External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G05000 295 / 3e-102
AtPFA-DSP1
plant and fungi atypical dual-specificity phosphatase 1, Phosphotyrosine protein phosphatases superfamily protein (.1.2)
AT4G03960 267 / 2e-91
AtPFA-DSP4
plant and fungi atypical dual-specificity phosphatase 4, Phosphotyrosine protein phosphatases superfamily protein (.1)
AT2G32960 268 / 9e-91
AtPFA-DSP2
plant and fungi atypical dual-specificity phosphatase 2, Phosphotyrosine protein phosphatases superfamily protein (.1)
AT3G02800 207 / 8e-68
AtPFA-DSP3
plant and fungi atypical dual-specificity phosphatase 3, Tyrosine phosphatase family protein (.1)
AT5G16480 199 / 1e-64
AtPFA-DSP5
plant and fungi atypical dual-specificity phosphatase 5, Phosphotyrosine protein phosphatases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031529
375 / 5e-134
AT1G05000 304 / 2e-106
plant and fungi atypical dual-specificity phosphatase 1, Phosphotyrosine protein phosphatases superfamily protein (.1.2)
Lus10029991
316 / 7e-110
AT1G05000 296 / 1e-102
plant and fungi atypical dual-specificity phosphatase 1, Phosphotyrosine protein phosphatases superfamily protein (.1.2)
Lus10035333
313 / 6e-109
AT1G05000 291 / 2e-100
plant and fungi atypical dual-specificity phosphatase 1, Phosphotyrosine protein phosphatases superfamily protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G159100
321 / 3e-112
AT1G05000 310 / 3e-108
plant and fungi atypical dual-specificity phosphatase 1, Phosphotyrosine protein phosphatases superfamily protein (.1.2)
Potri.002G224000
308 / 2e-107
AT1G05000 300 / 9e-105
plant and fungi atypical dual-specificity phosphatase 1, Phosphotyrosine protein phosphatases superfamily protein (.1.2)
Potri.011G021100
281 / 6e-97
AT4G03960 288 / 4e-100
plant and fungi atypical dual-specificity phosphatase 4, Phosphotyrosine protein phosphatases superfamily protein (.1)
Potri.004G002800
279 / 4e-96
AT1G05000 286 / 4e-99
plant and fungi atypical dual-specificity phosphatase 1, Phosphotyrosine protein phosphatases superfamily protein (.1.2)
Potri.013G086500
205 / 6e-67
AT3G02800 296 / 2e-103
plant and fungi atypical dual-specificity phosphatase 3, Tyrosine phosphatase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0031
Phosphatase
PF03162
Y_phosphatase2
Tyrosine phosphatase family
Representative CDS sequence
>Lus10015154 pacid=23149515 polypeptide=Lus10015154 locus=Lus10015154.g ID=Lus10015154.BGIv1.0 annot-version=v1.0
ATGAAATTCGATCACAATTTTACCGCCACGGGAGGAGGAGGAGGAGATCAACCGCCACGGTTCGAAGACGGCGAAGGATCGTCGTCCTCGGAGGAAGAGA
TGTGCAGGACGATCGAGGTCTCCTCCGTGATGATCAACCACCGCCGCGGCGAGGATCACTTATCGGATTCGCCGCCGCCGTTATGTGACGAGCTGAGTTT
CATCCCGCCGTTGAACTTCTCCATGGTTGATAATGGCATCTTCCGCTCCGGATTCCCTGACTCCGCCAACTTCTCCTTCCTTGAAACTCTCGGCCTCCGC
TCCATCATGGTTTCCAGATGCTTGTGCCCGGAGCCGTATCCCGAGGCGAACGCAGAGTTCCTCAAGGCCAACGGAATCAGGCTTTTCCAGTTTGGAATTG
AAGGTTACAAGGAGCCATTTGTAAACATCCCAGAGGATCCAATCCGCGAAGCTCTAGAAGTAGTTCTCGATGTGAAGAATCATCCGGTATTGATCCACTG
CAAACGCGGCAAGCATCGGACAGGTTGCGTCGTGGGGTGCTTGCGGAAACTGCAGAAGTGGTGCTTATCATCCATATTCGACGAGTACCAGAGGTTTGCA
GCAGCCAAGGCTAGAGTATCGGACCAAAGGTTTATGGAGCTGTTCGATGTTTCGAGCTTGAAGCACATTTCAATGTCGTTTTCATGCTCCAAGAGATAA
AA sequence
>Lus10015154 pacid=23149515 polypeptide=Lus10015154 locus=Lus10015154.g ID=Lus10015154.BGIv1.0 annot-version=v1.0
MKFDHNFTATGGGGGDQPPRFEDGEGSSSSEEEMCRTIEVSSVMINHRRGEDHLSDSPPPLCDELSFIPPLNFSMVDNGIFRSGFPDSANFSFLETLGLR
SIMVSRCLCPEPYPEANAEFLKANGIRLFQFGIEGYKEPFVNIPEDPIREALEVVLDVKNHPVLIHCKRGKHRTGCVVGCLRKLQKWCLSSIFDEYQRFA
AAKARVSDQRFMELFDVSSLKHISMSFSCSKR
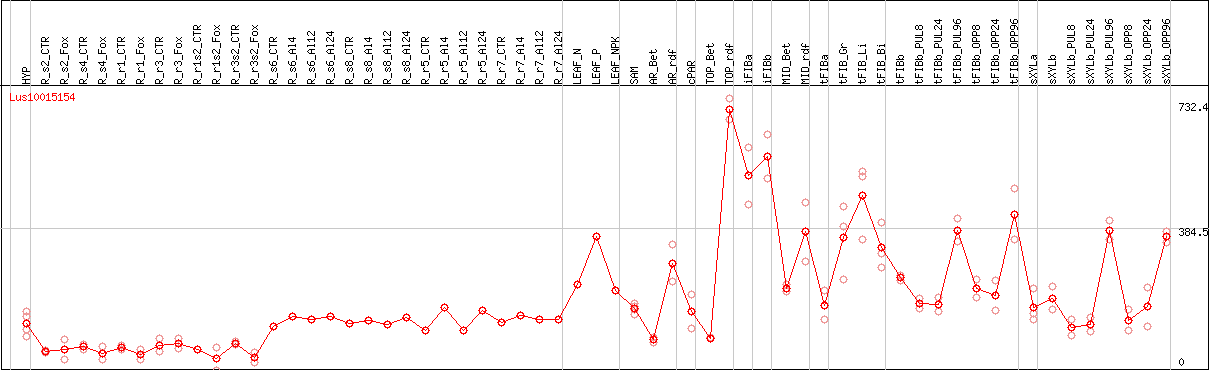
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10015154 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.