External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G15570 79 / 6e-17
Phototropic-responsive NPH3 family protein (.1)
AT1G52770 78 / 1e-16
Phototropic-responsive NPH3 family protein (.1)
AT5G67440 72 / 1e-14
MEL2, NPY3
NAKED PINS IN YUC MUTANTS 3, MAB4/ENP/NPY1-LIKE 2, Phototropic-responsive NPH3 family protein (.1.2)
AT3G08660 65 / 3e-12
Phototropic-responsive NPH3 family protein (.1)
AT3G26490 64 / 7e-12
Phototropic-responsive NPH3 family protein (.1)
AT3G08570 63 / 2e-11
Phototropic-responsive NPH3 family protein (.1)
AT2G14820 61 / 7e-11
MEL3, NPY2
NAKED PINS IN YUC MUTANTS 2, MAB4/ENP/NPY1-LIKE 3, Phototropic-responsive NPH3 family protein (.1)
AT5G48800 56 / 3e-09
Phototropic-responsive NPH3 family protein (.1)
AT2G47860 54 / 1e-08
SETH6
Phototropic-responsive NPH3 family protein (.1.2.3)
AT5G03250 53 / 4e-08
Phototropic-responsive NPH3 family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015169
326 / 2e-108
AT5G64330 264 / 7e-79
ROOT PHOTOTROPISM 3, NON-PHOTOTROPIC HYPOCOTYL 3, Phototropic-responsive NPH3 family protein (.1.2)
Lus10026046
82 / 5e-18
AT5G10250 639 / 0.0
DEFECTIVELY ORGANIZED TRIBUTARIES 3, Phototropic-responsive NPH3 family protein (.1)
Lus10014337
81 / 8e-18
AT5G10250 620 / 0.0
DEFECTIVELY ORGANIZED TRIBUTARIES 3, Phototropic-responsive NPH3 family protein (.1)
Lus10023274
68 / 4e-13
AT1G30440 910 / 0.0
Phototropic-responsive NPH3 family protein (.1)
Lus10038531
63 / 2e-11
AT1G30440 915 / 0.0
Phototropic-responsive NPH3 family protein (.1)
Lus10013799
61 / 1e-10
AT5G03250 672 / 0.0
Phototropic-responsive NPH3 family protein (.1)
Lus10037848
60 / 1e-10
AT3G15570 343 / 1e-115
Phototropic-responsive NPH3 family protein (.1)
Lus10011082
58 / 8e-10
AT5G67385 476 / 2e-162
Phototropic-responsive NPH3 family protein (.1)
Lus10025928
55 / 1e-08
AT5G48800 925 / 0.0
Phototropic-responsive NPH3 family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G024400
138 / 4e-38
AT5G10250 331 / 5e-106
DEFECTIVELY ORGANIZED TRIBUTARIES 3, Phototropic-responsive NPH3 family protein (.1)
Potri.005G075400
83 / 2e-18
AT5G10250 695 / 0.0
DEFECTIVELY ORGANIZED TRIBUTARIES 3, Phototropic-responsive NPH3 family protein (.1)
Potri.001G175400
81 / 1e-17
AT1G52770 507 / 8e-179
Phototropic-responsive NPH3 family protein (.1)
Potri.003G058800
80 / 1e-17
AT3G15570 499 / 1e-175
Phototropic-responsive NPH3 family protein (.1)
Potri.007G093000
80 / 3e-17
AT5G10250 706 / 0.0
DEFECTIVELY ORGANIZED TRIBUTARIES 3, Phototropic-responsive NPH3 family protein (.1)
Potri.017G048200
76 / 4e-16
AT5G64330 993 / 0.0
ROOT PHOTOTROPISM 3, NON-PHOTOTROPIC HYPOCOTYL 3, Phototropic-responsive NPH3 family protein (.1.2)
Potri.013G073700
76 / 6e-16
AT5G17580 409 / 4e-137
Phototropic-responsive NPH3 family protein (.1)
Potri.007G112600
72 / 9e-15
AT5G64330 989 / 0.0
ROOT PHOTOTROPISM 3, NON-PHOTOTROPIC HYPOCOTYL 3, Phototropic-responsive NPH3 family protein (.1.2)
Potri.010G223600
67 / 4e-13
AT5G03250 683 / 0.0
Phototropic-responsive NPH3 family protein (.1)
Potri.008G038600
64 / 6e-12
AT5G03250 723 / 0.0
Phototropic-responsive NPH3 family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03000
NPH3
NPH3 family
Representative CDS sequence
>Lus10015190 pacid=23149513 polypeptide=Lus10015190 locus=Lus10015190.g ID=Lus10015190.BGIv1.0 annot-version=v1.0
ATGGTGCATCAGGACTGCGAAAGGGGAGACTTGACTTCGAATCGATGGCTCGAAGAGGTTGCTTCTCTGCGAATCGATCATTTCGTTCGAGTTGTTGAGT
CAATGAAGTGGGAAGGAATGAAGCCAGAGATTGTAAGCTCATTTTTGGAACATTGGCTCTTGAAAGGGTCTCACCGGCTAGAAAATCTAACAATTGAGAC
ACTGAAAGTTGCCATAGAAGGACTGATAAGGCTGCTTCCTGAGCAAGAAACCTCTATACCTTGCAACATTCTGCTTTATCTTCTCAAGCTTGGAACTTCC
ATACGAGCCGACCCAGAGCTATTAGCGAGGAGTGAGGCGAGAGCGGTTCGTAAGCTAGATGGTTGTAGAATTTCGGACCTCCTGGTTCGAAATTCTACGG
GAGTCGGTGCTCTGTATGATGTGGAAATTGTTGCCAGAGTTGTGAAAGCATATGTTGCTTGTGTTGCAAGTAACCCAACATCGGGGTTGCCTAATGTTGG
AAGATTGGTTGATAAGTATTTGTGTTCGCGTACTTCTGCTTGGCGTACTCCTCCGCTGAAACAGCGGCAGCGAAGCGATTCCTCATTATCCTTTTTCTTA
TTCTTGACAATTGACTAA
AA sequence
>Lus10015190 pacid=23149513 polypeptide=Lus10015190 locus=Lus10015190.g ID=Lus10015190.BGIv1.0 annot-version=v1.0
MVHQDCERGDLTSNRWLEEVASLRIDHFVRVVESMKWEGMKPEIVSSFLEHWLLKGSHRLENLTIETLKVAIEGLIRLLPEQETSIPCNILLYLLKLGTS
IRADPELLARSEARAVRKLDGCRISDLLVRNSTGVGALYDVEIVARVVKAYVACVASNPTSGLPNVGRLVDKYLCSRTSAWRTPPLKQRQRSDSSLSFFL
FLTID
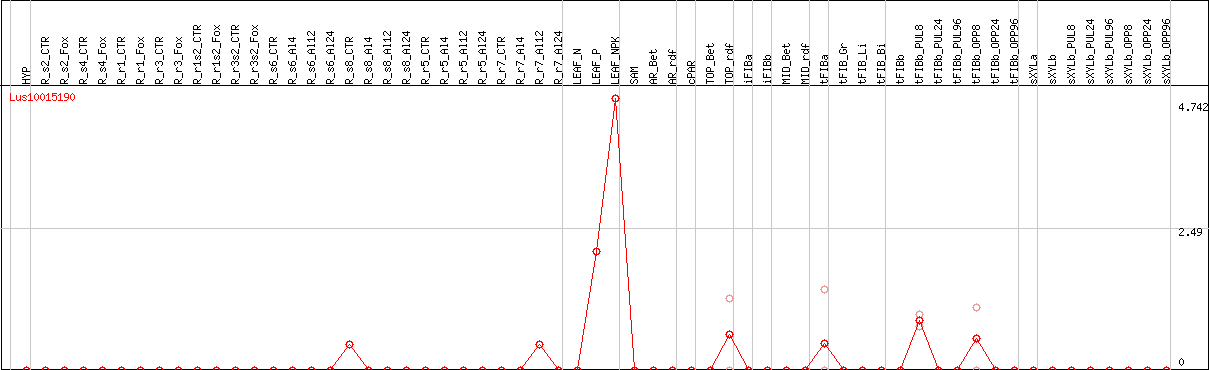
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10015190 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.