External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G18040 241 / 2e-81
LSP1, CUM1, AT.EIF4E1, EIF4E
eukaryotic translation Initiation Factor 4E1, CUCUMOVIRUS MULTIPLICATION 1, ARABIDOPSIS THALIANA EUKARYOTIC TRANSLATION INITATION FACTOR 4E1, eukaryotic translation initiation factor 4E (.1)
AT1G29550 209 / 2e-68
Eukaryotic initiation factor 4E protein (.1)
AT1G29590 206 / 6e-67
eIF4E3
eukaryotic translation Initiation Factor 4E3, Eukaryotic initiation factor 4E protein (.1.2)
AT5G35620 131 / 1e-38
eIFiso4E, EIF(ISO)4E, EIF(ISO)4E, EIF4E2, EIF(ISO)4E, LSP1, LSP, EIF(ISO)4E, EIF(ISO)4E, EIF(ISO)4E
LOSS OF SUSCEPTIBILITY TO POTYVIRUS 1, eukaryotic translation Initiation Factor isoform 4E, EUKARYOTIC TRANSLATION INITATION FACTOR 4E2, EUKARYOTIC INITIATION FACTOR \(ISO\)4E, Eukaryotic initiation factor 4E protein (.1.2)
AT5G18110 82 / 5e-19
NCBP
novel cap-binding protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015247
294 / 4e-102
AT4G18040 322 / 1e-112
eukaryotic translation Initiation Factor 4E1, CUCUMOVIRUS MULTIPLICATION 1, ARABIDOPSIS THALIANA EUKARYOTIC TRANSLATION INITATION FACTOR 4E1, eukaryotic translation initiation factor 4E (.1)
Lus10005410
292 / 2e-101
AT4G18040 318 / 3e-111
eukaryotic translation Initiation Factor 4E1, CUCUMOVIRUS MULTIPLICATION 1, ARABIDOPSIS THALIANA EUKARYOTIC TRANSLATION INITATION FACTOR 4E1, eukaryotic translation initiation factor 4E (.1)
Lus10011988
246 / 3e-83
AT4G18040 315 / 4e-110
eukaryotic translation Initiation Factor 4E1, CUCUMOVIRUS MULTIPLICATION 1, ARABIDOPSIS THALIANA EUKARYOTIC TRANSLATION INITATION FACTOR 4E1, eukaryotic translation initiation factor 4E (.1)
Lus10002785
241 / 5e-81
AT4G18040 313 / 7e-109
eukaryotic translation Initiation Factor 4E1, CUCUMOVIRUS MULTIPLICATION 1, ARABIDOPSIS THALIANA EUKARYOTIC TRANSLATION INITATION FACTOR 4E1, eukaryotic translation initiation factor 4E (.1)
Lus10011776
156 / 3e-48
AT5G35620 263 / 4e-90
LOSS OF SUSCEPTIBILITY TO POTYVIRUS 1, eukaryotic translation Initiation Factor isoform 4E, EUKARYOTIC TRANSLATION INITATION FACTOR 4E2, EUKARYOTIC INITIATION FACTOR \(ISO\)4E, Eukaryotic initiation factor 4E protein (.1.2)
Lus10023733
156 / 4e-48
AT5G35620 261 / 2e-89
LOSS OF SUSCEPTIBILITY TO POTYVIRUS 1, eukaryotic translation Initiation Factor isoform 4E, EUKARYOTIC TRANSLATION INITATION FACTOR 4E2, EUKARYOTIC INITIATION FACTOR \(ISO\)4E, Eukaryotic initiation factor 4E protein (.1.2)
Lus10020282
84 / 6e-20
AT5G18110 347 / 1e-122
novel cap-binding protein (.1)
Lus10005704
81 / 8e-19
AT5G18110 345 / 6e-122
novel cap-binding protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G077200
237 / 8e-80
AT4G18040 299 / 1e-103
eukaryotic translation Initiation Factor 4E1, CUCUMOVIRUS MULTIPLICATION 1, ARABIDOPSIS THALIANA EUKARYOTIC TRANSLATION INITATION FACTOR 4E1, eukaryotic translation initiation factor 4E (.1)
Potri.008G171100
172 / 1e-54
AT5G35620 249 / 5e-85
LOSS OF SUSCEPTIBILITY TO POTYVIRUS 1, eukaryotic translation Initiation Factor isoform 4E, EUKARYOTIC TRANSLATION INITATION FACTOR 4E2, EUKARYOTIC INITIATION FACTOR \(ISO\)4E, Eukaryotic initiation factor 4E protein (.1.2)
Potri.010G066700
164 / 2e-51
AT5G35620 254 / 1e-86
LOSS OF SUSCEPTIBILITY TO POTYVIRUS 1, eukaryotic translation Initiation Factor isoform 4E, EUKARYOTIC TRANSLATION INITATION FACTOR 4E2, EUKARYOTIC INITIATION FACTOR \(ISO\)4E, Eukaryotic initiation factor 4E protein (.1.2)
Potri.013G057000
85 / 2e-20
AT5G18110 358 / 6e-127
novel cap-binding protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0625
eIF4e
PF01652
IF4E
Eukaryotic initiation factor 4E
Representative CDS sequence
>Lus10015251 pacid=23178788 polypeptide=Lus10015251 locus=Lus10015251.g ID=Lus10015251.BGIv1.0 annot-version=v1.0
ATGGCGGTGGAGGAGTCGCAGAAACAGGTGACCACCGAAGAGGCCGCCGCGAACGCTAACCCTAAGGTCCAATCCGACGACGATGTTCTGGAGGAAGGAG
AGATAGTCGGAGGAGGCGGCGGAGAATTAATGTTACCTGAGCTTAATACATCTAATTCTCTAACTTTACCCCTCTGTTGCAGCTTGTACAATAACATTCA
TCATCCCAGCAAGCTTGCACAAGGAGCAGACTTCTATTGCTTTAAAAACAAGATTGAGCCCAAGTGGGAGGACCCTATCTGTGCAAATGGAGGAAGCTGG
AAGATAACCTATCCAAAAGGGAAATCTGATACACCATGGCTGTACACACTGCTAGCAATGATTGGTGAGCAGTTCGATCACGGTGACGAGATTTGCGGTG
CTGTTGTCAATGTAAGAGCCAGGCAGGAGAGGGTTTCTATTTGGACCAAGAATGCTGCAAACGAAGCTGCTCAGCTGAGTATCGGGAGACAGTGGAAGGA
GTTCCTCGATAACAACGACAGTATTGGGTTTGTCACTCACGACGATGCAAAGAAGAATGACAGAAACGCAAAGAATCGCTATAATGCCTAA
AA sequence
>Lus10015251 pacid=23178788 polypeptide=Lus10015251 locus=Lus10015251.g ID=Lus10015251.BGIv1.0 annot-version=v1.0
MAVEESQKQVTTEEAAANANPKVQSDDDVLEEGEIVGGGGGELMLPELNTSNSLTLPLCCSLYNNIHHPSKLAQGADFYCFKNKIEPKWEDPICANGGSW
KITYPKGKSDTPWLYTLLAMIGEQFDHGDEICGAVVNVRARQERVSIWTKNAANEAAQLSIGRQWKEFLDNNDSIGFVTHDDAKKNDRNAKNRYNA
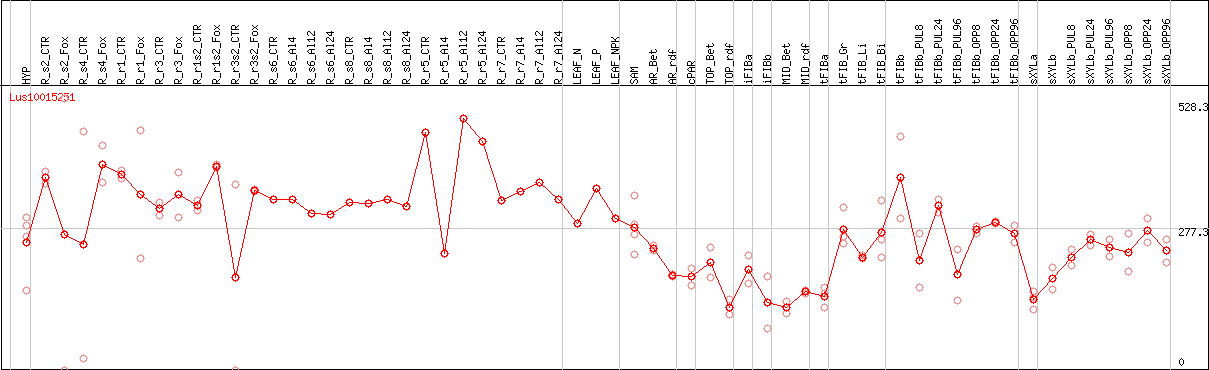
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10015251 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.