External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G26980 200 / 1e-63
ATSYP41, ATTLG2A, SYP41
syntaxin of plants 41 (.1.2)
AT4G02195 196 / 8e-62
ATTLG2B, ATSYP42, SYP42
syntaxin of plants 42 (.1)
AT3G05710 192 / 2e-60
ATSYP43, SYP43
syntaxin of plants 43 (.1.2)
AT5G46860 45 / 1e-05
SGR3, ATVAM3, ATSYP22, VAM3
SHOOT GRAVITROPISM 3, ARABIDOPSIS THALIANA VACUOLAR MORPHOLOGY 3, ARABIDOPSIS THALIANA SYNTAXIN OF PLANTS 22, Syntaxin/t-SNARE family protein (.1)
AT5G16830 42 / 0.0002
ATSYP21, ATPEP12, SYP21, PEP12P
syntaxin of plants 21 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031466
247 / 1e-81
AT3G05710 506 / 0.0
syntaxin of plants 43 (.1.2)
Lus10029864
235 / 6e-77
AT3G05710 510 / 0.0
syntaxin of plants 43 (.1.2)
Lus10020683
228 / 7e-75
AT3G05710 361 / 7e-126
syntaxin of plants 43 (.1.2)
Lus10000311
50 / 1e-07
AT5G46860 217 / 5e-72
SHOOT GRAVITROPISM 3, ARABIDOPSIS THALIANA VACUOLAR MORPHOLOGY 3, ARABIDOPSIS THALIANA SYNTAXIN OF PLANTS 22, Syntaxin/t-SNARE family protein (.1)
Lus10011064
50 / 8e-07
AT1G45616 366 / 2e-111
receptor like protein 6 (.1)
Lus10040094
49 / 9e-07
AT5G46860 419 / 1e-149
SHOOT GRAVITROPISM 3, ARABIDOPSIS THALIANA VACUOLAR MORPHOLOGY 3, ARABIDOPSIS THALIANA SYNTAXIN OF PLANTS 22, Syntaxin/t-SNARE family protein (.1)
Lus10030953
49 / 9e-07
AT5G46860 417 / 9e-149
SHOOT GRAVITROPISM 3, ARABIDOPSIS THALIANA VACUOLAR MORPHOLOGY 3, ARABIDOPSIS THALIANA SYNTAXIN OF PLANTS 22, Syntaxin/t-SNARE family protein (.1)
Lus10042527
47 / 7e-06
AT1G08560 391 / 2e-137
KNOLLE, syntaxin of plants 111 (.1)
Lus10021988
46 / 1e-05
AT1G08560 388 / 2e-136
KNOLLE, syntaxin of plants 111 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G020800
223 / 2e-72
AT3G05710 462 / 1e-164
syntaxin of plants 43 (.1.2)
Potri.013G011100
210 / 2e-67
AT3G05710 457 / 7e-163
syntaxin of plants 43 (.1.2)
Potri.002G200400
201 / 6e-64
AT5G26980 397 / 1e-139
syntaxin of plants 41 (.1.2)
Potri.001G138500
46 / 8e-06
AT5G46860 377 / 3e-133
SHOOT GRAVITROPISM 3, ARABIDOPSIS THALIANA VACUOLAR MORPHOLOGY 3, ARABIDOPSIS THALIANA SYNTAXIN OF PLANTS 22, Syntaxin/t-SNARE family protein (.1)
Potri.003G095300
45 / 2e-05
AT5G46860 368 / 2e-129
SHOOT GRAVITROPISM 3, ARABIDOPSIS THALIANA VACUOLAR MORPHOLOGY 3, ARABIDOPSIS THALIANA SYNTAXIN OF PLANTS 22, Syntaxin/t-SNARE family protein (.1)
Potri.019G030800
45 / 2e-05
AT1G08560 431 / 3e-153
KNOLLE, syntaxin of plants 111 (.1)
Potri.013G053200
44 / 5e-05
AT1G08560 430 / 8e-153
KNOLLE, syntaxin of plants 111 (.1)
Potri.014G145200
43 / 7e-05
AT5G46860 263 / 4e-88
SHOOT GRAVITROPISM 3, ARABIDOPSIS THALIANA VACUOLAR MORPHOLOGY 3, ARABIDOPSIS THALIANA SYNTAXIN OF PLANTS 22, Syntaxin/t-SNARE family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF05739
SNARE
SNARE domain
Representative CDS sequence
>Lus10015215 pacid=23149517 polypeptide=Lus10015215 locus=Lus10015215.g ID=Lus10015215.BGIv1.0 annot-version=v1.0
ATGGCGACGAGGAACCGGACGTTGATATTCAGAAAGTACAGAGACGCGTTGAAGAGCGTACGGATTCCGACGAGCTCCTCTGCCTCCGCGGCTTCGACGA
GCTCTTCTTCCGGAGGAGGTGGTCCTGTGATTGAGTTGGTTAGCACTTCGCTTCTCAATCATAAACGTTCTTATGCTCCTCTCAGTACTGAAGATCCTGG
CAATTCCAGTAAGGGAGCAGTAACAGTCGGTCTCCCACCTGCCTGGCGATCCCTTGCCACTGACCTCCAGAACCTTTCAATGGAGCTTCGCAAGAAACAA
TCGACTTACTTGAAGCGCCTCAGGCAGCAAAAAGAGGGTCAAGAAGGGGATGACTTGGAGATGAACCTTAATGGCAATAGATCCCGAATGGACGATGATG
ATGGCTTAGATGACATGGTCGTGGAGTCTGTAAATGAACTTGCTCAAATCATGAAGGACTTGTCAGTACTTGTGATAGACCAGGGTACGATCGTTGATAG
AATAGACTACAACGTTCAGAACGTCGCAACTACAGTCGAAGACGGGCTTAAACAGCTGCAGAAGGCCGAAAGATCGCAAAAGCAAGGTGGAATGGTAATG
TGTGCGACTGTGCTTGTGATCATGTGCTTCATCATGTTGGTTCTCCTGATCCTTAAGGAGATATTGTTCTGA
AA sequence
>Lus10015215 pacid=23149517 polypeptide=Lus10015215 locus=Lus10015215.g ID=Lus10015215.BGIv1.0 annot-version=v1.0
MATRNRTLIFRKYRDALKSVRIPTSSSASAASTSSSSGGGGPVIELVSTSLLNHKRSYAPLSTEDPGNSSKGAVTVGLPPAWRSLATDLQNLSMELRKKQ
STYLKRLRQQKEGQEGDDLEMNLNGNRSRMDDDDGLDDMVVESVNELAQIMKDLSVLVIDQGTIVDRIDYNVQNVATTVEDGLKQLQKAERSQKQGGMVM
CATVLVIMCFIMLVLLILKEILF
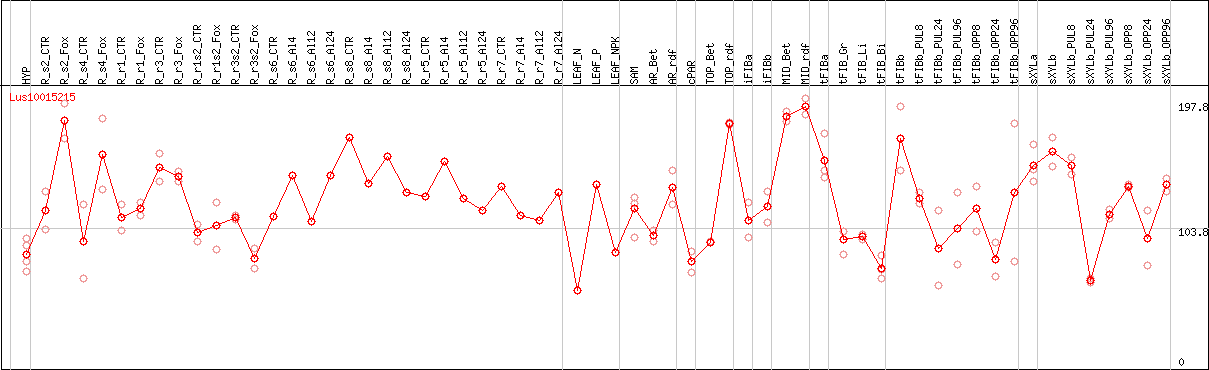
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10015215 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.