L10 B, Ribosomal protein L16p/L10e family protein (.1)
Lus10003055
115 / 9e-35
AT1G26910 135 / 2e-41
ribosomal protein L10 B, Ribosomal protein L16p/L10e family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G131900
226 / 9e-77
AT1G26910 410 / 6e-148
ribosomal protein L10 B, Ribosomal protein L16p/L10e family protein (.1)
Potri.013G159600
224 / 6e-76
AT1G26910 411 / 3e-148
ribosomal protein L10 B, Ribosomal protein L16p/L10e family protein (.1)
Potri.013G159301
134 / 1e-40
AT1G14320 222 / 2e-73
SUPPRESSOR OF ACAULIS 52, ribosomal protein L10 A, ribosomal protein L10, Ribosomal protein L16p/L10e family protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00252
Ribosomal_L16
Ribosomal protein L16p/L10e
Representative CDS sequence
>Lus10015106 pacid=23149494 polypeptide=Lus10015106 locus=Lus10015106.g ID=Lus10015106.BGIv1.0 annot-version=v1.0
ATGCTTTCGTGTGCCGGAGCTGATAGGCTCCAGACCGGTATGAGGGGTGCCTTCGGTAAGCCTCAAGGCGTGTGTGCTAGGGTTGCAATTGGCCAGGTTC
TCTTATCAGTACGTTGCAAGGACAGCAACAGTCAGCATGCTCAGGAGGCTCTTCGTCGTGCCAAGTTCAAGTTCCCTGGTCGTCAGAAGATCATCATCAG
CAGGAAATGGGGATTCACCAAGTTCAATCGAGCTGACTATGTGAAGTTGAAGCAAGAGAACAGGATTATGCCTGATGGTGTCAATGCTAAGCTTCTTGGA
TGCCATGGACCGTTGGCTAACAGACAACCTGGAAGAGCATTCGCACCAACTGCCACCGCATAG
AA sequence
>Lus10015106 pacid=23149494 polypeptide=Lus10015106 locus=Lus10015106.g ID=Lus10015106.BGIv1.0 annot-version=v1.0
MLSCAGADRLQTGMRGAFGKPQGVCARVAIGQVLLSVRCKDSNSQHAQEALRRAKFKFPGRQKIIISRKWGFTKFNRADYVKLKQENRIMPDGVNAKLLG
CHGPLANRQPGRAFAPTATA
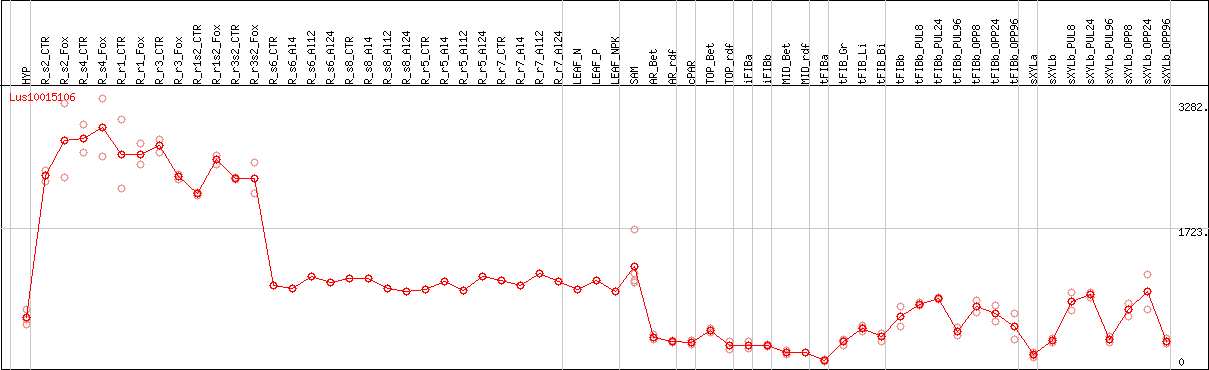
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10015106 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.